Many pharmacologically important molecules are being identified with potential activity against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative pathogen of coronavirus disease 2019 (COVID-19). Some use immunological mechanisms, while others inhibit crucial enzymes that are required for viral replication. A new study in the preprint server bioRxiv* published in November 2020 describes the discovery of acrylamide fragments that can effectively disable key enzymes in both the hand, foot and mouth virus and SARS-CoV-2 virus.
Enteroviruses are a family of RNA viruses from which many human pathogens have come. This includes the enteroviruses 68 and 71, rhinovirus, coxsackievirus B3 and the poliovirus. Enterovirus 71 (EV71) causes hand, foot and mouth disease, which can cause neurological damage as well as death in severe cases, though these are in the minority. Despite the availability of vaccines against EV71, the disease continues to be endemic in some parts of the world, and no effective therapy has been found.
EV71 is like SARS-CoV-2 in that its genome encodes a large polyprotein from which multiple functional proteins are derived by cleavage. In most enteroviruses, the protease involved in cleavage is the 3C protease (3CPro). Thus, many protease inhibitors have been developed, but none have been approved so far.
SARS-CoV-2 also uses two enzymes, main protease (MPro) and the papain-like protease (PLPro), to cleave its two polyproteins PP1a and PP1ab. MPro shares a common structure with 3CPro at its catalytic site, both being dependent on cysteine at the active site. Both rely on a catalytic triad with a 3C-like fold for their enzymatic activity. Thus, the current study was focused on exploring the possibility of identifying pharmacophores that bind to a range of viral proteases.
At present, there are many therapeutics that operate via a covalent mechanism of action, some of which use a Michael acceptor to bind irreversibly to cysteine on the targeted enzyme. The major issue in many such situations is to find the point of origin for such molecules. The current study offers a novel acrylamide-based MPro inhibitor scaffold for further design.
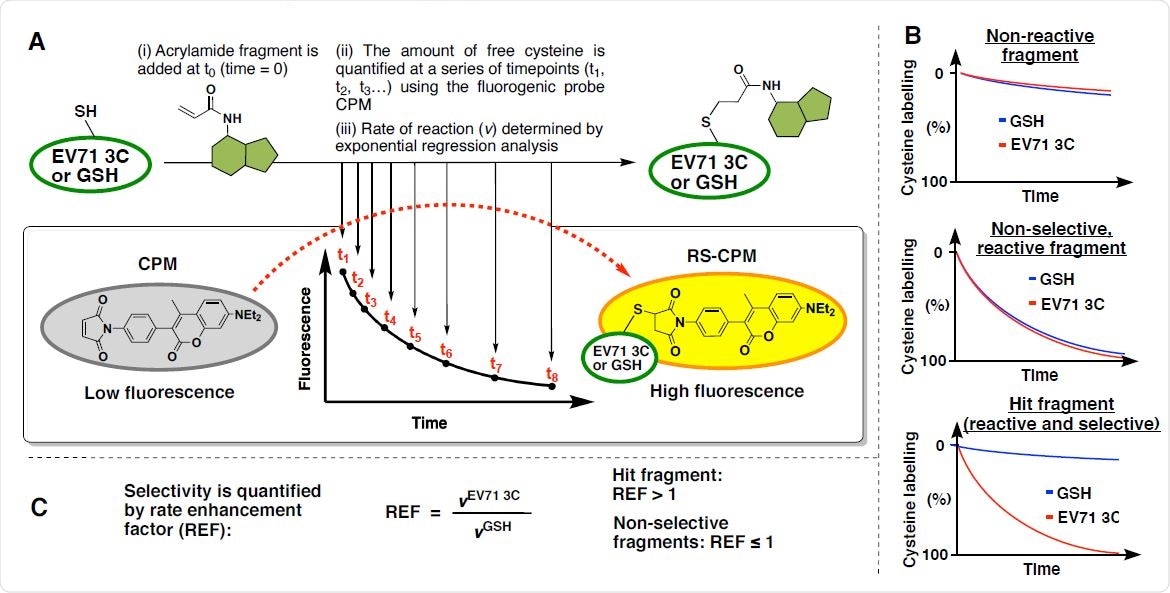
Quantitative Irreversible Tethering (qIT) screening platform. (A) Assay overview: The target thiol (5 μM), EV71 3Cpro or glutathione, is reacted with acrylamide fragments (0.5 mM) under pseudo-first order conditions. Reaction progress is followed by discrete measurements of free target thiol concentration using the fluorogenic probe CPM and the rate of reaction (v) are derived from exponential regression analysis. (B) Fluorescence intensity is converted into percentage cysteine modification by normalizing to DMSO control = 0%, no thiol = 100%. Fragments are characterized as (i) non-reactive, (ii) reactive but non-selective or (iii) reactive and selective by comparing the reactivity profiles between EV71 3Cpro and GSH. (C) Kinetic selectivity is quantified by the rate enhancement factor (REF) which is used to identify and prioritise hit compounds.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Using qIT to identify selective covalent inhibitors
One approach has been to use fragments to initiate such binding, and this shows promise in targeting SARS-CoV-2 MPro. The current study takes this forward using a method newly developed by the researchers, called quantitative irreversible tethering (qIT). Using this, covalent fragments that show selective binding to a given cysteine on the target protein can be identified.
The advantages of qIT include the ability to prioritize hits and to minimize the number of false positives and negatives. This is achieved by normalizing the rate at which proteins are modified by the intrinsic reactivity of the compound.
Using qIT, the researchers examined over a thousand covalent fragments, adding chemical groups that reacted with cysteine, mostly acrylamides, to the core structures to functionalize them. Acrylamides are used in many covalent drugs to disrupt target molecules because they are mild electrophiles without significant non-specific reactivity and, thus, without toxicity.
In this way, they identified inhibitory fragments that formed covalent bonds with cysteine at the active site of the EV71 3CPro. Using the rate enhancement factor (REF), they also measured the selectivity of the fragments for the 3CPro to prioritize the hits.
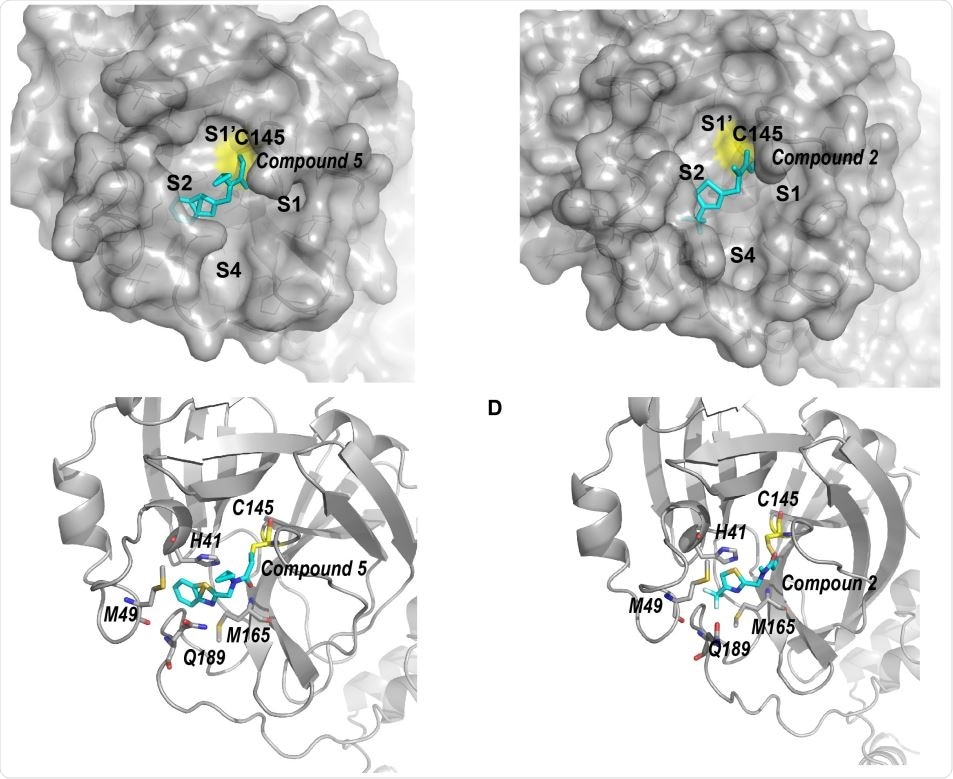
Structure of SARS-CoV-2 Mpro complexed with compound 2 & 5. (A) Surface plot of SARS-CoV-2 Mpro complexed by compound 5 (cyan). Compound 5 occupies pockets S1’ and S2. (B) Ribbon model of SARS-CoV-2 Mpro-5. Residues surrounding the benzothiazole moiety of 5 are shown with stick model. (C) Surface plot of SARS-CoV-2 Mpro complexed with 2 (cyan). The trifluoromethyl thiazole moiety of 2 occupies the S2 pocket. (D) Ribbon model of SARS-CoV-2 Mpro-2. Residues surrounding the trifluoromethyl thiazole moiety are shown with stick model.
Binding induces conformational rearrangements
They examined the structure by co-crystallization and found that these fragments bound to the enzyme via a novel hidden pocket, rather than to the central pocket as expected. The thiazole ring appears to be the main component of this pharmacological binding.
This binding induced a large rearrangement of the conformation of a catalytic loop in 3CPro, containing the critical cysteine residue C147. This loop forms the upper wall of the central pocket. When this loop is bound by a ligand, it is held in the active conformation and allows the formation of an oxyanion hole between C145 and C147. This hole is able to bind a tetrahedral intermediate anion.
However, when this loop is rearranged by acrylamide fragment binding, it resulted in the collapse of the substrate recognition pocket. This led to the disappearance of several other pockets while generating the cryptic pocket. The researchers discovered that some chemical functional groups acted better to produce efficient binding than others.
Successful inhibition of SARS-CoV-2 MPro
The same acrylamide fragments were also found to target the MPro active site cysteine and disable the enzyme, while one fragment also broke up the quaternary protein structure of MPro as well. Thus these fragments inhibited both the enzymes studied, with MPro inhibition being more effective than 3CPro inhibition.
Acrylamide fragment 5 was found to have two distinct mechanisms of inhibition: covalent linkage to the catalytic site and occupation of the substrate-binding pockets, along with features that prevent the formation of the active dimeric MPro. Thus, several hits turned up, some of which markedly rearrange the protease structure.
Implications
Thus, the study demonstrates the use of this screening method in addition to or instead of crystallography, which does not show up the conformational changes, unlike the former. Not only does the use of qIT allow structural rearrangements to be seen, but it provides a preliminary list of hit ligands targeting both EV71 3CPro and SARS-CoV-2 MPro, by covalent binding to the catalytic cysteine residues on both enzymes.
These ligands thus provide “excellent candidates for the development of potent protease inhibitors.” The study also explains how these inhibitors achieve their action and thus promote the design of more selective drugs based on this mechanism.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Qin, B. et al. (2020). Acrylamide Fragment Inhibitors that Induce Unprecedented Conformational Distortions in Enterovirus 71 3C and SARS-CoV-2 Main Protease. bioRxiv preprint. doi: https://doi.org/10.1101/2020.11.06.370916, https://www.biorxiv.org/content/10.1101/2020.11.06.370916v1
- Peer reviewed and published scientific report.
Qin, Bo, Gregory B. Craven, Pengjiao Hou, Julian Chesti, Xinran Lu, Emma S. Child, Rhodri M. L. Morgan, et al. 2022. “Acrylamide Fragment Inhibitors That Induce Unprecedented Conformational Distortions in Enterovirus 71 3C and SARS-CoV-2 Main Protease.” Acta Pharmaceutica Sinica B 12 (10): 3924–33. https://doi.org/10.1016/j.apsb.2022.06.002. https://www.sciencedirect.com/science/article/pii/S2211383522002684.