An international team of researchers has warned that individuals with acute malaria infection generate high levels of antibodies that cross-react with the viral spike protein of the novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) that causes coronavirus disease 2019 (COVID-19).
Amy Bei from Yale School of Public Health in New Haven, Connecticut, and colleagues say the routine use of spike-based assays for surveillance of COVID-19 disease risk and burden may overestimate exposure to SARS-CoV-2 and herd immunity in malaria-endemic countries.
“False-positive results could cause an individual to underestimate their risk of future infection,” they write. “At a population level, an overestimate of prior SARS-CoV-2 exposure could lead to overestimates of immunity, meaning larger proportions of people may remain susceptible to infection than serological surveillance might indicate, allowing for the continued spread of SARS-CoV-2.”
A pre-print version of the research paper is available on the medRxiv* server, while the article undergoes peer review.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Serological studies have reported higher seroprevalence than would be expected
Serological surveillance studies aim to provide a fundamental understanding of previous exposure to infectious pathogens such as SARS-CoV-2.
Recently, such studies in Africa have reported higher overall seroprevalence of SARS-CoV-2 than is predicted based on case counts, which has prompted various hypotheses regarding why seropositivity might be higher than expected.
What did the researchers do?
Using an enzyme-linked immunosorbent assay (ELISA), the team tested a total of 741 samples from 617 individuals for SARS-CoV-2 spike protein seropositivity.
Samples were collected across eight countries before the first case of COVID-19 had been reported.
The researchers identified a high degree of cross-reactivity to subunit 1 (S1) of the spike protein among individuals with acute malaria infection. Spike S1 contains the receptor-binding domain (RBD) and the N-terminal domain (NTD) – the two main regions that are targeted by neutralizing antibodies.
Rapid exoproteome antigen profiling of the samples showed that malaria-induced antibodies did not bind to the spike RBD.
Furthermore, linear peptide epitopes tested against bacterial display libraries did not show any significant binding of the antibodies to SARS-CoV-2 peptides.
Taken together, these data imply that the cross-reactivity may be targeted to the NTD of S1 (and not the RBD) or to a post-translational modification such as carbohydrates, says the team.
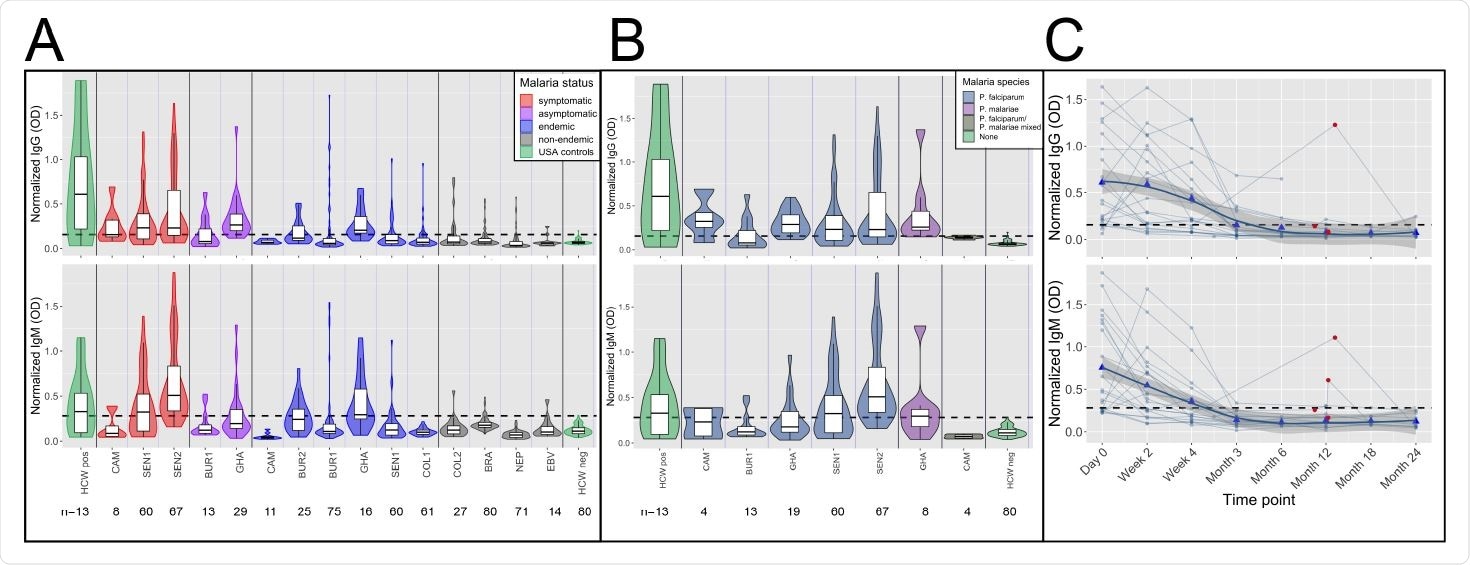
High frequency of cross-reactive antibodies to SARS-CoV-2 Spike protein from Plasmodium-infected individuals. In A. and B., Violin plots showing normalized IgG and IgM responses among patients from different cohorts. A. Among all subjects, those in non-malaria endemic areas had significantly lower IgG and IgM than others (t-test IgG p-value<0.0001 and IgM p-value<0.0001.) Subjects with acute malaria infection (symptomatic and asymptomatic) had significantly higher IgG and IgM than uninfected subjects living in malaria-endemic areas (t-test p-value<0.0001 for both IgG and IgM). B. Normalized IgG was significantly higher among subjects with P. falciparum, P. malariae, and mixed infections than among negative controls (Welch Two Sample t-test p-value<0.0001 for P. falciparum, p-value= 0.044 for P. malariae and p-value=0.008 for mixed infections), and normalized IgM was significantly higher among subjects with P. falciparum and mixed infections but not P. malariae than among negative controls (Welch Two Sample t-test p-value<0.0001 for P. falciparum, p-value= 0.106 for P. malariae, and p-value=0.018 for mixed infections). C. Normalized IgG and IgM over time in 21 subjects with P. falciparum monoinfection on Day 0. Both IgG and IgM peaked between Day 0 and Week 4 for all subjects. Reinfection, shown by red circles, boosted IgG response in 1 of 4 subjects and IgM response in 2 of 4 subjects. Bold trend line based on local regression (LOESS). In A. B. and C., normalized IgG or IgM calculated by IgG or IgM OD divided by IgG or IgM of positive control (camelid monoclonal chimeric nanobody VHH72 antibody was IgG control, and pooled convalescent serum from SARS-CoV-2 patients was IgM control). Black dashed lines represent cutoffs for positivity, calculated from normalized IgG and IgM values from 80 RT-qPCR negative HCWs (mean + 3 SDs).
Testing the specificity of the cross-reactivity
To test the specificity of the cross-reactivity to either structural epitopes such as the NTD or glycosylation, the researchers treated S1 to alter its structure and composition of glycans and carbohydrates.
ELISAs showed that the cross-reactivity of immunoglobulin G antibodies was maximally reversed following treatment with the enzyme neuraminidase, which cleaves terminal sialic acids from glycosylated sites.
Treatment with Peptide:N-glycosidase F – which removes all N-linked glycans – also significantly reduced cross-reactivity.
“Cross-reactive antibodies specifically recognized the sialic acid moiety on N-linked glycans of the spike protein,” write the researchers.
Investigating any protective effects of the cross-reactivity
Next, the team investigated whether these malaria-induced antibodies could neutralize SARS-CoV-2 and potentially protect against infection.
However, in vitro neutralization assays of 11 samples with cross-reactive antibodies found that none demonstrated significant neutralizing activity against SARS-CoV-2 at any dilution tested.
“Cross-reactivity was not neutralizing, giving no evidence that malaria exposure protects against SARS-CoV-2 infection through antibody-mediated viral neutralization,” writes Bei and colleagues.
What are the implications of the study?
“Our study reveals that acute malaria infection can cause cross-reactivity to the S1 Spike protein through antibody binding to terminal sialic acids of complex glycans,” say the researchers.
The team warns that while serological surveys remain critical tools in understanding disease burden, cross-reactivity in malaria-endemic regions, especially in patients with acute malaria, could lead to false positive antibody tests and overestimates population-level exposure.
These overestimates would lead to underestimates of the risk of severe disease among individuals exposed to SARS-CoV-2.
“High rates of false positives could preclude using serological tests as correlates of immunity in malaria-endemic areas, hampering policy decision making with respect to risk stratification and implementation of non-pharmaceutical interventions,” says Bei and colleagues.
“These results further highlight the need to validate diagnostics in populations with different disease exposures and optimize such diagnostics so that serological surveillance tools can accurately track SARS-CoV-2 exposure,” they conclude.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Bei A, et al. Plasmodium infection induces cross-reactive antibodies to carbohydrate epitopes on the SARS-CoV-2 Spike protein. medRxiv, 2021. doi: https://doi.org/10.1101/2021.05.10.21256855, https://www.medrxiv.org/content/10.1101/2021.05.10.21256855v1
- Peer reviewed and published scientific report.
Lapidus, Sarah, Feimei Liu, Arnau Casanovas-Massana, Yile Dai, John D. Huck, Carolina Lucas, Jon Klein, et al. 2022. “Plasmodium Infection Is Associated with Cross-Reactive Antibodies to Carbohydrate Epitopes on the SARS-CoV-2 Spike Protein.” Scientific Reports 12 (1): 22175. https://doi.org/10.1038/s41598-022-26709-7. https://www.nature.com/articles/s41598-022-26709-7.