The current pandemic of COVID-19 is due to the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) which enters the human host cell via the receptor molecule called the angiotensin-converting enzyme 2 (ACE2). A new study published on the preprint server bioRxiv* in August 2020 reports a novel engineered molecule based on human and other mammalian orthologues of ACE2 fused with the Fc part of the antibody. This molecule interacts more tightly with the viral receptor-binding domain (RBD) to constitute a powerful immunoadhesin that targets the virus for destruction.
The significant impact of the non-pharmaceutical interventions implemented to stop the virus from spreading in an unbridled fashion has caused both economic and social disruption globally. Thus, scientists have been on the job to identify effective antivirals and vaccines that can cut short the deadly course of the pandemic.
Immunotherapy Targeting the Virus
One promising avenue is immunotherapy, where viruses can be directly targeted by the therapeutic molecule, as well as by the activated immune effector cells that destroy the infected cells and thus prevent successful propagation of the virus. This approach is already in use in other viral conditions such as the monoclonal antibody developed against the respiratory syncytial virus (RSV), and broadly-neutralizing antibodies against HIV. Convalescent serum from patients who have recovered from COVID-19 is currently being used to treat other patients to overcome the infection.
Immunoadhesins
The current study focused on developing a targeted immunotherapeutic molecule. The researchers looked at immunoadhesins, which are molecules that look very much like antibodies but have a sticky patch moored to the specific antigen detecting Fc part of an antibody. They aimed at using the human ACE2 receptor as their sticky patch, or binding domain. However, prior research shows that the virus, being primarily an animal virus, may show a higher binding affinity for its orthologous cell receptors in other species than in humans. They, therefore, set out to build an immunoadhesin specific for the SARS-CoV-2, with a higher binding affinity for the virus than the human ACE2 receptor shows.
The researchers say, “Such an engineered ACE2 may be better suited for therapeutic applications than the human ACE2.” The current team of researchers has already provided proof-of-concept by constructing Arenacept which is a very potent arenavirus-targeting immunoadhesin.
Improving ACE2-RBD Affinity
The ACE2 binds to the RBD of the virus, recognition being accomplished mostly through a long N-terminal helix. With over 200 mammalian ACE2 receptors being compared, it is clear that many of the residues at the recognition site are not conserved, leaving ample space for modification to achieve greater binding affinity.
To find the best alterations concerning this objective, the researchers selected 70 ACE2 genes from orthologous species, all with over 80% sequence identity. Using modeling techniques, they evaluated the stability, the binding energy, the interface packing, and the complementarity of shape for the RBD. The top 20 models brought up with this approach were then assessed visually to find those mutations that seemed most likely to achieve closer contact with the RBD than the human ACE2 could achieve.
They found numerous possible sequences but rejected tryptophan mutations since these were likely to show nonspecific interactions. They also discarded those mutations that were already shown to impair RBD binding. Finally, they decided on three mutations at the first N’-terminal helix, which enhances packing with the hydrophobic residues of the viral RBD and stabilizes the receptor-RBD interaction. Another three mutations nearby favored receptor-RBD binding and packing. With these six changes, the energy of binding improved significantly.
A few other mutations removed one glycosylation site on human ACE2 that might cause steric hindrance for RBD binding and disabled the enzymatic activity of the molecule to enable its safe use in humans as a therapeutic.
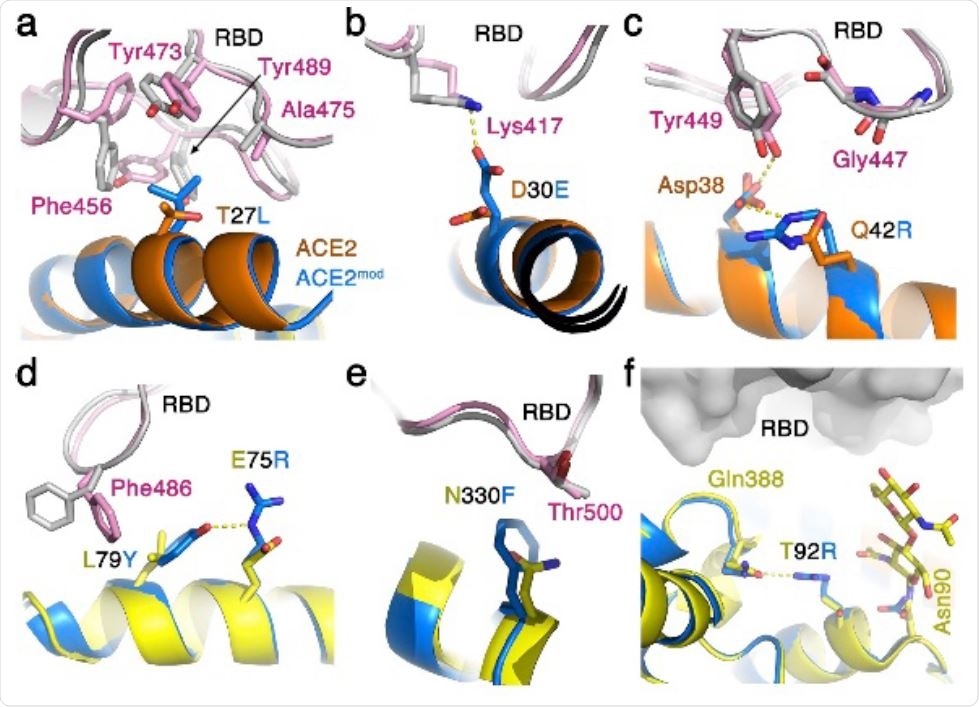
Optimized ACE2 interface for improved binding of SARS-CoV-2. The interfaces of SARS-CoV-2 RBD/human-ACE2 (grey and orange/yellow, respectively) and of the SARS-CoV-2 RBD/modified-ACE2 (pink and light blue, respectively) are shown. a. Leucine, instead of threonine in position 27, makes better Van der Waals interactions with hydrophobic residues on the SARSCoV- 2 RBD. b. Glutamic acid in position 30 of ACE2 can make a salt bridge with Lys417 of SARSCoV- 2, but not an aspartic acid that is present in the human-ACE2. c. Arginine in position 42 can form a salt-bridge with Asp38 of ACE2 to stabilize it in a configuration that allows it to make a hydrogen bond with the hydroxyl of Tyr449 from SARS-CoV-2 RBD. An arginine in this position can also assume a different rotamer that will allow it to form electrostatic interaction with the main-chain carbonyl oxygen of Gly447 of SARS-CoV-2 RBD. d. A double replacement of Leu79 and Glu75 with tyrosine and arginine respectively allows favorable interaction between Phe486 of SARS-CoV-2 RBD and Tyr79 that is stabilized through a hydrogen bond by Arg75. e. Phenylalanine in position 330 of ACE2 is predicted to pack better against the aliphatic portion of Thr500 from SARS-CoV-2 RBD. f. A replacement of Thr92 with arginine abrogates the glycosylation site on Asn90, which bears a glycan that can sterically interfere with the binding of SARS-CoV-2 RBD. An arginine in position 92 of ACE2 can form a hydrogen bond with the nearby Gln388.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Constructing the Immunoadhesin
They then used this engineered protein to produce a chimeric protein comprising part of the human ACE2 ectodomain with the Fc part of the IgG1, both with and without the eight mutations mentioned above. They found that the modified construct had no catalytic activity, and had good binding to the RBD. The binding affinity, in fact, was two orders of magnitude higher than that of the unmodified ACE2 construct, mainly because of a drop in the dissociation rate by two orders.
The next step was to assess the effect of this increased affinity in biological terms. They carried out a pseudovirus neutralization assay. They found that the modified ACE2 construct had dramatically better neutralization potency than the other, with both the half-maximal and the 80 percent neutralization capability (IC50 and IC80 respectively) being ten times better with the former.
Implications
The findings show that it is possible to overcome the handicap posed by the limited potency of immunoadhesins based on ACE2, as well as the fact that the natural ACE2 receptor stoichiometrically inhibits them on human cells. Instead, a more efficient binding molecule can be used to block the viral RBD and thus prevent infection of the human cell. This is the mechanism of some of the monoclonal antibodies that act against the virus.
A benefit of this engineered immunoadhesin over such antibodies is that. In contrast, the latter have binding sites that overlap the RBD only in part, the modified protein binds only with those residues that constitute the viral binding site for ACE2. The problem with the former antibodies is that leaving some of the RBD open allows for escape mutations, whereby the virus can still bind to the ACE2 and infect the cell using other recognition sequences.
The immunoadhesin is thus a highly promising molecule in terms of better binding affinity. The modified construct also has a higher capacity to recognize the spike protein of the virus. In addition, they say, “Anti SARS-CoV-2 immunoadhesin that binds to cell-surface displayed spike complexes might recruit beneficial immune factions via its Fc portion.”
The researchers sum up, “Altogether, we present a promising immunoadhesin that we now term Coronacept, which is a candidate immunotherapeutic agent for COVID-19.”

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources