Researchers have developed a blood test that could accurately detect up to 50 types of cancers, and some of these detections could be even before the symptoms appear. The test would help physicians detect the origin or the source of cancer within the body explain the researchers from the European Society of Oncology. The study titled, "Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA," was published in the latest issue of the journal Annals of Oncology.
The researchers noted that this new test has a 0.7 percent false-positive rate for cancer detection and soon could be part of the cancer screening programs in different nations. They found that 96 percent of the time, the test could predict the tissue from which cancer had started, and the accuracy rate was 93 percent.
What was this study about?
The researchers and all cancer experts have time and again emphasized upon detection of cancers early, which could help begin treatment before cancer has invaded major areas of the body and thus improve prognosis. This was a prospective case-control study that looked at a special test called the "targeted methylation analysis of circulating cell-free DNA (cfDNA)" which would help detect cancer cells originating from different parts or cells of the body. It was part of the Circulating Cell-free Genome Atlas (CCGA) study.
The team explained that every tumor sheds individual DNA particles in the blood, and these are called cfDNA. The task of looking for a cfDNA that has come from a tumor cell and from a regular cell is difficult, they wrote. This new blood test can do just that. It looks at the chemical change called methylation on the DNA. These methylation processes can control the gene expressions in normal DNA. Abnormal patterns of these DNA methylations could be detected using this test, and this can demarcate a cfDNA that has come from a tumor. This process could then help detect the location of the origin of the tumor.
Senior author of the paper, Dr. Michael Seiden (MD, Ph.D.), President of US Oncology (Texas, USA), explained, "Our earlier research showed that the methylation approach outperformed both whole-genome and targeted sequencing in the detection of multiple deadly cancer types across all clinical stages, and in identifying the tissue of origin. It also allowed us to identify the most informative regions of the genome, which are now targeted by the refined methylation test that is reported in this paper."
What was done?
For this study, the team of researchers included 6,689 participants, of which 2,482 had cancers of over 50 different types, and 4,207 participants did not have cancer. The whole group of participants was divided into two sets – a training set and a validation set. There were 3,052 participants in the training set with 1,531 cancer patients and 1,521 participants without cancer. In the validation set, there were 1,264 participants, 654 with cancer, and 610 without cancer.
For each of the participants, they collected the cfDNA, and on these samples, they performed bisulfite sequencing to find out over 100,000 informative methylation regions. The blood test targets approximately one million of the 30 million methylation sites in the human genome. A machine learning classifier (an algorithm) was used to predict the presence of cancer and the type of cancer based on the patterns of methylation in the cfDNA shed by tumors. The classifier was trained using a methylation database of cancer and non-cancer signals in cfDNA. The database is believed to be the largest in the world and is owned by the company involved in this research, GRAIL, Inc. (California, USA).
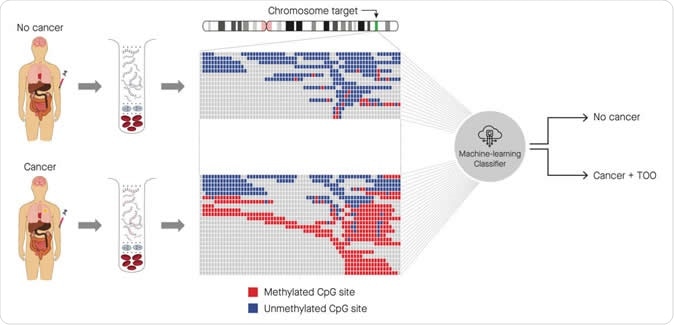
Identification of cancer status for more than 50 cancer types, as well as tissue of origin localization, from a single blood draw. Cell-free DNA is isolated from blood samples drawn from a patient without cancer (top) or with cancer (bottom), and subjected to a targeted methylation sequencing assay. Sequencing results identifying methylated (red) or unmethylated (blue) CpG regions are fed into a machine-learning classifier that can identify the presence or absence of cancer, as well as identify the tissue of origin (TOO).
What was found?
In both training and validation sets, the team noted that the test performed equally. The false positivity rate was 0.7 percent among the validation set of samples. This meant that in less than one person per 100 tested; there may be a mistaken positive result in the absence of cancer. The specificity of the test to detect the tissue of origin (TOO) was 99.3 percent wrote the researchers.
The researchers wrote that in 12 types of cancers, "anus, bladder, colon/rectum, esophagus, head and neck, liver/bile-duct, lung, lymphoma, ovary, pancreas, plasma cell neoplasm, stomach," the sensitivity of the test or ability to detect cancer correctly was 67.3 percent in stage 1 cancers. These cancers constitute the reason for 63 percent of deaths in the United States per year, said the researchers. The sensitivity was 43.9 percent for all cancer types. With increasing cancer stages, the sensitivity rose with 18 percent, 43 percent, 81 percent, and 93 percent in stages I, II, III, and IV, respectively. TOO could be predicted in 96 percent of all samples that were flagged with suspicion of cancer. Localization of the tissue of origin for these cancers was accurately predicted in 93 percent cases.
Conclusions
Researchers wrote in conclusion, "cfDNA sequencing leveraging informative methylation patterns detected more than 50 cancer types across stages."
Dr. Seiden said, "These data support the ability of this targeted methylation test to meet what we believe are the fundamental requirements for a multi-cancer early detection blood test that could be used for population-level screening: the ability to detect multiple deadly cancer types with a single test that has a very low false-positive rate, and the ability to identify where in the body the cancer is located with high accuracy to help healthcare providers to direct next steps for diagnosis and care."
Future directions and implications
Dr. Seiden added, "Considering the burden of cancer in our society, it is important that we continue to explore the possibility that this test might intercept cancers at an earlier stage and, by extension, potentially reduce deaths from cancers for which screening is either not available or has poor adherence. To our knowledge, this is the largest clinical genomics study, in participants with and without cancer, to develop and validate a blood test for early detection of multiple cancers." The authors wrote, "Considering the potential value of early detection in deadly malignancies, further evaluation of this test is justified in prospective population-level studies." They speculate that this test could be a part of the screening programs for cancer and could benefit large populations in the future.
Editor-in-chief of Annals of Oncology, Professor Fabrice André, Director of Research at the Institut Gustave Roussy, Villejuif, France, in a statement said, "This is a landmark study and a first step toward the development of easy-to-perform screening tools. Earlier detection of more than 50% of cancers could save millions of lives every year worldwide and could dramatically reduce morbidity induced by aggressive treatments. While numbers are still small, the performance of this new technology is particularly intriguing in pancreatic cancer, for which mortality rates are very high because it is usually diagnosed when it's at an advanced stage."
Journal reference:
Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA Liu, M.C.Liu, Minetta C.Cummings, Steven R.Absalan, FarnazCurtis, Christina et al. Annals of Oncology, https://www.annalsofoncology.org/article/S0923-7534(20)36058-0/fulltext