Within a year of the start of the COVID-19 pandemic, scientists have discovered that it is far more complex than expected. With over 31 million cases and almost a million fatalities, the achievement of herd immunity is still far away. Meanwhile, hundreds of research teams are working to find a vaccine as fast as possible, focused chiefly on the viral spike protein using many different strategies.
A new study by researchers at Clover Biopharmaceuticals and Tsinghua Institute of Multidisciplinary Biomedical Research, Tsinghua University and published in the preprint server bioRxiv* in September 2020 reports on the structure of this protein, as revealed by cryo-electron microscopy (cryoEM).
Vaccine Criteria
For any vaccine to be successful, four criteria must be met. Namely, it must be safe, effective, scalable, and must be developed fast. Protein subunit vaccines do meet the first three, as demonstrated by the human papillomavirus (HPV) vaccine Gardasil and the herpes zoster vaccine Shingrix. Still, they do take a very long time to be produced, running into decades.
One big reason is the difficulty of ensuring that the protein subunit antigen retains its structure very close to the native form throughout the manufacturing process. In the case of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the spike protein is a trimeric surface antigen that is essential for viral entry into the host cell. The challenge here is to stabilize this in trimeric conformation while also introducing a mutation in the viral antigen so that it ‘freezes’ in the prefusion state. The first is by using a non-covalent trimer-foldon from bacterial phage GCN4 protein. The second is usually by preventing spike cleavage through a mutation at the furin cleavage site.
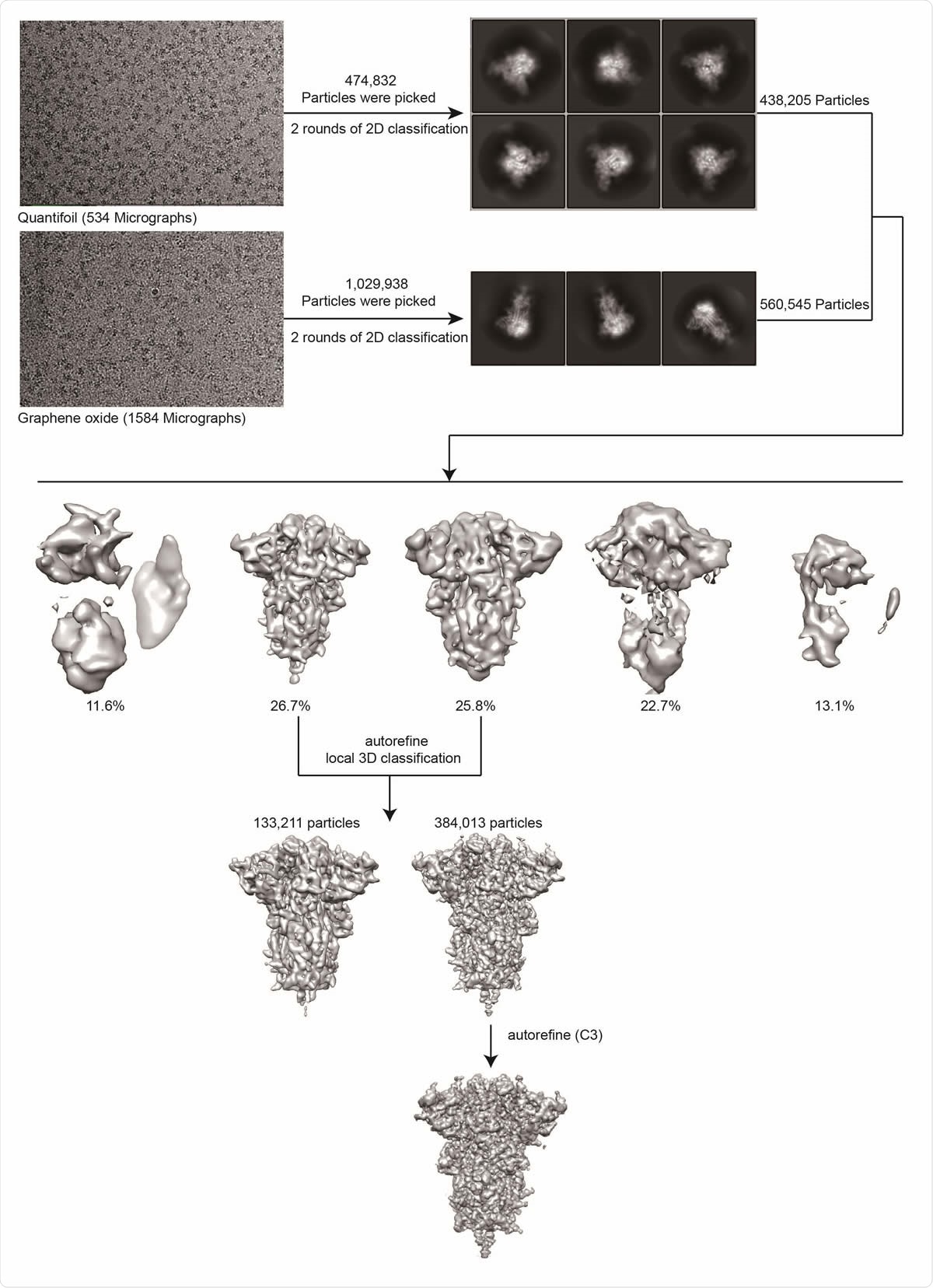
Cryo-EM data processing workflow for the MT S-Trimer.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Trimer-Tag Technology
Though these techniques have been used over decades of research on HIV and RSV vaccines, it is noteworthy that both have not yet yielded a single successful vaccine. The current research, therefore, used another route, namely, Trimer-Tag, to produce both soluble wild-type (WT) and a furin site mutant (MT) forms of S-Trimer in a bioreactor with large yields of 0.5-1 g/L.
This exceeds previously reported antigen yields by three orders of magnitude concerning foldon derived S proteins with a double-Pro mutation, thus proving its scalability.
The CryoEM structure of the S-trimer shows an ectodomain from residue 1-1211, fused to the C-terminal region of human type I collagen to form a homotrimer via disulfide bonds. The researchers used a high-affinity collagen receptor fusion protein to bind to the Trimer-Tag in an affinity purification set up so that the secreted S-trimers were almost homogeneously purified in one step.
Higher Infectivity due to Furin Cleavage
Analysis of this protein reveals a metastable WT S protein in trimeric form, partially cleaved between S1 and S2. When the single point mutation was introduced in the MT S trimer, the protein remained intact. Interestingly, the WT SARS-CoV S protein is also uncleaved by furin protease.
All three spike variants discussed so far have a similar high affinity to ACE2-Fc at nanomolar concentration. The researchers comment that this shows that “the much higher infectivity of SARS-CoV-2 is more likely attributed to furin cleavage of the spike protein that is largely absent in that of SARS-CoV-1,” rather than being due to the difference in their respective affinities to the receptor.
Tightly Closed Structure
Both the WT and MT S trimers are revealed to be homogeneous particles under negative staining electron microscopy. The internal structure was clear, and they observed that the Trimer-Tag was not seen in the molecular structure because of the linker’s flexibility. The linker is the sequence that joins the soluble S to the C-prodomain of the collagen molecule.
They also found that the RBD domains in all three protomers were in the down conformation, unlike that of the previously observed S protein with a double-Pro mutation. On the alignment of the S2 domains, they found that all three S1 domains shifted towards the threefold axis relative to their position in the S-2P structure. This led to a tightly closed structure.
PS80/Fatty Acid Interactions Stabilize Closed State
On the other hand, there was unexpected density in both the N-terminal domain and the RBD of the S1 domain. The former was likely due to the presence of the detergent PS80, buried deep in the hydrophobic pocket residues while some hydrophilic residues of the spike protein form hydrogen bonds with the PS80 hydroxyl group.
The elongated nature of the EM density in the RBD suggested it may be due to oleic or linoleic acid in the culture medium. The high-quality EM mapping feature allows both of these molecules (P80 and the oleic or linoleic acid) to be fitted well into the density of these domains, and their identity was confirmed by mass spectrometry analysis. Thus, PS80 probably lends stability and order to the NTD disordered loops.
The oleic acid in the hydrophobic pocket of the RBD is bound via a salt bridge to the adjacent protomer, so that both RBD domains come in close proximity to each other, accounting for the closed conformation.
Acidic Conditions Favor Closed State
The S trimer is stable under acidic conditions, as confirmed by a more homogeneous appearance of these particles at a pH of 5.5 compared to the physiological pH. However, at this pH, the NTD was not as well seen in the MT S as in the WT S trimer.
The researchers observed a switch element, which they called pH switch 2, in the C-terminal domain-1. In the MT structure, this underwent pH-dependent structural rearrangement. At an acidic pH, it forms a helical repeating structure, whereas it was disordered at physiological pH.
It is thought that this is because protonation occurs at lower pH. This results in hydrophobic interactions instead of the earlier disorderly arrangement, with an ordered repeating arrangement.
This structural motif comes in contact with the pH switch 1 of the neighboring protomer and stabilizes the trimer in a tightly closed state, where all three RBDs are also in close proximity to each other. This observation is supported by earlier studies and indicates that the change in conformation of the MT vs. WT trimeric S protein is triggered by the change in pH rather than the furin cleavage mutation.
D614G Favors Open RBD Conformation
Overall, therefore, the researchers found three states of the S protein. This includes a tightly closed state, a loosely closed state, and an open state, with the switch from the first to the third state via the conformational or pH switch, also called the fusion-peptide proximal region (FPPR), which is highly ordered.
It has been observed that the D614G mutation becomes dominant everywhere it emerges due to its increased infectivity. The D614 is responsible for salt bridge formation involving a residue in the tightly closed RBD conformation at the switch region. For this to open, it has to undergo a marked change to a disorderly arrangement.
When the D614G mutation occurs, this salt bridge is no longer formed. The change causes a flipping of the residue that formerly interacted with the salt bridge so that in its new position, it interacts with other residues on the CTD1. The result is a downward shift of the S1 relative to the S2. The final downward movement of the CTD1 domain results in the open conformation of the RBD that allows it to bind to the host cell ACE2 receptor.
Implications
The researchers in this study used Trimer-Tag to generate both the wild-type (WT) and the furin site mutant (MT) trimeric spike protein variants. They examined both protein structures with cryoEM. They found that both are in a tightly closed conformation, but structurally they are almost identical to the WT full-length spike protein when visualized in a detergent-containing solution.
The researchers point out that this is the first time a cryo-EM structure has been described for the WT S trimer protein in soluble metastable form. The structural integrity of the described protein allows for vaccine candidates to be developed based on the WT spike trimer protein rather than, as formerly, the furin cleavage mutation variant or the S-2P variant.
This should be associated with broader protection and greater vaccine efficacy. The researchers say, “Trimer-Tag technology that has been proven here to be able to rapidly produce large quantities of native-like S-Trimer antigen, may offer a platform technology for subunit vaccine development for enveloped RNA viruses that use ubiquitous trimeric antigens to invade host cells.”
The researchers say they are not sure if the furin cleavage site in this structure is cleaved or intact, not could they rule out other conformational states in the WT sample. Despite this, they say the purified WT S trimer is mostly in a prefusion state, whereas the full-length WT spike protein is found to adopt both pre- and post-fusion states in the presence of detergent.
Preclinical studies show this vaccine candidate is able to rapidly induce high levels of neutralizing antibodies, with a beneficial Th1-skewed cellular immune response in rodents and non-human primates. The latter was also completely protected from SARS-CoV-2 infection after immunization. Such a subunit vaccine is now being tested in clinical trials.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Ma, J. et al. (2020). Cryo-EM structure of S-Trimer, A Subunit Vaccine Candidate For COVID-19. bioRxiv preprint. doi: https://doi.org/10.1101/2020.09.21.306357. https://www.biorxiv.org/content/10.1101/2020.09.21.306357v1
- Peer reviewed and published scientific report.
Ma, Jiahao, Danmei Su, Yinyan Sun, Xueqin Huang, Ying Liang, Linqiang Fang, Yan Ma, Wenhui Li, Peng Liang, and Sanduo Zheng. 2021. “Cryo-Electron Microscopy Structure of S-Trimer, a Subunit Vaccine Candidate for COVID-19.” Edited by Tom Gallagher. Journal of Virology 95 (11). https://doi.org/10.1128/jvi.00194-21. https://journals.asm.org/doi/10.1128/JVI.00194-21