The male testes are important areas for biological innovation, as well as being a factory for sperm. In exciting new biological research a team of scientists has found that the testes are also a hotspot for the production of new genes.
The researchers at Rockefeller University used fruit flies to get important information into how nature’s attempts at innovation manifests during sperm development. Published in the journal eLife, the study shows how mapping DNA mutations at the single-cell level, and new gene activity from these changes, could shed light on evolution.
“Our work offers an unprecedented perspective on a process that enables living things to adapt and evolve, and that ultimately contributes to the diversity of life on Earth,” Li Zhao, lead researcher and assistant professor at Rockefeller University, said.
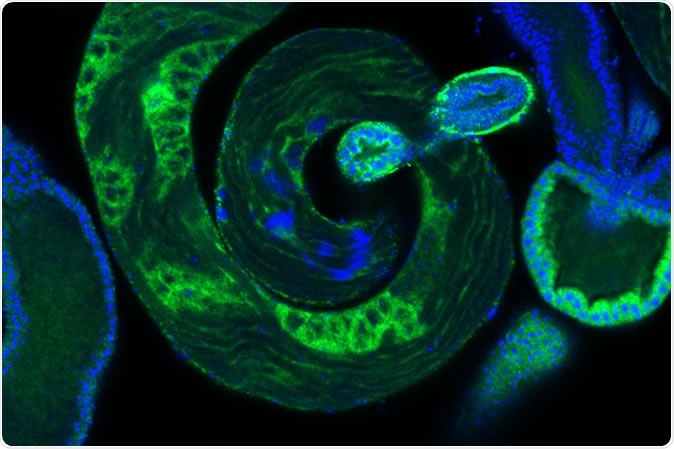
Developing sperm, in blue, within a fruit fly testis. IMAGE CREDIT Laboratory of Evolutionary Genetics and Genomics at The Rockefeller University
Hotspot for newly originated genes
A hypothesis states that the testis catalyzes the birth and preservation of novel genes. These novel functional genes that contain beneficial products are selectively preserved. In time, they evolve more advanced regulatory programs. Over the past ten years, many research studies pointed out that young genes, including the de novo originated genes, which are genes born from ancestrally noncoding DNA, tend to be biased toward the testis.
Moreover, in recent years, studies in flies and even humans have shown many young genes arising from the testes. Hence, many scientists state that the testes rank among the most productive sites in the body for genetic innovation.
However, there are risks involved in the mass production of genetic novelties. For instance, a father’s sperm gets two to three times more mutations that than of a mother’s egg cell. Hence, during development, the sperm acquires more genetic mistakes, and this can harm the offspring when the egg gets fertilized. Put simply, males can lose an important thing in the evolution game – the ability to proliferate their gene pools into the next generations.
Study findings
To arrive attheir findings, the team of researchers leveraged single-cell RNA-sequence and unsupervised clustering to determine all the main types and classes of cells in the sperm lineage. They identified populations of somatic cells, which include hub cells, stem cells, and terminal epithelial cells.
The team found that the overall gene expression is very active in early spermatogenesis, the production or development of mature sperm cells, and lowers throughout the process of spermatogenesis. Aside from these, when the team studied de novo genes, they revealed complex patterns. For example, some genes showed up primarily in some cell types, but not in others.
In fact, about 15 percent of the genes appeared early on during the stem cell stage. This is a surprising discovery since scientists in the past thought that the new genes rarely show up during the early stage of development. They found that the most active de novo genes happened in the middle of the process, otherwise known as the spermatocyte phase of developing sperm.
“Our finding that young duplicated genes have different expression patterns than de novo genes merits further study. Young duplicated genes are more likely to be bimodally expressed than de novo genes of a similar age,” the researchers wrote in the study.
“Our result indicates that cell-type-specific mutational load can be estimated from single-cell RNA-seq data with reasonable accuracy. Overall, we provide novel insights into the dynamics of mutation, repair, and de novo gene expression profiles in the male germline,” they concluded.
Now, scientists want to understand the purpose of de novo genes when they first develop. They suspect these new genes play a pivotal role in sperm cell maturation.
“Precisely what these de novo genes are doing to move development along is an exciting open question,” Zhao said.
Journal reference:
Witt, Evan, Benjamin, S., Svetec, N., and Zhao, L. (2019). Testis single-cell RNA-seq reveals the dynamics of de novo gene transcription and germline mutational bias in Drosophila. eLife. https://elifesciences.org/articles/47138