A team of researchers in the United States have developed highly specific and sensitive assays that could improve understanding of the antibody response in coronavirus disease 2019 (COVID-19) and help to determine the effectiveness of vaccines.
The assays uncovered dynamic changes in the level of antibodies against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). They could be vital in helping researchers better understand the humoral response and potential treatments for COVID-19.
Using the assays, the researchers were able to correlate levels of antibodies specific to SARS-CoV-2 Spike protein with neutralization titers. They could, therefore, be used as a proxy for neutralization in the clinical setting, say Derya Unutmazy (University of Connecticut School of Medicine) and colleagues.
The Spike protein is the surface structure that binds angiotensin-converting enzyme 2 (ACE2) on host cells before undergoing proteolytic cleavage to enable viral fusion and entry across the cell membrane.
A pre-print version of the paper is available on the server medRxiv*, while the article undergoes peer review.
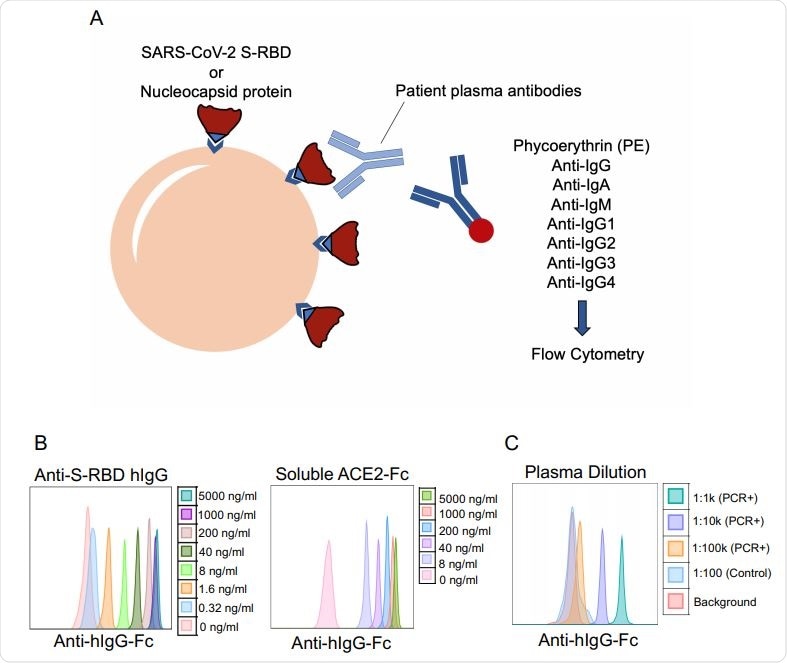
SARS-CoV-2 specific antibody detection assay. a. Illustration of antibody detection assay. Biotinylated S-RBD or Nucleocapsid proteins are captured by streptavidin-coated beads, then incubated with plasma samples and stained with PE-conjugated anti-IgG, IgA, IgM, IgG1, IgG2, IgG3, IgG4 antibodies. Fluorescence intensity analyzed by flow cytometry. b. Histogram overlays demonstrating the detection of anti-S-RBD human IgG antibody (left) and soluble ACE2-Fc (right) as positive controls for plasma antibody assay. c. Representative patient plasma titration. Healthy control plasma at 1:100 dilution was used as a negative control. Serial dilutions used were used in the flow cytometry overlay.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Challenges faced in predicting the immune response
Antibodies that target the Spike protein have the potential to neutralize viral entry and are considered key in the protective immune response against SARS-CoV-2 infection and COVID-19.
Predicting this immune response requires a clear understanding of the quantity, quality, and duration of the antibody responses during different stages of COVID-19 and in the convalescent period.
“In this regard, assessing the level of neutralizing antibodies (NAbs) that block viral entry into cells could be a critical parameter in determining protection from CoV-2 and management of convalescent plasma therapies, which are being tested as a COVID-19 treatment option,” explain the authors.
The team says current techniques for detecting SARS-CoV-2 antibodies lack the dynamic range and sensitivity needed to determine the magnitude of the humoral immune response accurately.
Assessing neutralization activity in plasma also requires assays that tend to rely on live virus replication, meaning a high-level biohazard security level laboratory is needed.
Sensitive and robust antibody and virus neutralization assays that enable the testing and monitoring of large numbers of infected or recovered patients are therefore needed.
Now, Unutmazy and colleagues have developed a highly sensitive bead-based fluorescent immunoassay for the measurement of SARS-CoV-2-specific antibody levels and a robust Spike protein pseudovirus to enable the measurement of neutralizing antibodies (Nab) levels in the plasma of patients with COVID-19.
Highly sensitive and detection
The team reports that antibodies specific to the receptor-binding domain (RBD) of the Spike protein were detected with very high sensitivity, at up to a 100,000-fold dilution of the plasma.
Spike RBD-specific immunoglobulin G (IgG) antibodies at titers of more than 1:100,000 were detectable in all cases of infection and were not detectable in virus-negative controls.
The total antibody level and neutralization titers differed significantly between patients with severe COVID-19 and outpatient or convalescent donors.
“Hospitalized patients had up to 3000-fold higher antibody and neutralization titers compared to outpatients or convalescent plasma donors,” writes the team.
Antibody and neutralization titers also significantly differed according to the presence of risk factors for severe outcomes. Patients with the most risk factors for severe outcomes such as older age and co-morbidity had the highest antibody titers and neutralization activity. This was also the case among patients with fatal COVID-19.
“It is, therefore, possible that surviving COVID-19 may require non-antibody dependent factors or that producing too much antibody may even have deleterious effects,” say Unutmazy and colleagues.
Would plasma transfer necessarily benefit patients?
Notably, the majority of convalescent donors’ levels of neutralizing antibodies were at least one order of magnitude lower than among hospitalized patients. They would be considered suitable recipients of plasma transfer from such donors.
“This finding raises the question of whether convalescent plasma transfers may actually provide benefit to severe COVID-19 patients by providing neutralizing antibodies,” writes the team. “It may perhaps be more beneficial to identify donors with much higher neutralizing antibody titers for the plasma donation.”
The researchers point out that this highlights the importance of having assays with a broad dynamic range that can accurately measure antibody responses in patients or seropositive individuals.
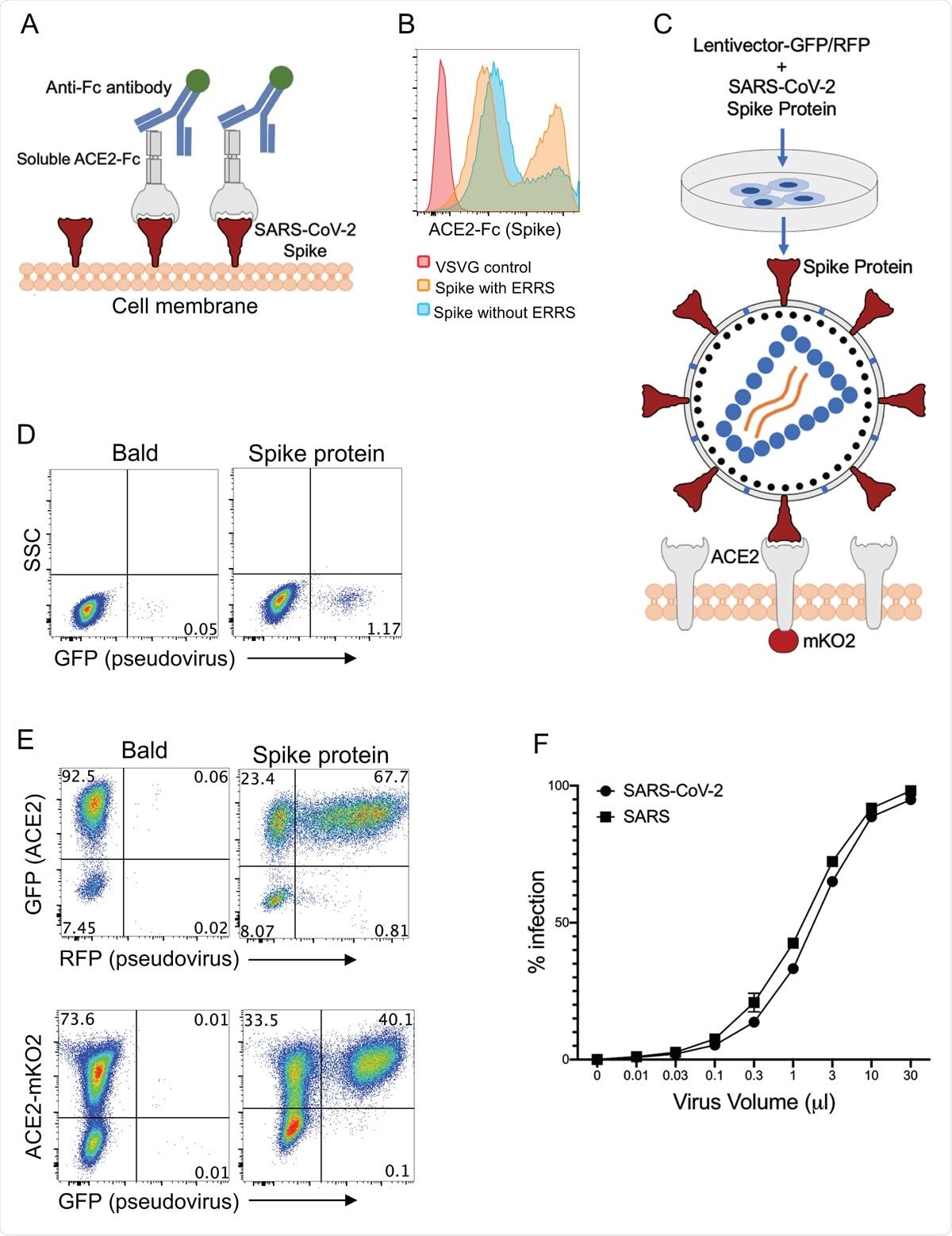
Development of CoV-2 and SARS Spike protein pseudotyped lentiviruses. a. Schematic illustration of Spike protein expression on the cell surface and soluble ACE2-Fc staining followed by an anti-Fc antibody staining. b. 293 cells transfected with Spike protein with or without endoplasmic reticulum retention signal (ERRS) and VSV-G as a negative control. The cells were stained with ACE2-Fc and anti-Fc-APC secondary antibody, flow cytometry data overlays are shown. c. Schematic representation of the generation of Spike protein pseudovirus and the interaction with ACE2-expressing host cells. A lentivector plasmid and a Spike protein over-expressing envelope plasmid are used to co-transfect 293 cells to generate Spike pseudovirus that in turn infect engineered cells over-expressing wild type ACE2 or ACE2 fused to mKO2. d. Infection of wild type 293 cells with either bald lentiviruses generated without envelope plasmid or Spike protein pseudovirus. e. Infection of 293-ACE2 cells with bald and Spike lentiviruses. GFP and mKO2 markers are used to determine ACE2 over-expressing cells in ACE2- IRES-GFP and ACE2-mKO2, respectively. f. The titration of CoV-2 and SARS Spike protein pseudoviruses encoding RFP. ACE2-IRES-GFP expressing 293 cells were incubated with 3-fold serial dilutions of virus supernatant and analyzed for RFP expression by flow cytometry on day 3 post-infection. Percent infection is % RFP+ cells after gating on GFP+ cells (i.e. ACE2+).
The antibody response might be relatively short-lived
The researchers also found that the antibody titers were lower among donors that were further out from the diagnosis of the infection, indicating that the antibody response to COVID-19 might be relatively short-lived.
“Thus, understanding the mechanism of survival from COVID-19 and immune response dynamics will be critical in better prediction of outcomes as well as assaying for a protective response to potential vaccines,” they write.
The team says the high sensitivity and specificity of their high-throughput immunoassay has enabled them to correlate the Spike protein RBD-specific antibody levels with neutralization titers (which showed very high concordance) and “thus can be utilized as a proxy for neutralization in a clinical setting.”
“The assays developed herein can have a utility in uncovering dynamic changes in the antibody levels in SARS-CoV-2 infected subjects over time, in responses to vaccines, and as potential clinical determinants for plasma or antibody therapies for COVID-19 patients,” conclude Unutmazy and colleagues.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources