Introduction
Antibiotic resistance (AMR)
Why Soil is Important for Antibiotic Discovery
Historical Role of Soil in Medicine
Why Discovery Slowed Down
Renewed Interest in Soil Microbiomes
How Soil Microbes Produce Antibiotics
Modern Technologies Driving Discovery
Examples of Recent Discoveries
Challenges and Limitations
Broader Implications for Health and Environment
Future Directions
Conclusions
References
Further reading
Beneath our feet lies a hidden pharmaceutical frontier: soil microbes are yielding new antibiotic leads just as scientists race to outpace the global rise of antimicrobial resistance.
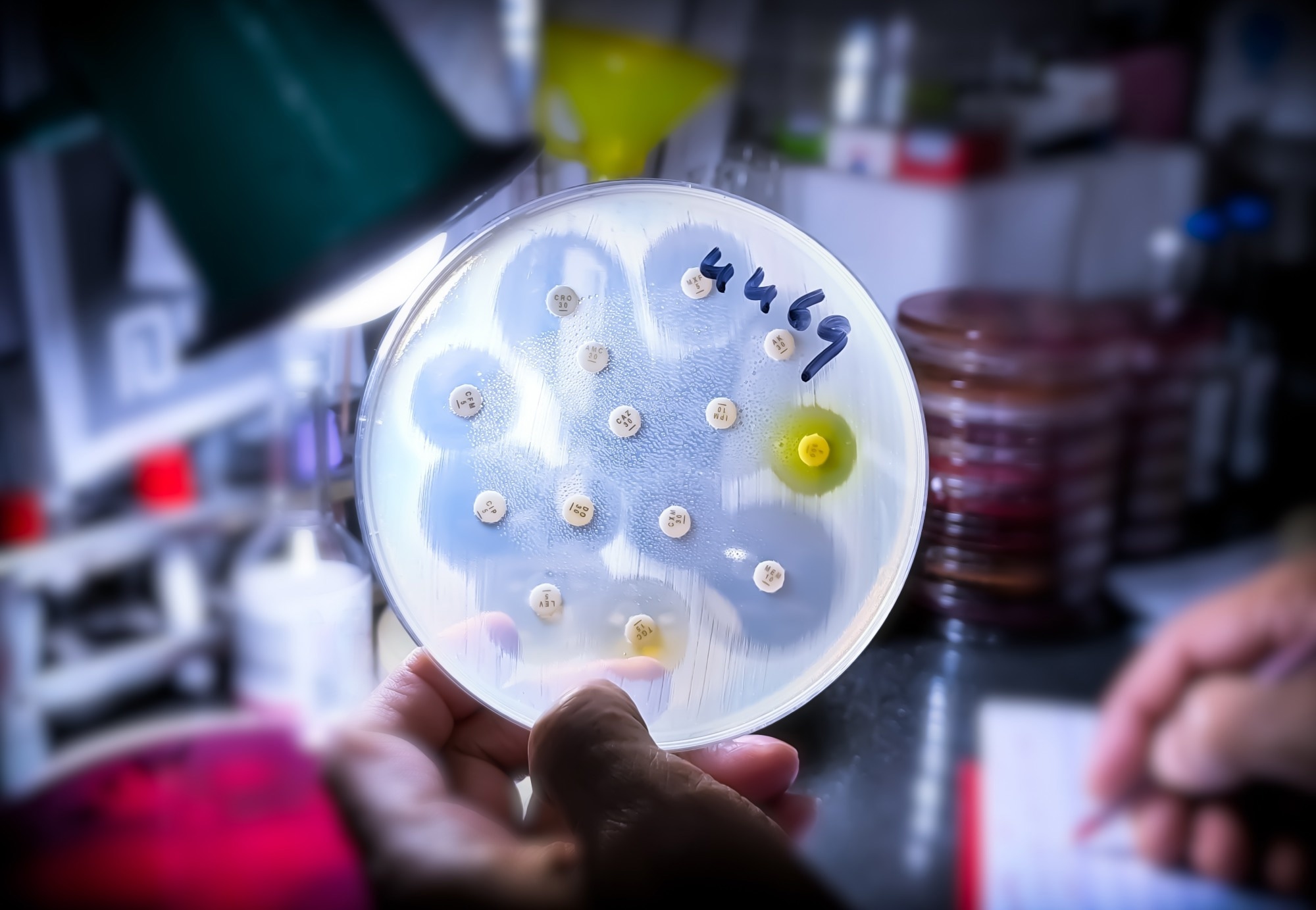
Antimicrobial susceptibility testing in petri dish. Image Credit: Saiful52 / Shutterstock
Introduction
Antimicrobial resistance (AMR) is one of the most serious threats to modern medicine, with recent reviews citing roughly 4.95 million deaths associated with antibacterial resistance globally in 2019.1,3
This burden is rising while the supply of genuinely innovative antibiotics remains limited, especially for difficult-to-treat Gram-negative pathogens.3,9
Decades of research implied that the "Waksman platform" - the traditional method of culturing soil microbes - seemed exhausted, leading to a “discovery void” of mechanistically distinct pharmacological interventions against bacterial strains resistant to conventional antibiotic therapies.
However, technological advances in genomic sequencing, in situ cultivation technologies like the isolation chip (iChip), and, most recently, AI-assisted antibiotic discovery have revealed that the "Golden Age" of antibiotics barely scratched the surface of microbial "dark matter."3,6
This article synthesizes research on how soil biodiversity, cultivation technology, and genome-guided discovery are renewing the search for antibiotics while also highlighting soil’s parallel role as a reservoir of resistance genes.1,3,5
Antibiotic resistance (AMR)
Antibiotic resistance (AMR) has evolved into a global health threat that endangers the foundations of modern infectious disease management. Reviews in the reference set note that antibacterial resistance was associated with about 4.95 million deaths globally in 2019, including a smaller subset directly attributable to resistance.3
This ongoing public health crisis is compounded by a thin development pipeline: one 2025 review notes that only 13 new antibacterial drugs were approved between 2017 and 2023, and only two met at least one WHO innovation criterion, such as novelty of class, target, mode of action, or lack of cross-resistance.3
Why Soil is Important for Antibiotic Discovery
Modern medical research, however, is increasingly investigating soil as the next frontier of antimicrobial biomolecule discovery, primarily because soil microbes have been observed to produce natural inhibitory compounds that enable them to compete and survive in hyper-competitive environments.1 Many existing antibiotics, including the major classes like tetracyclines and aminoglycosides, were originally derived from these soil organisms.2
Soil remains especially important because it is among the richest known microbial habitats and has historically yielded many clinically useful natural products, even though only a small fraction of its biosynthetic potential has been explored.1,3,5
Recent genome-mining work also shows that soil- and fermented-food-associated Bacillus strains harbor abundant biosynthetic capacity: in one study, 83% of analyzed genomes encoded the siderophore bacillibactin clusters, while others encoded surfactins, fengycins, iturins, kurstakins, bacitracin, and several previously uncharacterized non-ribosomal peptide synthetase clusters.4
Historical Role of Soil in Medicine
The mid-20th century is popularly considered the “Golden Age” of antibiotic discovery, a period defined by the systematic screening of soil microorganisms. This strategy yielded many of the microbial natural products that still underpin antibacterial therapy today.3,5
Unfortunately, despite these early Golden Age successes, the discovery rates for novel antibiotics sharply declined by the late 1960s, a phenomenon now attributed to the "re-isolation problem," in which traditional phenotypic screening methods repeatedly identified the same known compounds from ubiquitous soil species. This observation prompted researchers of the time to erroneously assume that the accessible biosynthetic potential of soil had been exhausted.5
Why Discovery Slowed Down
Current microbial research attributes these declines in antibiotic discovery rates to a technical limitation known as “the culturability crisis” – approximately 99% of environmental microorganisms are estimated to be unculturable under standard laboratory conditions.5,8 Studies have shown that standard agar plates fail to provide the variable nutrients and ecological signaling molecules that most soil-based bacteria require to grow.6
High rediscovery rates of biomolecules (antimicrobial scaffolds) are suggested to have further discouraged financial and resource investment in soil-centric bioprospecting, prompting the pharmaceutical industry to shift its focus toward synthetic drug development.5
That shift did not solve the problem: target-based and synthetic screening often produced compounds active in vitro but lacking whole-cell activity, particularly against Gram-negative bacteria whose outer membranes and efflux systems restrict compound accumulation.3
Renewed Interest in Soil Microbiomes
Modern advances in next-generation sequencing technologies, particularly genomics and metagenomics, are now enabling the study of previously unculturable microbes. These technologies have demonstrated that soil is an abundant reservoir of bioactive compounds synthesized by cryptic gene sequences that remain unexpressed under standard laboratory conditions.4
Furthermore, advances in computing power and the simultaneous development of previously computationally restrictive bioinformatics tools now allow for the identification of natural product biosynthetic pathways directly from metagenomic data, bypassing the need for initial laboratory-based cultivation or isolation.4
Together, metagenomics, genome mining, and resistome analysis have renewed interest in soil microbiomes as both a source of new antibiotics and a system for understanding how natural and acquired resistance evolve in the environment.1,3,7
How Soil Microbes Produce Antibiotics
Microbe-derived antibiotics are naturally synthesized as secondary metabolites – biomolecules which are not essential for primary growth but are used for ecological competition and inter-species signaling.7
These compounds include non-ribosomal peptides produced by non-ribosomal peptide synthetases (NRPSs), as well as ribosomally synthesized and post-translationally modified peptides (RiPPs) such as lasso peptides.4,9
NRPS systems function as modular assembly lines that can incorporate both proteinogenic and non-proteinogenic building blocks, helping generate the structural diversity that makes soil microbes such valuable sources of bioactive molecules.4,9
Modern Technologies Driving Discovery
Next-generation sequencing approaches and their derivatives have revolutionized modern drug discovery. Metagenomic sequencing and environmental DNA (eDNA) allow for the identification of potential antibiotic-producing species directly from soil samples.4 Genome mining and BGC analysis are subsequently used to prioritize novel chemical scaffolds before their industrial-scale synthesis.3
Emergent soil microbial cultivation techniques, particularly the isolation chip (iChip) – a high-throughput device that enables the cultivation of microorganisms that cannot be cultured using traditional agar plate-based approaches – are capable of simulating natural environments by using novel semipermeable membranes.6
In the classic iChip workflow, a diluted soil sample is loaded so that roughly one bacterial cell occupies a chamber; the device is then enclosed by semipermeable membranes and returned to soil, where natural nutrients and growth factors diffuse into the device during incubation.6,8
This in situ strategy was developed to recover slow-growing or previously uncultured organisms that are often missed by conventional plate cultivation.5,6,8
Examples of Recent Discoveries
These modern technologies have already led to the identification of new antibiotic classes from soil bacteria. For example, Teixobactin is a novel antibiotic isolated from the bacterium Eleftheria terrae via the application of the iChip. It inhibits cell wall synthesis by binding to highly conserved lipid precursors, including lipid II and lipid III, and the original report did not recover resistant mutants of Staphylococcus aureus or Mycobacterium tuberculosis under the tested laboratory conditions.8
Another recent example is lariocidin, a lasso peptide produced by Paenibacillus sp. M2. It binds to a unique site on the small ribosomal subunit, interferes with protein synthesis, shows broad-spectrum activity, and displays a low propensity for spontaneous resistance, together with efficacy in a mouse model of Acinetobacter baumannii infection.9
Challenges and Limitations
Despite recent surges in the number of soil bacteria-derived antibiotic candidates, reviews highlight that translating these discoveries into clinically useful drugs remains a significant hurdle, with discovery and development failure rates still extremely high.3
Major obstacles include rediscovery of known chemistry, difficulties obtaining sufficient compound quantities, toxicity and pharmacokinetic problems, and poor penetration into Gram-negative bacteria.3,5
Even promising leads must be interpreted cautiously: for example, teixobactin showed strong preclinical properties in the original paper, but broader clinical translation still requires further optimization and testing.8
Broader Implications for Health and Environment
Recent research emphasizes that soil functions as a source of life-saving antibiotics, but also a reservoir of resistance genes – antibiotic residues, resistant bacteria, and resistance genes can enter soils through manure, sewage sludge, reclaimed wastewater, and other agricultural inputs.1,2,7
Reviews in the reference set therefore frame soil within both “planetary health” and “One Health” perspectives: it is a reservoir of antimicrobial innovation, but also a pathway through which resistance determinants can move among soil, water, food, animals, and humans.1,4,7
Future Directions
AI-assisted methods are increasingly being explored in the modern drug discovery pipeline to help prioritize and screen millions of virtual molecules for potential antibiotic activity.3
Future progress will likely depend on combining AI-assisted screening with metagenomics, genome mining, and improved cultivation platforms that expand access to previously uncultured microbes.3,5,6
On the environmental side, current reviews also discuss interventions such as tighter antibiotic-use regulation, manure management, soil monitoring, and remediation approaches including biochar application to help reduce agricultural ARG burdens.7
Getting the dirt: A global atlas of antibiotic resistance genes in soil microbes
Conclusions
The evidence reviewed here shows that advances in cultivation, sequencing, and genome mining have reopened soil as a productive source of antibiotic discovery, helping address long-standing barriers such as rediscovery and microbial unculturability. These technologies are overcoming the historical challenges of re-isolation and the technical limits of culturability, providing novel scaffolds that can bypass prevalent resistance paradigms.3,5,6
At the same time, the same soil systems that yield antibiotics also store and disseminate resistance genes, so future success will depend on pairing bioprospecting with environmental stewardship. Soil still matters in medicine not only because it contains new drug leads, but because soil health and microbial ecology shape the long-term sustainability of antibiotic therapy itself.1,3,7
References
- Zhu, Y.-G., et al. (2019). Soil biota, antimicrobial resistance and planetary health. Environment International, 131, 105059. DOI – 10.1016/j.envint.2019.105059. https://www.sciencedirect.com/science/article/pii/S0160412019310177?via%3Dihub
- Cycoń, M., Mrozik, A., & Piotrowska-Seget, Z. (2019). Antibiotics in the Soil Environment - Degradation and Their Impact on Microbial Activity and Diversity. Frontiers in Microbiology, 10. DOI – 10.3389/fmicb.2019.00338. https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2019.00338/full
- Cardona, S. T., Rahman, A. S. M. Z., & Novomisky Nechcoff, J. (2025). Innovative perspectives on the discovery of small molecule antibiotics. Npj Antimicrobials and Resistance, 3(1). DOI – 10.1038/s44259-025-00089-0. https://www.nature.com/articles/s44259-025-00089-0
- Waongo, B., Ndayishimiye, L., Tapsoba, F., Zongo, W.-S. A., Li, J., & Savadogo, A. (2025). Prospection for potential new non-ribosomal peptide gene clusters in Bacillus genus isolated from fermented foods and soil through genome mining. Frontiers in Microbiology, 16. DOI – 10.3389/fmicb.2025.1515483. https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2025.1515483/full
- Hegemann, J. D., Birkelbach, J., Walesch, S., & Müller, R. (2022). Current developments in antibiotic discovery. EMBO Reports, 24(1). DOI – 10.15252/embr.202256184. https://link.springer.com/article/10.15252/embr.202256184
- Chabib, L., et al. (2025). Harnessing iChip technology for novel antibiotic discovery from peat soil microbiomes to combat antimicrobial resistance. Frontiers in Microbiology, 16. DOI – 10.3389/fmicb.2025.1530273. https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2025.1530273/full
- Zeng, Y., Feng, R., Huang, C., Liu, J., & Yang, F. (2025). Antibiotic Resistance Genes in Agricultural Soils: A Comprehensive Review of the Hidden Crisis and Exploring Control Strategies. Toxics, 13(4), 239. DOI – 10.3390/toxics13040239. https://www.mdpi.com/2305-6304/13/4/239
- Ling, L. L., et al. (2015). A new antibiotic kills pathogens without detectable resistance. Nature, 517, 455–459. DOI – 10.1038/nature14098. https://www.nature.com/articles/nature14098
- Jangra, M., et al. (2025). A broad-spectrum lasso peptide antibiotic targeting the bacterial ribosome. Nature, 640(8060), 1022–1030. DOI – 10.1038/s41586-025-08723-7. https://pmc.ncbi.nlm.nih.gov/articles/PMC12497486/
Further Reading
Last Updated: Apr 17, 2026