Researchers in the UK have provided important new insights into the early stages of infection with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) – the agent that causes coronavirus disease 2019 (COVID-19).
The team’s quantitation of SARS-CoV-2 replication dynamics at the single-cell level revealed a previously unrecognized variability among cells in supporting SARS-CoV-2 replication.
The researchers were also surprised to find that the SARS-CoV-2 variant of concern B.1.1.7 (Alpha) that first emerged in the UK replicates more slowly than the Victoria isolate – an early variant that is related to the original strain isolated in Wuhan, China.
Ilan Davis and colleagues say this finding suggests that a novel mechanism contributes to the increased transmissibility of B.1.1.7.
A pre-print version of the research paper is available on the bioRxiv* server, while the article undergoes peer review.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
More about the SARS-CoV-2 genome
The SARS-CoV-2 genome consists of a positive-sense single-strand RNA of 30 kilobases that encodes a plethora of viral proteins.
The virus primarily targets the respiratory tract and infection is mediated by the viral spike protein when it binds to the host cell receptor angiotensin-converting enzyme (ACE2).
Following host cell entry, the first step in the replicative life cycle is the translation of the genomic RNA (gRNA) to form the replicase complex, enabling the synthesis of further positive gRNA copies. In addition, a series of shorter sub-genomic (sg)RNAs is also synthesized that encodes the spike, nucleocapsid and envelope structural proteins, as well as non-structural proteins.
Despite their essential role in establishing productive infection, these early steps in the SARS-CoV-2 replicative life cycle are currently poorly understood.
Typically, analyses of viral replication are performed using “in-bulk” approaches such as reverse transcription-quantitative polymerase chain reaction (RT-qPCR) and conventional RNA sequencing.
“While very informative, these approaches lack spatial information and do not allow single-cell analyses,” says Ilan and colleagues.
“The kinetics of SARS-CoV-2 RNA replication and transcription during the early phase of infection are not well understood and lack quantitative, spatial and temporal information on the genesis of gRNA and sgRNAs,” writes the team.
Concerns surrounding variants of concern
Since the COVID-19 outbreak began in Wuhan, China, in late December 2019, several variants of concern (VOC) have emerged containing mutations that increase transmissibility and confer immune escape.
“Given the current status of the pandemic, there has been a global effort to understand the biology of emergent VOC with high transmission rates and possible resistance to neutralizing antibodies,” say the researchers.
The majority of studies have so far focused on mutations within the spike protein, since they can alter host cell entry and susceptibility to infection- or vaccine-induced immunity.
“However, some of the mutations map to non-structural proteins, and it is thus plausible that they impact viral replication dynamics,” writes the team.
What did the current study involve?
Ilan and colleagues developed a single-molecule fluorescence in situ hybridization (smFISH) method that enabled them to visualize SARS-CoV-2 RNAs with high sensitivity and spatial precision.
Using this powerful new approach, the team quantified positive-sense RNA genomes with 95% detection efficiency while simultaneously visualizing negative-sense genomes, sub-genomic RNAs, and viral proteins.
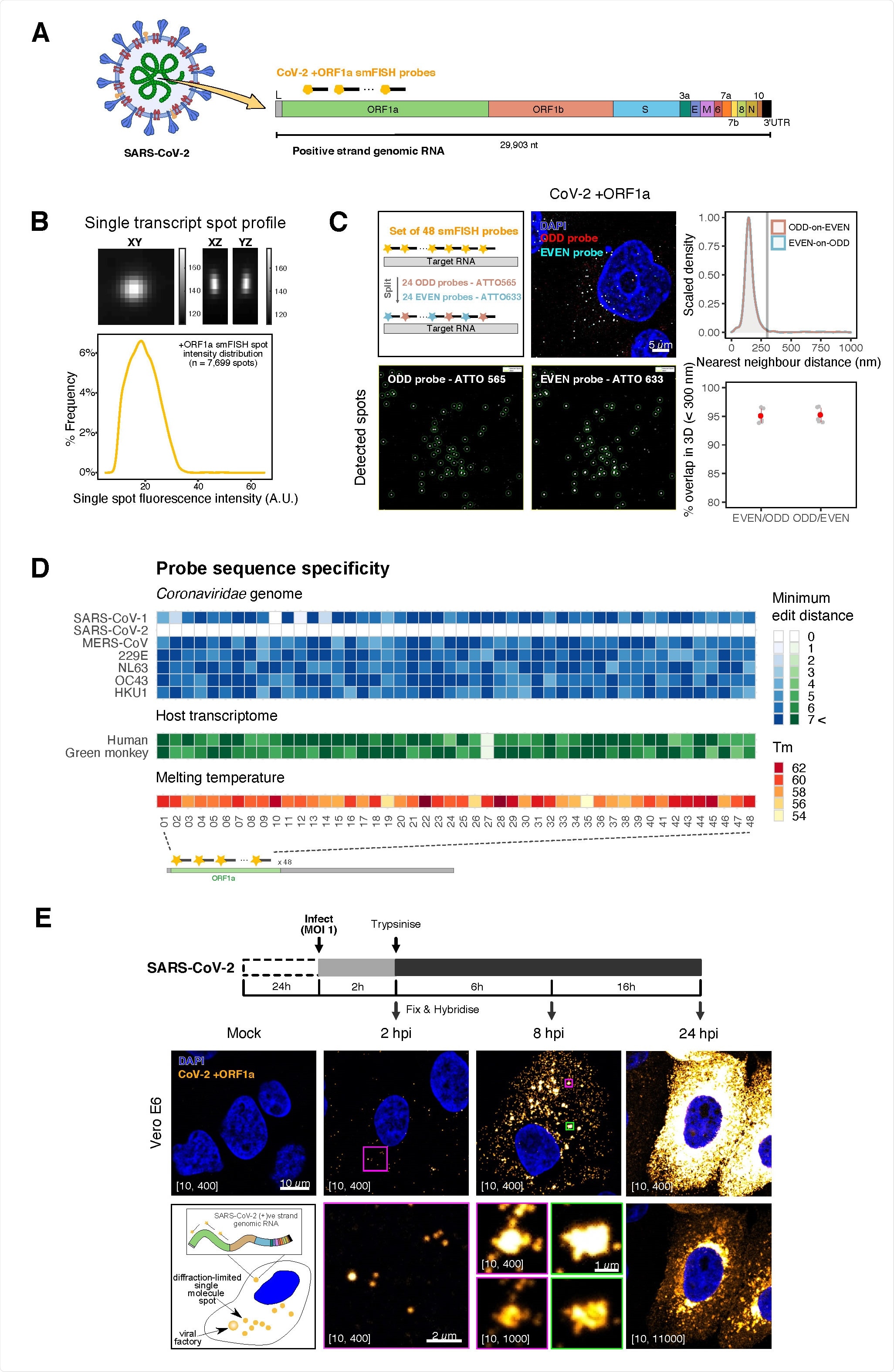
Sensitive single-molecule detection of SARS-CoV-2 genomic RNA in infected cells. (A) Schematic illustration of single-molecule fluorescence in situ hybridisation (smFISH) for detecting SARS-CoV-2 positive-strand genomic RNA (+gRNA) within infected cells. (B) Reference spatial profile of a diffraction-limited +ORF1a smFISH spot. The gradient legend represents relative fluorescence intensity (top). Frequency distribution of smFISH spot intensities, exhibiting a unimodal distribution (bottom). (C) Assessment of smFISH detection sensitivity by a dual-color co-detection method. Maximum intensity projected images and corresponding FISH-quant spot detection views of ODD and EVEN probe sets are shown. Scale bar = 5 µm. Density histogram of nearest11 neighbour distance from one spectral channel to another (top). Vertical line indicates 300 nm distance. Percentage overlap between spots detected by ODD and EVEN split probes, calculated bidirectionally (bottom). (D) Heatmap of probe sequence alignment against various Coronaviridae and host transcriptomes. Each column represents individual 20 nt +ORF1a probe sequences. The minimum edit distance represents mismatch scores, where ‘0’ indicates a perfect match. Melting temperatures of each probe at the smFISH hybridization condition are shown. (E) Experimental design for visualizing SARS-CoV-2 gRNA with smFISH at different time points after infection of Vero E6 cells. Cells were seeded on cover-glass and 24 h later, inoculated with SARS-CoV-2 (Victoria strain at MOI 1) for 2 h. Non-internalized viruses were removed by trypsin digestion and cells fixed at the timepoints shown. Representative 4 µm maximum intensity projection confocal images are shown. Numbers at the bottom left corner indicate dynamic contrast range used to display the image. Magnified view of insets in the upper panels are shown in lower panels. Scale bars = 10 µm or 2 µm.
What did they find?
The team found that SARS-CoV-2 gRNA persisted in the presence of the antiviral drug remdesivir, indicating a long half-life.
The researchers say this could reflect the high secondary structure of the RNA genome that may be refractory to degradation by cellular nucleases.
The study revealed that SARS-CoV-2 replication was highly variable between cells, with only a small proportion displaying a high burden of viral RNA.
The researchers say that recent sequencing studies of SARS-CoV-2-infected bronchial cultures identified ciliated cells as the primary target. However, only a minority of these cells contained viral RNA that may reflect variation in susceptibility.
Ilan and colleagues say that since the human respiratory tract encompasses the nasal passage, large and small airways, and bronchioles, knowledge of the specific cell types and their SARS-CoV-2 RNA burden is limited.
“Applying smFISH to clinical biopsies and experimentally-infected animal samples will allow us to address this important question,” they write.
An unexpected finding about B.1.1.7
Unexpectedly, the analysis revealed that the B.1.1.7 (Alpha) variant exhibited significantly slower replication kinetics than the Victoria isolate, yielding a lower number of gRNA and sgRNA copies per cell, fewer viral replication factories, and a lower frequency of “super-permissive” cells.
The team says a recent longitudinal study of nasopharyngeal swabs showed that B.1.1.7 was associated with longer infection times while showing similar peak viral loads to non-B.1.1.7 variants.
The authors of that study suggested that this extended duration of viral shedding may contribute to increased transmissibility, which Ilan and colleagues say would be consistent with their data showing reduced replication of B.1.1.7 at the single-cell level.
“Replication fitness will be defined by the relationship of the virus with its host cell,” say the researchers.
While aggressive replication may trigger cellular antiviral sensors, reduced replication may enable the virus to replicate and persist for longer before host antiviral sensors are triggered.
“Such differences, and their impact on host antiviral responses, are likely to be of key importance for our understanding of the success of viral variants to spread through the population,” concludes the team.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Davis I, et al. Absolute quantitation of individual SARS-CoV-2 RNA molecules: a new paradigm for infection dynamics and variant differences. bioRxiv, 2021. doi: https://doi.org/10.1101/2021.06.29.450133, https://www.biorxiv.org/content/10.1101/2021.06.29.450133v1
- Peer reviewed and published scientific report.
Lee, Jeffrey Y, Peter AC Wing, Dalia S Gala, Marko Noerenberg, Aino I Järvelin, Joshua Titlow, Xiaodong Zhuang, et al. 2022. “Absolute Quantitation of Individual SARS-CoV-2 RNA Molecules Provides a New Paradigm for Infection Dynamics and Variant Differences.” ELife 11 (January). https://doi.org/10.7554/elife.74153. https://elifesciences.org/articles/74153.