Due to the long-term ramifications and widespread demographic impact of the pandemic, the origins of the medieval Black Death pandemic from AD 1346 to 1353 have been a subject of ongoing inquiries. Unfortunately, despite the extensive multidisciplinary investigations, the location where the second plague pandemic originated is unknown.
To date, the most contentious archaeological evidence for the start of the Black Death pandemic has come from cemeteries around Lake Issyk-Kul in modern-day Kyrgyzstan. Since tombstone inscriptions precisely dated to 1338-1339 mention "pestilence" as the cause of death, these sites likely housed victims of a 14th-century epidemic.
About the study
In the present work, the researchers document ancient deoxyribonucleic acid (aDNA) information from seven people exhumed from the Burana and Kara-Djigach cemeteries near Lake Issyk-Kul to investigate probable evidence associated with the origin of the second plague pandemic. Excavations of these cemeteries around 1885 and 1892 discovered a distinctive archaeological assemblage that could be linked to a 14th-century epidemic that affected the area.
 Lake Issyk-Kul in the summer day, Kyrgyzstan. Image Credit: V. Smirnov / Shutterstock
Lake Issyk-Kul in the summer day, Kyrgyzstan. Image Credit: V. Smirnov / Shutterstock
To learn more about the circumstances of the Burana and Kara-Djigach cemeteries, the team translated and studied the existing archival evidence from the excavations of these graves. Then, they used a hybridization capture of roughly 1.24 million ancestry-informative single-nucleotide polymorphisms (SNPs) to generate human genomic data from seven individuals (five from Kara-Djigach and two from Burana). This process resulted in four individuals with adequate genomic coverage for population genetic evaluations, i.e., >30,000 SNPs. In addition, the scientists used ancestry modeling and component analysis to analyze the generated genomic data from the seven individuals.
Shotgun metagenomic details obtained from the seven individuals were taxonomically sorted utilizing heuristic operations for pathogen screening (HOPS) pipeline to assess signs of ancient pathogen DNA that could elucidate the cause of the presumed epidemic. The authors also analyzed the SNP patterns of the BSK003 and BSK001 Y. pestis genomes to see if they represented different bacterial strains. They used the MALT software to perform taxonomic-informed metagenomic screening to reduce variant calls resulting from environmental contamination, which was significant considering the high number of multi-allelic domains found in both genomes.
The investigators compared the Kara-Djigach genomes to previously documented historical and presently circulating Y. pestis variety in an SNP analysis. They also calculated Faith's phylogenetic diversity (FPD) indices and the mean pairwise distances (MPDs) in 203 genomes from the entire modern dataset and 130 genomes from branches 1–4 to calculate the amount of present-day Y. pestis genetic diversity that originated from the branch 1–4 polytomy. The scientists examined the likelihood of a local emergence relative to the BSK001/003 strains' introduction into the Chüy Valley from another area to resolve current hypotheses about the geographical origins of the Black Death pandemic.
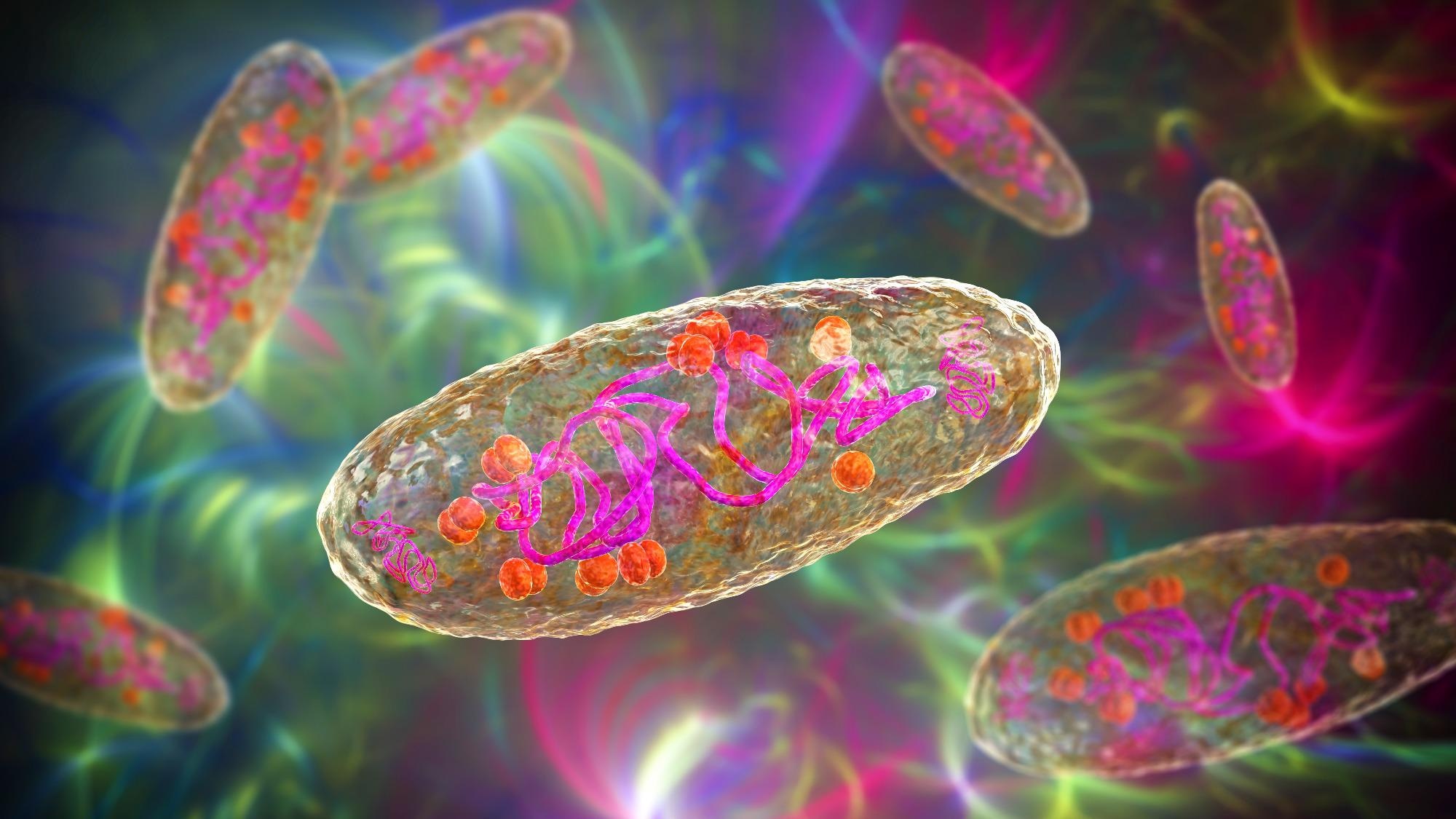 Plague bacterium Yersinia pestis, scientifically accurate 3D illustration showing a structure of the cell with DNA, plasmids and ribosomes. Image Credit: Kateryna Kon / Shutterstock
Plague bacterium Yersinia pestis, scientifically accurate 3D illustration showing a structure of the cell with DNA, plasmids and ribosomes. Image Credit: Kateryna Kon / Shutterstock
Results and conclusions
The current analysis of historical, ancient, and archaeological genomic evidence reveals that the plague bacterium Yersinia pestis was involved in the Black Death pandemic. The most recent shared ancestor of a prominent diversification generally linked with the pandemic's development, here dated to the initial half of the 14th century, was identified as two rebuilt ancient Y. pestis genomes (BSK003 and BSK001), which form a single strain.
The local arousal of the recovered ancient sequence was supported by comparisons with current-day diversity from Y. pestis reservoirs in the wider Tian Shan area. The present data support a source for the second plague pandemic in central Eurasia in the early 14th century, based on various lines of proof.
Overall, the current research presents ancient Y. pestis evidence from central Eurasia that suggest a 14th-century outbreak; thus, the previous outbreak attributions necessitate further investigations. Currently, the study's narrow-focused sampling does not permit an evaluation of the BSK003/001 strain's spread. Previous analyses have demonstrated that Y. pestis may spread quickly without accumulating genetic variation, allowing for the simultaneous existence of the same sequence over a wide geographic area.
The current study provides no proof to infer links between environmental elements and the Kara-Djigach epidemic. However, the authors believe that the new precise 1338 to 1339 date would be a benchmark for future archaeological, environmental, and historical studies concentrating on the incidents that caused the Y. pestis emergence in humans and provoked the second plague pandemic.
The scientists point out that past and current experiences showed that determining the origin of a pandemic was a difficult undertaking that cannot be completed by a single scientific discipline. Although the ancient Y. pestis genomes revealed in this publication offer biological data to settle an old controversy, the scope and value of the current investigation were defined by the distinct historical and archaeological circumstances. As a result, the team expects future synergies to provide crucial information for a thorough reconstruction of the dynamics that led to the second plague pandemic.