As the coronavirus disease 2019 (COVID-19) pandemic continues to cause many difficulties for public health authorities, research is ongoing to understand how the virus can cause fatal effects on those infected and thereby suggest effective preventive and therapeutic measures against it.
A new study conducted by researchers at the Houston Methodist Hospital, Rutgers University, and other institutions has been published on the preprint server medRxiv.* Its findings show the central role that innate immunity can play in controlling the viral load of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative pathogen of COVID-19. In doing so, the researchers have underscored the importance of timing in the therapeutic treatment of COVID-19 in determining the outcome.

Modeling Virus-Receptor Interactions

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Mathematical modeling of the SARS-CoV-2 infection has shown its importance in arriving at a better understanding of the viral dynamics and the disease's pathophysiology. Such models are based on the fact that the angiotensin-converting enzyme 2 (ACE2) receptor is required for SARS-CoV-2 infection of the host cell and is shaped by current knowledge of immune responses. The aim is to predict the viral load in different organs of the body.
ACE2 messenger RNA (mRNA) is found in very high levels on the alveolar epithelium surface of the lungs and small intestine enterocytes. With an incubation period of 3 to 17 days, a significant percentage of asymptomatic infection, and a large proportion of patients with only mild nonspecific infection, it is challenging to diagnose COVID-19 based on symptoms.
Therefore, such a diagnosis must be supported by the detection of viral RNA by RT-PCR testing (specimens from both upper and lower respiratory tracts) and pulmonary CT scans. The significant biomarker of COVID-19 is the viral load in nasal and oral swabs, and therapeutic interventions aim to reduce it to undetectable levels.
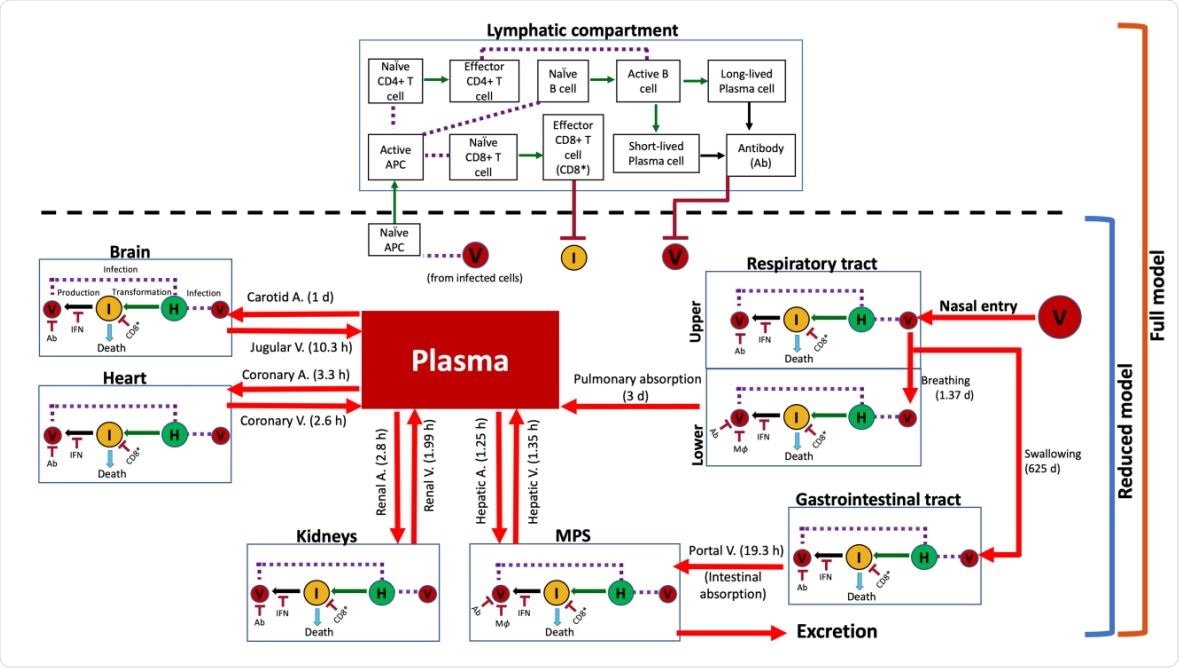
Model schematic showing system interactions. Connectivity diagram between compartments indicating viral transport mechanisms, cell populations, immune system agents, and their interactions. Estimated characteristic times of the various transport processes are given in parentheses alongside red arrows. Notation: (V) virus, (H) healthy cells, (I) infected cells, (IFN) interferon, (Ab) antibody, (CD8* ) effector CD8+ cells, and (APC) antigen presenting cells. Solid red arrows indicate transport of virus; solid green arrows indicate transformation of a cell into another type; solid black arrows indicate production of an agent; purple dashed lines indicate interaction between two agents; and solid dark red arrows with a flat head indicate inhibition.
Utility of Mathematical Modeling in COVID-19
With limited data on the viral dynamics in living organisms, mathematical modeling at various scales has been widely adopted to fill out the current knowledge of virus-host interactions at molecular, cellular, tissue, organ, and organism levels.
Earlier models have provided useful information – such as showing that antivirals are most effective when used early in the course of the disease – before symptoms arise. Others have modeled the interactions between both the infected and the immune cells as well as the relationship between infectiousness and severity of the disease.
Multiscale Model
The current study uses a "multiscale mechanistic model" designed to include processes from viral infection through viral dissemination and destruction of infected cells to immune responses. One fundamental assumption was that the rate of bile production was the rate-limiting step in the excretion of the virus through the bile duct into the feces.
The unknown parameters were estimated from known parameters, from published in vivo and clinical data. The model was then calibrated against published experimental data. The model assumes that viral entry into various tissues is dependent on the permeability of the tissue rather than the flow of blood to the tissues.
The size of the endothelial pores thus decides how long the virus takes to enter each organ. Thus, it takes 20 times longer to penetrate the brain compared to the mononuclear phagocytic system (MPS) because of the large pores in the capillaries of the latter.
Viral Load Varies with Immune Response
The model predicted the kinetics of the viral load in various body compartments on the basis of the earlier estimated parameters. Thus, the predicted persistence of the virus in organs other than the lungs and in the blood is for around 17 to 20 days after the earliest symptom, which matches published studies and compares well with the known duration of viral load in the upper and lower respiratory tract infections. This indicates that the use of nasal swabs to reflect the infection status of the patient is valid.
Antibody Production Undetectable Beyond 100 Days?
The model also predicted that antibodies in plasma are detectable for 100 or more days, but then sink below the limit of detection. This, they say, "suggests the lack of indefinite antibody protection against reinfection, and highlights the need for vaccine boosters to achieve long-lasting immunity." It is also important to note that the data comes from subjects in the absence of pharmacological treatment. This makes the predictions more reliable concerning viral load distribution and clearance.
Effects of Reduced Immunity on Viral Load
The researchers also looked at the effect of immunity on the viral load. Each component, namely, cellular, humoral, adaptive, and innate immunity, was removed from the model, to model immunocompromised states due to various chronic conditions (cancer, diabetes, autoimmune diseases), in the absence of any antiviral therapy. The viral kinetics were modeled up to the point when the viral load became undetectable.
With the removal of one or more components, the viral concentration in every compartment exceeded the baseline. They also found that interferons were the primary mechanism whereby the virus is suppressed in the upper airway, but macrophages are primarily responsible for viral clearance in the lungs, MPS, and other systemic sites, though interferons do also play a role.
Interferons Promote Viral Clearance
This agrees with earlier findings, such as that interferon deficiency can cause excessive viral load and a potentially lethal case of COVID-19. A subset of macrophages was also absent in severe COVID-19 but present in mild disease, indicating they are primarily responsible for controlling infection, just as in chronic obstructive pulmonary disease. However, the model cannot predict whether or not the disease itself affects macrophage availability and function.
Another finding is that adaptive immunity, both humoral and cellular, is not a major player in limiting infection. However, this does not mean that passive immunity or T-cell therapy is useless. It does highlight the importance of immunity in limiting viral replication.
If immunity is defective, viral clearance depends on the excretion of the virus through the bile duct, which can take weeks to achieve with undetectable viral loads.
How treatment affects viral load
The investigators suggest that their findings indicate the importance of manipulating interferon concentrations and viral replication to limit the viral load following infection. Targeting both parameters is more efficient than monotherapy, which agrees with clinical reports. The model also suggests that viral clearance depends on the rate of cytopathic death of the infected cells, and the rate of viral replication, but not so much on interferon production.
Viral transport to the lungs and between the blood and MPS, are the chief determinants of viral clearance from the plasma and, therefore from other organs. The rate at which lung and MPS macrophages clear the virus also affects the viral load in both the lungs and the plasma, as does the rate at which viral agents travel from the lungs to the plasma.
Simulating antiviral therapy with a remdesivir-like hypothetical agent, or with interferon beta-1a, they found the model predicted significant reduction of the viral load, positively related to the timing of initiation. The combination of both produced the most rapid clearance of the virus, bringing the viral load below the limit of detection 2-3 days earlier compared to monotherapy.
Conclusion
The researchers sum up, "Importantly, using our modeling platform, we can identify treatment strategies for effective viral load suppression under various clinically relevant scenarios. We also expect it to be applicable to other viruses that have shared similarities in mechanisms of infection and physical dimensions." As more data comes in regarding the changes in viral load in organs other than the lung, the model can be fine-tuned to test more therapeutic approaches.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Dogra, P. et al. (2020). Innate Immunity Plays A Key Role in Controlling Viral Load In COVID-19: Mechanistic Insights from A Whole-Body Infection Dynamics Model. medRxiv preprint. doi: https://doi.org/10.1101/2020.10.30.20215335, https://www.medrxiv.org/content/10.1101/2020.10.30.20215335v1
- Peer reviewed and published scientific report.
Dogra, Prashant, Javier Ruiz-Ramírez, Kavya Sinha, Joseph D. Butner, Maria J. Peláez, Manmeet Rawat, Venkata K. Yellepeddi, et al. 2020. “Innate Immunity Plays a Key Role in Controlling Viral Load in COVID-19: Mechanistic Insights from a Whole-Body Infection Dynamics Model.” ACS Pharmacology & Translational Science 4 (1): 248–65. https://doi.org/10.1021/acsptsci.0c00183. https://pubs.acs.org/doi/10.1021/acsptsci.0c00183.