A recent study from France showed that the UK variant of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) – also known as B.1.1.7 variant – can replicate much more efficiently in a reconstituted human bronchial epithelium, which may explain why it spreads so rapidly in the human population. The study is currently freely available on the bioRxiv* preprint server.
Since its emergence in 2019, circulating populations of SARS-CoV-2 are known to acquire genetic diversity continuously. Consequently, within a few months, D614G spike mutations were pervasive in all viral populations, however, without evidence for higher coronavirus disease 2019 (COVID-19) severity or mortality.
In September 2020, a variant named 20I/501Y.V1 from the lineage B.1.1.7 emerged in the United Kingdom, with a subsequent rapid spread around Western Europe and the United States. Many lines of evidence show that this so-called 'UK variant' can spread much more swiftly and efficiently when compared to pre-existing European strains – particularly in younger individuals.
Furthermore, new studies even show potentially increased mortality due to the UK strain, which in combination with increased spread may give rise to emergent disease waves. Nonetheless, salient biological evidence of different phenotypic characteristics in comparison to original European variants is still lacking.
In this study, a research group from Aix-Marseille University in France (led by Dr. Franck Touret) decided to appraise the replication ability of the UK variant in different cellular models, utilizing an ancestral D614G European strain (lineage B1) for comparison purposes.
Highly specific/realistic cell models
In this study, the replication ability of the 20I/501Y.V1 viral variant (strain UVE/SARS-CoV-2/2021/FR/7b isolated in February 2021 in Marseille, France) was assessed in vitro and ex vivo, by using the lineage B.1 BavPat D614G strain that circulated in Europe in February/March 2020 for comparison purposes.
The initial experiments were conducted in two cell lines frequently used for SARS Cov-2 culture: VeroE6/TMPRSS2 (African green monkey kidney cells) and Caco-2 (i.e., human colorectal adenocarcinoma cells).
After revealing highly similar replication kinetics, the researchers subsequently assessed the replicative fitness of both strains by utilizing a model of reconstituted human airway epithelial cells of bronchial origin.
More specifically, after inoculating the epithelia through their apical side at a multiplicity of infection of 0.1 (which was done in order to imitate the natural infection route of infection), the researchers monitored the excretion of new virions at the apical side 2-4 days after infection and measured the intracellular yield of viral genetic material at day 4.
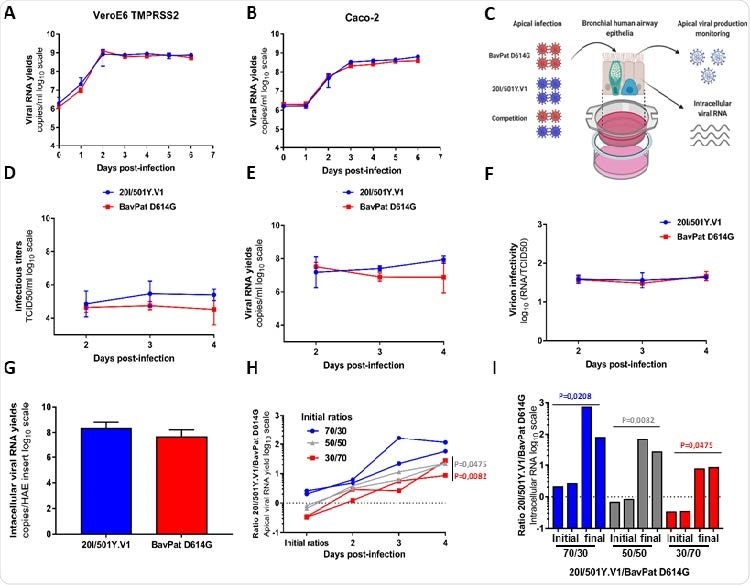
In vitro and ex vivo replication ability of a 20I/501Y.V1 variant in comparison with a lineage B.1 D614G strain. (A-B) Replication kinetics in VeroE6 TMPRSS2 (A) and Caco-2 (B) cells. Viral replication was assessed using a RT-qPCR assay. (C) Graphical representation of experiments with reconstituted human airway epithelium (HAE) of bronchial origin. (D-E) Kinetics of virus excretion at the apical side of the epithelium measured using a TCID50 149 assay (D) and a RT-qPCR assay (E). (F) Estimation of virion infectivities (i.e., the ratio of the number of infectious particles over the number of viral RNA). (G) Intracellular viral RNA yields measured at 4 dpi using a RT-qPCR assay. (A-G) Data represent mean±SD of a triplicate. No statistical difference was observed between both viral strains (p>0.05; unpaired Mann-Whitney test). (H) Follow-up of the 20I/501Y.V1/BavPat D614G ratios at the apical side. Each line represents results from an HAE insert. (I) Individual 20I/501Y.V1/BavPat D614G ratios estimated from intracellular viral RNAs at 4dpi (I). (H-I) p-values were determined against the initial ratios using Kruskal-Wallis test followed by uncorrected Dunn’s post-hoc analysis. The graphical representation was created with BioRENDER.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
New strain outcompeting the old one
The study has shown that infectious titers and yields of viral genetic material at the apical side of the cells were slightly higher for the UK variant at days 3 and 4 of the infection. The same was valid for intracellular yields of viral genetic material on day 4 of the infection.
Nonetheless, these differences were not significant, while estimated relative virion infectivity (which was calculated as the ratio of the number of infectious particles over the number of viral RNA or genetic material) has been similar for both viral strains at all sampling times.
This study's significant highlight was when both viruses were put in competition in a human reconstituted bronchial epithelium because the UK variant subsequently outcompeted the ancestral strain – regardless of their initial ratio used in this study.
Understanding strain replacement
"Our results demonstrated that the 20I/501Y.V1 is more fit than the BavPat D614G in a reconstituted bronchial human epithelium", say study authors in this bioRxiv paper. "This may be explained by the presence of the N501Y mutation in the receptor-binding domain (RBD) of the spike protein, which enhances viral particle binding to the ACE2 receptor", they add.
In any case, this may render certain fitness advantages to the virus, as demonstrated recently in research projects that used engineered viral strains. More specifically, similar findings have been observed with the D614G mutation, where new G614 strains overcame the original D614G strains when put in competition.
Everything said this study could contribute to our improved understanding of the progressive replacement of circulating strains by the SARS-CoV-2 20I/501Y.V1 (or the UK) variant. The emergence of different variants raises numerous questions concerning the future course of this pandemic. Thus more research endeavors in this field are highly needed.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Touret, F. et al. (2021). Replicative fitness SARS-CoV-2 20I/501Y.V1 variant in a human reconstituted bronchial epithelium. bioRxiv. https://doi.org/10.1101/2021.03.22.436427, https://www.biorxiv.org/content/10.1101/2021.03.22.436427v1
- Peer reviewed and published scientific report.
Touret, Franck, Léa Luciani, Cécile Baronti, Maxime Cochin, Jean-Sélim Driouich, Magali Gilles, Laurence Thirion, Antoine Nougairède, and Xavier de Lamballerie. 2021. “Replicative Fitness of a SARS-CoV-2 20I/501Y.V1 Variant from Lineage B.1.1.7 in Human Reconstituted Bronchial Epithelium.” Edited by Peter Palese. MBio 12 (4). https://doi.org/10.1128/mbio.00850-21. https://journals.asm.org/doi/10.1128/mBio.00850-21.