The team’s SARS-BLOCK™ peptides competitively inhibit the receptor-binding domain (RBD) on the SARS-CoV-2 spike protein – the surface structure that binds the host cell angiotensin-converting enzyme 2 (ACE2) receptor, to gain viral entry.
As well as blocking this viral entry pathway, the peptides are also designed to bind to the neutralizing antibodies generated against SARS-CoV-2 following infection, with the aim of inducing a specific neutralizing antibody response.
“In contrast to neutralizing antibody therapies and other approaches that seek to target the virus, we developed a biomimetic virus decoy peptide technology that would compete for viral binding to cells, and expose the virus for binding to neutralizing antibodies,” write Dr. Andre Watson and colleagues.
The researchers say they also discovered that ACE2 prevents the binding of RBD to neutralizing antibodies and that the SARS-BLOCK™ peptides can reverse this immune-evasive mechanism of the virus.
A pre-print version of the paper is available in the server bioRxiv*, while the article undergoes peer review.
The challenges faced in developing treatments
Since the COVID-19 outbreak began in Wuhan, China, late last year, SARS-CoV-2 has spread at an unprecedented rate and now infected approximately 19.73 million people and caused more than 728,000 deaths globally.
The long-term health effects of SARS-CoV-2 infection are not yet known but are thought to involve outcomes ranging from respiratory, neurological, and vascular to renal, respiratory, and even autoimmune.
Although various vaccines are in different stages of development, the long-term immune responses to vaccination and infection remain unclear, and the alternative “herd immunity” approach would significantly compromise global health and pose a significant burden in terms of the long-term associated health problems.
One potential treatment approach is the administration of neutralizing antibodies as a way of halting disease progression, but the transient nature of this therapy leaves the patient vulnerable to reinfection.
“An ideal therapeutic strategy must take into account the root cause of infection, rather than attempting to treat specific downstream symptoms, given the measurable effects of various portions of the virus on extensive and specific cascades of physiological, intracellular, and biological systems,” said Watson and team.
Mimicking the RBD of the viral spike protein
The researchers used computational techniques to design, simulate, and synthesize model peptides that mimic the SARS-CoV-2 spike RBD for ACE2 and exhibit neutralizing antibody binding motifs.
Next, the team conducted binding and competitive association assays and infected ACE2-expressing cells with pseudotyped lentivirions displaying the SARS-CoV-2 spike protein.
After modeling the key interacting motifs of the spike protein and its RBD, the team was able to design and synthesize SARS-BLOCK™ peptides that potently and competitively inhibited the binding of ACE2 to the RBD, which reduced cellular infection by as much as 95 to 100%, without causing any toxicity.
As well as preventing this RBD-ACE2 binding, the peptides bound neutralizing antibodies with micromolar affinity, says the team.
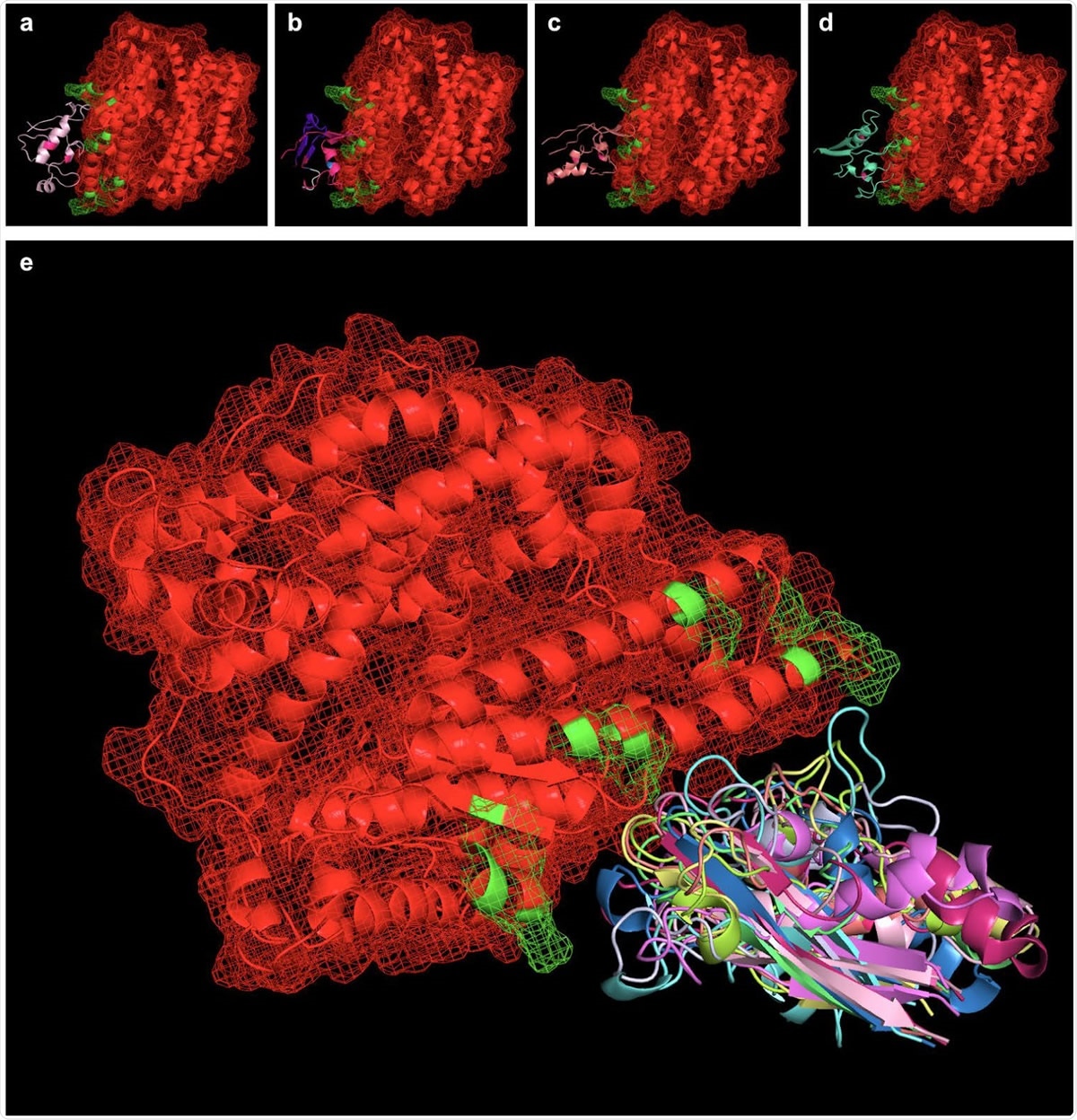
Peptides simulated via RaptorX were aligned with the SARS-CoV-2 binding interface of ACE2 (ACE2 in red, with PDBePISA-predicted binding interfaces in green). Shown from left to right (top) are SARS-BLOCK™ Peptides 1 (a), 4 (b), 5 (c), and 6 (d) bound to ACE2. Of note, all peptides exhibited two mutations introducing a disulfide bond to recreate the beta sheet structure of the SARS-CoV-2 receptor binding motif (RBM). Otherwise, Peptides 1 and 4 utilized the wildtype sequence, while Peptide 5 utilized MHC-I and MHC-II epitopes, and Peptide 6 utilized a GSGSG linker (white) in one of its non-ACE2-interfacing loop regions. Peptides 4, 5 and 6 exhibited additional, proprietary modifications to their sequences to facilitate appropriate folding, while Peptide 1 lacked this modification. Taking into account the 9 possible folded states generated for each peptide, we utilized PyMOL align commands which take into account multiple potential conformations of each peptide and may serve as a basis for future studies exploring more advanced molecular dynamics approaches for relaxing and simulating intramolecular interactions at the binding interface (e). In essence, the overlay of many possible folded states represents an electron distribution cloud of possible states that can be simulated for the minimal interfacial free energy of binding, and this approach requires vastly fewer computational resources than typically required for modeling binding pockets of de novo peptides or protein-protein interfaces without existing interfacial structures.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
ACE2 stopped spike being recognized by the immune system
The researchers also discovered that soluble ACE2 blocks the binding of neutralizing antibodies by forming an “immune cloak” on the spike RBD, thereby preventing the RBD from being recognized by the adaptive immune system.
The researchers found that the spike RBD bound to ACE2 with an approximate 2.3nM affinity and that ACE2 could stop a neutralizing antibody binding with the RBD that would otherwise have a bound with an affinity of around 6nM.
Next, the team showed that the SARS-BLOCK™ peptides were able to “uncloak” the ACE2 viral coating mechanism, thereby reversing this immune-evasive property and stimulating an immune response by enabling the binding of neutralizing antibodies to the viral surface.
“In contrast to neutralizing antibody therapies and other approaches that seek to target the virus, we developed a biomimetic virus decoy peptide technology that would compete for viral binding to cells, and expose the virus for binding to neutralizing antibodies,” writes the team.
The peptides could provide a new generation of antivirals that can be quickly produced
The researchers say the synthetic nature, in silico screening, and precise conformation of the peptides enable their rapid production, without the limitations that are associated with other antiviral approaches or vaccine approaches.
The team says the SARS-BLOCK™ peptides are a promising COVID-19 antidote that effectively provides a new generation of prophylactic and reactive antiviral therapeutics.
“Because SARS-BLOCK™ mimics the virus, rather than binding to it, and also due its ability to displace ACE2 from cloaking the virus spike protein, we believe that these multifunctional peptides will prove to be an effective immune-enhancing strategy in infected patients, with additional potential to serve as a prophylactic vaccine,” concludes the team.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.