Somatic mutations in healthy cells advance the development of cancer and have been speculated to be contributors to aging. Decade-long hypothesis on the rates of somatic mutation correlates cancer risk to body mass. Another theory cites the role of these mutations in aging.
Until recently, it had been challenging to study the somatic mutations in normal tissues as it is arduous to detect the mutations in single cells or small clones. But, interestingly, recent technological advancements have enabled the study of normal tissues for somatic mutations.
In healthy tissues, the pattern and rates of these mutations remain largely unknown, except for humans.
 Study: Somatic mutation rates scale with lifespan across mammals. Image Credit: Konstantin Faraktinov / Shutterstock
Study: Somatic mutation rates scale with lifespan across mammals. Image Credit: Konstantin Faraktinov / Shutterstock
The study
A recent study published in the journal Nature aimed to elucidate, through a comparative analysis, the diversity in mutagenesis across different species, and the long-established hypothesis regarding evolution in the rates of somatic mutation, as well as their role in cancer and aging.
To investigate the landscape of somatic mutation across 16 mammalian species, whole-genome sequencing was carried out of 208 intestinal crypts from 56 multicellular living beings – belonging to different species with wide variations in lifespans and anatomical dimensions. These species were namely – humans, black-and-white colobus monkeys, ferrets, mice, cats, dogs, naked mole-rats, rats, ring-tailed lemurs, harbor porpoises, rabbits, cows, horses, giraffes, lions, and tigers.
Findings
The results of whole-genome sequencing of the colorectal crypts showed some significant similarities in the mutational spectrum of these organisms, despite differences in their diet and life histories. Furthermore, the analysis suggested that the three signatures dominating the somatic mutagenesis in humans, namely SBS1, SBSB, and SBSC, were also dominant in other mammals but with a variable contribution.
Based on this, it could be argued that somatic mutagenesis in different mammalian species is dominated by a set of preserved mutational processes.
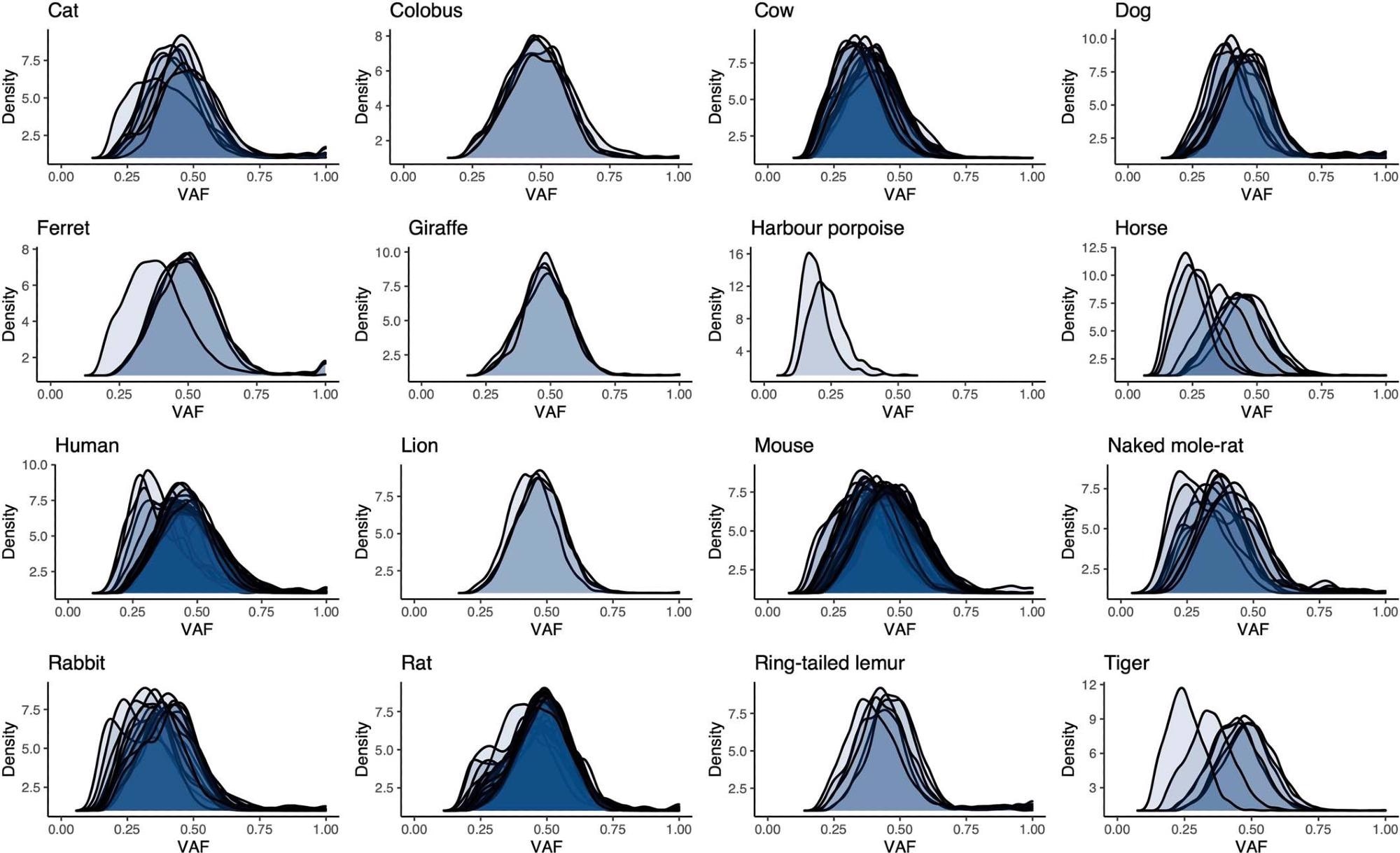 Distributions of variant allele fraction (VAF) for somatic substitutions in each crypt for each species. Each distribution refers to the variants in a single sequenced crypt.
Distributions of variant allele fraction (VAF) for somatic substitutions in each crypt for each species. Each distribution refers to the variants in a single sequenced crypt.
The most significant finding of this study was the inverse scaling or rates of somatic mutation and lifespan – a well-established prediction of the theory of aging via somatic mutation.
The mechanistic and evolutionary models of aging predict that its process is dependent on several factors. It states that various forms of molecular and cellular damage contribute to the aging of the organism due to its evolutionary constraints to the selection, which again act upon the rate of aging. This finding was supported by studies that reported more efficient DNA repair in species with longer lifespans.
Moreover, somatic mutations affect aging in various ways. This study brings forth the possibility that somatic mutations could contribute to aging through clonal expansions of the functionally altered cells in human tissues, for example – the possible association of mutations in liver disease and insulin resistance.
This phenomenon can be further comprehended by an in-depth understanding of somatic mutations and clonal expansions in age-related diseases and phenotypes. Similar inverse correlations with different forms of aging-related molecular damage and lifespans are expected and previously reported, such as telomere shortening and protein turnover.
There exists a need to consider other non-casual inverse correlations between lifespan and somatic mutations. One such interpretation is the possible scaling of cell division rates with lifespan and explaining the observed rates of somatic mutation. However, the cell-division rate estimates available reject this. Another explanation could be the fact that the rate of cell division in humans is not a detrimental factor for somatic mutation rates.
An additional possible cause for the observed inverse correlation might be that selection reduces the rates of a germline mutation in species with comparatively longer reproductive spans. The selective pressure on germline mutations might have some influence on the somatic mutation rates. Yet, germline mutation rates firmly determining the somatic mutation rates is not likely since, in humans, somatic mutation rates are 10-20 times higher when compared to the germline mutation rates and exhibit variability in different cell types.
Conclusion
Therefore, it can be concluded that the somatic mutation rates have themselves been constrained by an evolutionary process, perhaps owing to the selection of multiple DNA-repair pathways.
On the whole, this study gives a detailed account of the mutational processes and contrasts the common and variable features across mammals. Further research will help expand our knowledge regarding somatic mutations and their influence on disease, aging, and evolution.