The millions of bacteria residing in the gut play a very important role in health and in disease. However, a constant issue has been the lack of understanding of the actual composition of the healthy human gut microbiome.
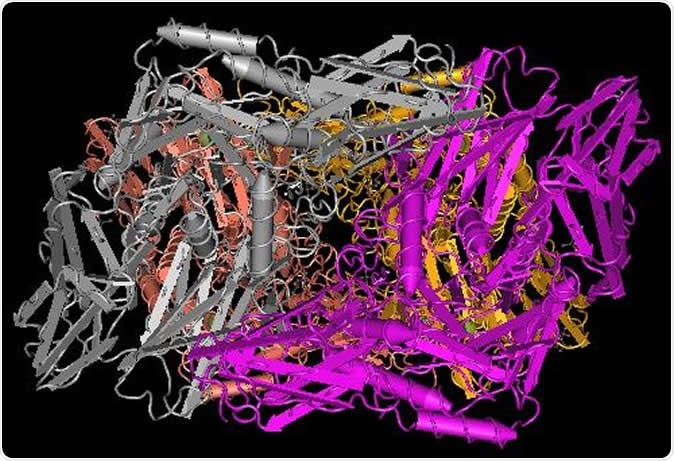
Crystal structure of putative beta-galactosidase from Bacteroides fragilis. Image Credit: National Institutes of Health
Micro-organisms, or microbes, have been around for a very long time, and shape both the external and internal environment of human beings. The word “microbiome” was coined by Joshua Lederberg in 2001, as a way to focus the attention of scientists on the multiple interactions between the microbes dwelling on and in our bodies, and their interplay with our human physiology. The term has now been defined as “a multi-species community of microorganisms in any environment: host, habitat, or ecosystem.” Far from being simply invaders bent on our destruction, the human microbiome comprises a whole living world in itself, bringing a complete and highly diverse set of genes that interact and change, and also have an impact on human health. This microbial genetic mix is called the metagenome. The Human Microbiome Project (HMP) took off in 2008 and has helped catalyse greater characterization and understanding of how these communities function.
Gut microbiome research requires high-throughput accurate data collection and analysis, as well as facilities to integrate the processed data in an organized fashion for storage, sharing and access among research groups. Most earlier studies focused on specific genes or groups of organisms, leaving out large segments of the microbial genomes. Also, different reference standards have led to a variety of statements on the gut composition.
Bacteria Bacteroides fragilis, one of the major components of normal microbiome of human intestine, 3D illustration Credit: Kateryna Kon / Shutterstock
In fact, most studies on the metagenome use only a small reference set of nucleic acid sequences from already selected microbes or microbial genes. This is due to the difficulty of pairing the experimental data with the complete nucleotide database available with the NCBI (NCBI-nt). However, new algorithms can now make use of the latter to permit a more accurate analysis of experimental data to yield a microbial abundance profile.
The present work builds on this foundation to form a Gut Feeling Knowledge Base–GutFeelingKB - with samples from a healthy set of participants. These gut microbiota samples were sequenced to get a picture of what a healthy gut metagenome looks like. The sample number was filled out using 50 more randomly selected sequences from the HMP.
The researchers also collected assembled contiguous sequences, or contigs, which do not correspond to any NCBI-nt sequence but can be detected in healthy fecal samples. Contigs are thus dark matter, not recognizable by any known nucleic acid sequence, but which can be built into sequences of 10,000 nucleotides or longer. This length was chosen to reduce the number of foreign (artefactual) dark matter contigs while still including microbial dark matter. The GutFeelingKB is thus a comprehensive knowledge base on the healthy human gut microbiome.
This was then used as a reference to construct a standard reporting template where individual microbiomes can be reported, to allow direct comparison of results between studies and samples.
The scientists also fashioned a new workflow using several computer programs and a filtered gut microbial sequence database called Filtered-nt, containing almost 35,000,000 sequences that allow biologically relevant interpretation of sample sequences, while offering assurance that the whole of the known sequence space has been included.
- The CensuScope algorithm first identifies the organisms to provide a microbial profile of the sample. If any of the organisms identified are not already in GutFeelingKB they are added in.
- Secondly, HIVE-hexagon is used to create a read-map for all the reads in each sample by aligning short reads quickly and accurately with known sequences. This gives the abundance profiles.
- Finally, IDBA-UD is used to assemble the reads. If they are 10 000 nucleotides in length or longer, they are included in the dark matter of the metagenome. Adding this to the previous analysis gives the sum of the known and unknown organisms in the microbiome.
- MaAsLin was also used to associate dietary factors with the bacterial abundance profile, factoring in variations in the diet of each individual from day to day as well as between different individuals.
Thus, GutFeelingKB represents a thoroughly curated collection of nucleotide sequences with metadata from 157 organisms in 60 genera.
The healthy human microbiome thus contains members from 8 phyla, 18 families, 60 genera and 109 species, mostly from the Firmicutes (40%) and Bacteroidetes (20%) phyla. Another 20% come from Actinobacteria. Among Firmicutes, over half are Clostridia, closely followed by Bacteroides, Bifidobacteria, Enterobacteria and Lactobacilli.
All the samples were positive for 84 out of the 109 organisms, which perhaps represents the core species list.
However, it is important to note that around the world, specific organisms that are not on the GutFeelingKB have been mapped, such as Fusobacteria, certain Actinobacter and Bacteroides species. The function of this platform will be to act as a launching pad to compare the results of sample analysis from healthy individuals, and give more insights into the variations observed under the influence of dietary factors, illnesses and medications.
Organisms in the gut
Bacteroides is the single most abundant genus in many countries, in health. These are generally beneficial inside the gut but if they escape, they seize the chance to cause infections that are often drug-resistant and may lead to a 20% fatality rate. However, in the gut they are protective against other pathogens and help break down carbohydrates in the diet.
Similarly, Bifidobacteria are among the first settlers of the gut, are often found in probiotics, and produce the important short-chain fatty acid (SCFA) acetate that strengthens the gut epithelial barrier against infection. One strain of Bifidobacterium longum has been found in one individual in a particularly long-lived people group in China.
Diet and nutrients – how do they associate with gut bacteria?
Bifidobacterium numbers increase with a higher protein intake, and especially with vegetable protein. Dietary soluble fiber also promotes its growth. Akkermansia is linked to saturated fat and linoleic acid, but negatively associated with polyunsaturated fatty acids (PUFA). Bacteroides ovatus grows in number with increased food intake, obesity and waist circumference. These examples help us understand how these numbers can be used to take health action to correct microbiome imbalances in the future.
FecalBiome template
The reporting template published in the current study is intended to replace the non-standardized formats used by different commercial and research groups, which hampers their interpretation and comparison. It converts the research data into a clinical report, helping to make it immediately actionable.
FecalBiome has three domains, Sample, Patient and Result, resembling a metabolic panel report. It also allows collaborators to share a lot of information quickly, when research happens across a wide range of locations. The threshold for reporting can be individually set as per the purpose of the study. It reports the abundance, the average abundance and information on the microbes present.
Summary
The present report thus allows gut bacterial research to be connected to understandable health information, making it relevant for medical practice and the patient. It also allows easy comparison across studies. It will help gut replacement products to be reviewed more rapidly, and help evidence-based medicine to progress.
The study was published on September 11, 2019, in PLOS ONE.
Journal reference:
Baseline human gut microbiota profile in healthy people and standard reporting template. Charles H. King, Hiral Desai, Allison C. Sylvetsky, Jonathan LoTempio, Shant Ayanyan, Jill Carrie, Keith A. Crandall, Brian C. Fochtman, Lusine Gasparyan, Naila Gulzar, Paul Howell, Najy Issa, Konstantinos Krampis, Lopa Mishra, Hiroki Morizono, Joseph R. Pisegna, Shuyun Rao, Yao Ren, Vahan Simonyan, Krista Smith, Sharanjit VedBrat, Michael D. Yao, & Raja Mazumder. PLOS ONE. https://doi.org/10.1371/journal.pone.0206484. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0206484