Following the outbreak of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which is the virus responsible for the coronavirus disease 2019 (COVID-19), there has been a significant increase in research focused on zoonoses.
Although the majority of these studies have been focused on bats, there should be a greater focus on rodents, particularly rats. Rodents are known to be important carriers or reservoirs for several zoonotic viruses such as Western equine encephalitis, lymphocytic choriomeningitis virus (LCMV), hemorrhagic fever with renal syndrome, Apoi virus disease, Omsk hemorrhagic fever, and hepatitis E cowpox. Rats are also found in almost every part of the earth; therefore, humans are most likely to interact with these animals as compared to any other wildlife species.
A new review published in the journal Viruses aimed to describe the current state of knowledge on Sialodacryoadenitis virus (SDAV), which is a rat coronavirus, along with its characteristics and potential zoonotic threat.
Classification of coronaviruses
Coronaviruses are known to belong to the order Nidovirales. Nidovirales are characterized by enveloped, positive-sense, non-segmented RNA viruses.
Viruses with the largest capacity for causing pandemics and epidemics belong to the suborder Cornidovirineae of order Nidovirales. A subfamily of Cornidovirineae, Coronaviridae is further subdivided into two families, Letovirinae and Orthocoronavirinae.
The Orthocoronavirinae comprises four genera: Alpha, Beta, Gamma, and Delta coronaviruses. The Alpha and Beta coronaviruses are known to infect bats and mammals, while the Gamma and Delta coronaviruses infect birds and few marine mammals.
An overview of rat coronaviruses
The first report of a rat coronavirus (RCoV) appeared in 1960 after laboratory rats were found to experience dacryoadenitis, destructive sialadenitis, and transmission disease of the lower respiratory tract. Following this, agents related to mouse hepatitis virus (MHV) were found in rat sera in 1964.
Parker’s rat coronavirus (RCV-P) was also isolated from the lungs of asymptomatic rats. A second strain is known to cause sialoadacryoadenitis (SDA) that was antigenically identical to RCV-P and MHV was also recognized. Thereafter, studies evaluated several other strains of RCoVs.
RCoVs belong to Beta coronaviruses, especially subgenera Embecovirus. Among the RCoVs, the China Rattus Coronavirus (ChRCoV) represents the murine lineage of Betacoronavirus 1 and their ability to undergo interspecies transmission from rodents to other mammals. SDAV and other RCoVs belong to the same genus as human coronaviruses that were capable of causing pandemics and epidemics throughout the world.
SDAV transmission, symptoms, and diagnosis
Experimental studies have indicated that RCoVs are capable of remaining infectious when dried on solid surfaces. RCoV infections spread easily, either by direct contact with infected individuals or through aerosol.
RCoVs are known to cause symptomatic and asymptomatic infections. There are also two models of infection. The first model involves the development of breeding colonies that further leads to epizootic development in young non-immune individuals.
The second model involves a sudden onset of episcleritis in naive rats from weaning to adulthood. Other common signs of SDAV infections include edema of the submaxillary salivary glands, ocular discharge, photophobia, lacrimation, corneal ulcers, corneal opacities, and cervical swelling due to inflammation. Additional ancillary effects may include weight loss, transient anorexia, and disruption of estrus.
Detection of SDAV and other RCoVs is achieved through serological tests and immunohistochemical techniques. Reverse-transcriptase polymerase chain reaction (RT-PCR) is also used for the detection of the M, N, and pol genes collected from infected tissue samples. Individuals who are diagnosed as positive are either eliminated or quarantined for 5 to 8 weeks.
Structure of SDAV and biological functions of proteins
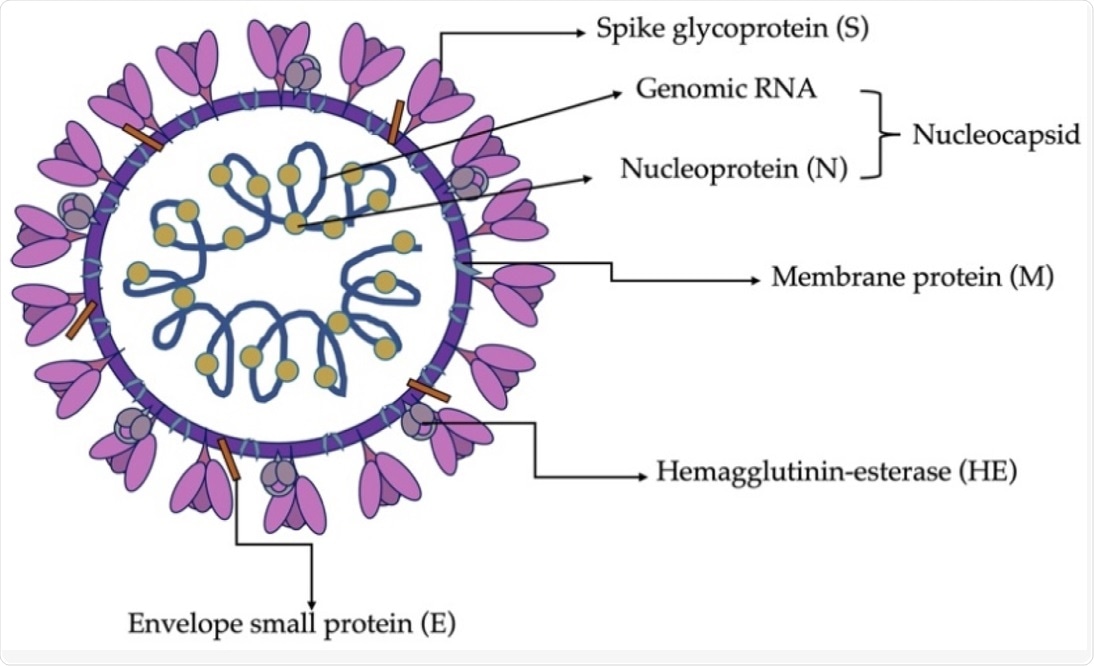 Schematic structure of SDAV virion (Bartak et al., 2021).
Schematic structure of SDAV virion (Bartak et al., 2021).
The SDAV virion is spherical with a diameter of 80 to 180 nanometers (nm). It comprises a genomic core surrounded by a membrane envelope and tentacles like spike proteins emanating from the surface. The viral genome comprises four major structural proteins that are encoded towards the 3’ end of the genome, of which include the membrane (M), spike (S), nucleocapsid (N) proteins, as well as the hemagglutinin esterase (HE).
The S protein is responsible for the entry of the virion into the host cell. The gene regions of the S protein are highly variable, heterogenous, and are responsible for tissue tropism and change of virulence.
The M protein is responsible for promoting membrane curvature by adapting the membrane region for virion assembly, as well as uptake of structural protein residues. The protein E works in association with the M protein during virion assembly and morphogenesis. It also helps in the formation of pentameric lipid pores in the host membrane through which ion transport can take place.
The N protein aids in the viral replication process and limits translation in host cells. The M, E, and N proteins work together to bring about the folding and budding of a newly assembled virion. The role of the HE protein is virion attachment, wherein it functions as a supporting binding molecule in addition to the S protein.
Cell infection and virus replication
SDAV is capable of infecting a wide variety of cell types such as epithelial cells of the respiratory tract, central nervous system (CNS) cells, or mononuclear cells in lymphoid organs. The exact receptor that promotes SDAV cell entry has not been confirmed yet. It is assumed that SDAV utilizes the mechanism of the human Betacoronavirus-1 and attaches to the host sialic acid receptor through their S protein to promote cell entry.
Following fusion with the host membrane with the help of S and HE proteins, proteolytic cleavage of the S protein takes place. Cleavage of the S protein generates two different subunits, S1 and S2.
Cleavage is known to take place at two different points, the first one induced by furin and the second one induced by TMPRRS2 protease, cathepsin, or any other protein. This results in the formation of the S2’ fusion peptide that fuses the host and viral membrane, thus introducing the viral RNA into the cytoplasm.
This is followed by translation of the replicase gene that helps in viral replication. Translation of the replicase gene results in sixteen non-structural proteins (nsps) that assemble into the replicase-transcriptase complex (RTC) and carry out viral replication.
The replication involves a negative sense genomic RNA that is used as a template for the synthesis of the positive strand. Replication is followed by transcription, which generates a subgenomic positive sense messenger RNA (mRNA).
The mRNA then undergoes translation to generate the structural and accessory proteins. This is followed by the generation of mature virions and their exocytosis. The exact mechanism of egress for SDAV has not been described yet.
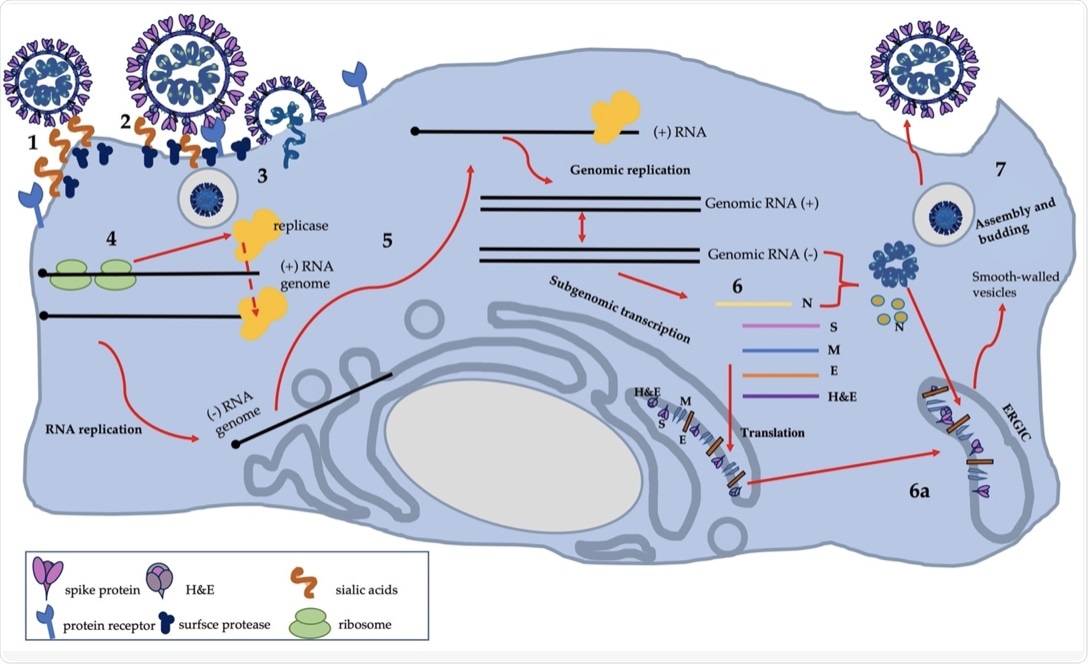 The scheme of SDAV replication: own work based on available knowledge (Bartak et al., 2021).
The scheme of SDAV replication: own work based on available knowledge (Bartak et al., 2021).
Genomic structure and gene functions of SDAV
Characterization of the genomic structure of SDAV was achieved by sequencing the 3’ terminal 9.8 kb of the genomic RNA. Seven cDNA fragments were generated by RT-PCR complementary DNA (cDNA) cloning that represented the 3’ terminal of the genome. The coding sequence for S, M, and N structural proteins, as well as non-structural proteins, NS2, and HE gene, were confirmed to belong to nine open reading frames (ORFs).
The coding sequence for ORF5a, non-structural protein, M protein, and ORF7b, small internal ORF was present in the +1 frame. The coding sequence for polymerase 1b, 15k nonstructural protein, small membrane protein, and S protein was present in the +2 frame.
The sequences for N protein, nonstructural protein NS2, and HE protein were present in the +3 frame. Furthermore, each of the identified protein genes was found to contain a short intergenic consensus sequence.
SDAV propagation and role of in vitro studies
Previously SDAV was used in evaluating the response of alveolar epithelial cells to infection. Recently, it was identified that the uninfected alveolar type I cells were able to produce chemokine in response to interleukin 1 (IL-1) produced by SDAV infected cells.
Chemokine production was also observed in alveolar type II cells and airway epithelial cells post SDAV infection. Alveolar type I epithelial cells was therefore detected as the primary target for SDAV infection.
Current studies also show that SDAV in vitro propagation is now possible in various established cell lines, such as murine cell lines and rat kidney cell cultures that make experimental studies easier.
Conclusion
The growing number of new coronaviruses is of concern due to the close relationship between the coronaviruses with distantly related animals. In spite of having commonalities with zoonotic viruses, the zoonotic potential of SDAV has not been explored yet. The major reason for this can be the insufficient knowledge of the entire replication cycle of the virus.
Symptomatic SDAV infection can be treated easily, while asymptomatic or mild cases are difficult to treat and can provide the virus a suitable environment to mutate. Poor hygienic conditions in rat or mouse cages are responsible for SDAV transmission.
Although current studies on respiratory infections of SDAV have been described, further research needs to be carried out for the identification of virus-cell interactions in different cell models. Further studies are also needed to characterize SDAV infections and determine their zoonotic potential that can pose a threat to humans in the future.