Researchers from the National Institutes of Health (NIH) and Cornell University in the U.S. just described several new inhibitors of human coronavirus spike proteins by screening a library of approved drugs with SARS-S and MERS-S pseudotyped particle entry assays. Their exciting findings are currently available on the bioRxiv* preprint server.
In the months since the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) rose from a regional outbreak to a pandemic of coronavirus disease (COVID-19), small and large drug manufacturers have scrambled to find a solution to halt this global health emergency.
It is well known that all viruses rely on host cells for replication and propagation; therefore, cell entry represents the initial step of any viral life cycle. However, that also makes it a prime target for drug intervention, and both pathogen-specific and broad-spectrum inhibitors of viral entry have been identified for emerging viruses.
Consequently, a wide array of drug repurposing and virtual screens have been described for SARS-CoV-2. Still, it is commonly observed that the potencies determined in drug repurposing research are not high enough to achieve clinical relevance in comparison to the human plasma concentrations at approved and accepted dosing regimens.
All of this means we need to up the ante. Developing BSL-2 compatible SARS-CoV-2 compound screening assays can be considered as an alternative and more simplistic approach for high-throughput screening and subsequent drug development. Viral entry assays that utilize pseudotyped particles (P.P.) are cell-based BSL-2 viral assays that are ideal for this purpose.
This was recently tried by the researchers from the NIH and Cornell when they decided to apply phenotypic SARS-S and MERS-S PP entry assays for drug repurposing screens in order to identify viral entry inhibitors with diverse mechanisms of action.
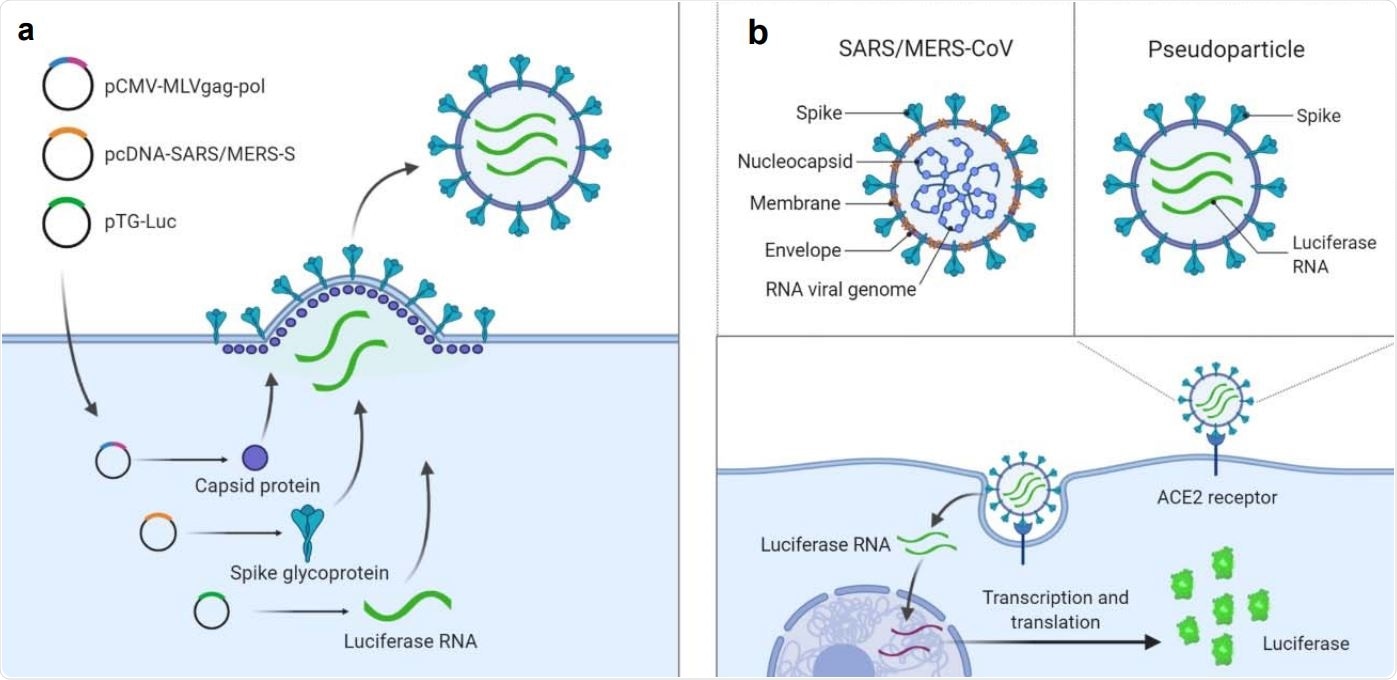
Illustration of pseudotyped particle generation and entry assay. (a) Three plasmids (pCMV-MLVgag-pol, pcDNA-SARS-S/MERS-S, and pTG-Luc) are co-transfected into HEK5 293T/17 cells. The plasmids express MLV core gag-pol polyprotein, coronavirus spike glycoproteins, and luciferase RNAs, which together assemble into pseudotyped particles. (b) Comparison of SARS/MERS-CoV and pseudotyped particle, showing shared spike proteins to facilitate entry into the target cell. Once cell entry occurs, RNAs of pseudotyped particles are released into cell, where they are reverse transcribed into DNAs, integrated into the genome, and express luciferase reporter enzyme. Illustrations were made with BioRender.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
A comprehensive drug repurposing screening endeavor
Their study reports parallel drug repurposing screens to identify a set of broad-spectrum coronavirus entry inhibitors. SARS-CoV-2 live virus cytopathic effect assay has been employed to confirm the inhibition of SARS-CoV-2 entry.
To achieve adequate optimization and miniaturization, both SARS-S and MERS-S PP were produced by a co-transfection with three plasmids to yield particles with spike protein (SARS-S or MERS-S), capsid protein of murine leukemia virus, and luciferase ribonucleic acid (RNA).
The NCATS pharmaceutical collection (NPC) of 2,678 compounds, which is essentially a storehouse of either approved or investigative drugs, was utilized for drug repurposing screening of both SARS-S and MERS-S PP entry assays. Those that were considered as 'hits' were included in further analysis.
"In our secondary assays, we retested the 106 cherry-picked hits in the same SARS-S and MERS-S PP entry assays, along with ATP content cytotoxicity assays at 11 concentrations with 1:3 titration", study authors further explain their stringent methodological approach.
Several cell cultures were used to appraise cytopathic effects – Vero E6 (kidney cells derived from the African green monkey), Huh7 (specific type of human liver cells), and Calu-3 (human lung cancer cell line).
Three 'musketeers' that inhibit SARS-CoV-2 cell entry
"In this study, we identified seven coronavirus spike-driven entry inhibitors out of a library of 2,678 approved drugs", say study authors. "We identified three compounds (cepharanthine, abemaciclib, and trimipramine) that rescued the cytopathic effect of SARS-CoV-2 infection", they add.
Albeit the exact mechanism for such entry, inhibition is still unclear, these three compounds inhibited coronavirus cell entry with greater potency when compared to the control cell entry with vesicular stomatitis virus G glycoprotein, highlighting, in turn, corona-specific inhibitory effect.
Of these three, cepharanthine (already known for its broad antiviral activity) and abemaciclib (previously tested for the treatment of breast cancer) have been reported to rescue cytopathic effect of SARS-CoV-2 in Vero E6 cells, and thus are considered more promising candidates than trimipramine (which is otherwise an oral antidepressant) who did not display that effect.
A combination therapy as an optimal approach
In a nutshell, by employing a very specific high-throughput screening approach, this research group identified three compounds that act as broad-spectrum inhibitors for spike-mediated viral entry. It is not often the case that one study reveals more than one promising drug candidate.
"This work should contribute to the development of effective treatments against the initial stage of viral infection, thus reducing viral burden in COVID-19 patients", study authors accentuate the significance of their findings for the ongoing pandemic.
Still, the question remains what will be an optimal approach in the long runt. Current experience shows that these coronavirus-specific viral entry inhibitors may be used in combination therapy with other anti-SARS-CoV-2 drugs that exhibit different modes of action – such as already approved remdesivir or various promising autophagy inhibitors.
Consequently, the synergistic effect of combination therapy with two or three different drugs may indeed augment the therapeutic effect with lower doses of individual drugs, reducing, in turn, potential adverse effects. As with many other viruses, this could be a way forward with SARS-CoV-2; hence, new studies are anticipated with excitement.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources