Introduction
ESKAPE pathogens
Mechanisms of antibiotic resistance
Biofilm formation
Enzymatic drug inactivation
Efflux pumps
Challenges in treatment and control
Emerging strategies to combat ESKAPE pathogens
References
Further reading
ESKAPE pathogens are a group of opportunistic bacteria that frequently evade antibiotics through resistance mechanisms such as drug inactivation, efflux pumps, biofilms, and altered drug targets. These organisms drive difficult-to-treat hospital infections and are increasingly viewed through a One Health lens because resistance can spread through healthcare, food, water, animals, and the environment.
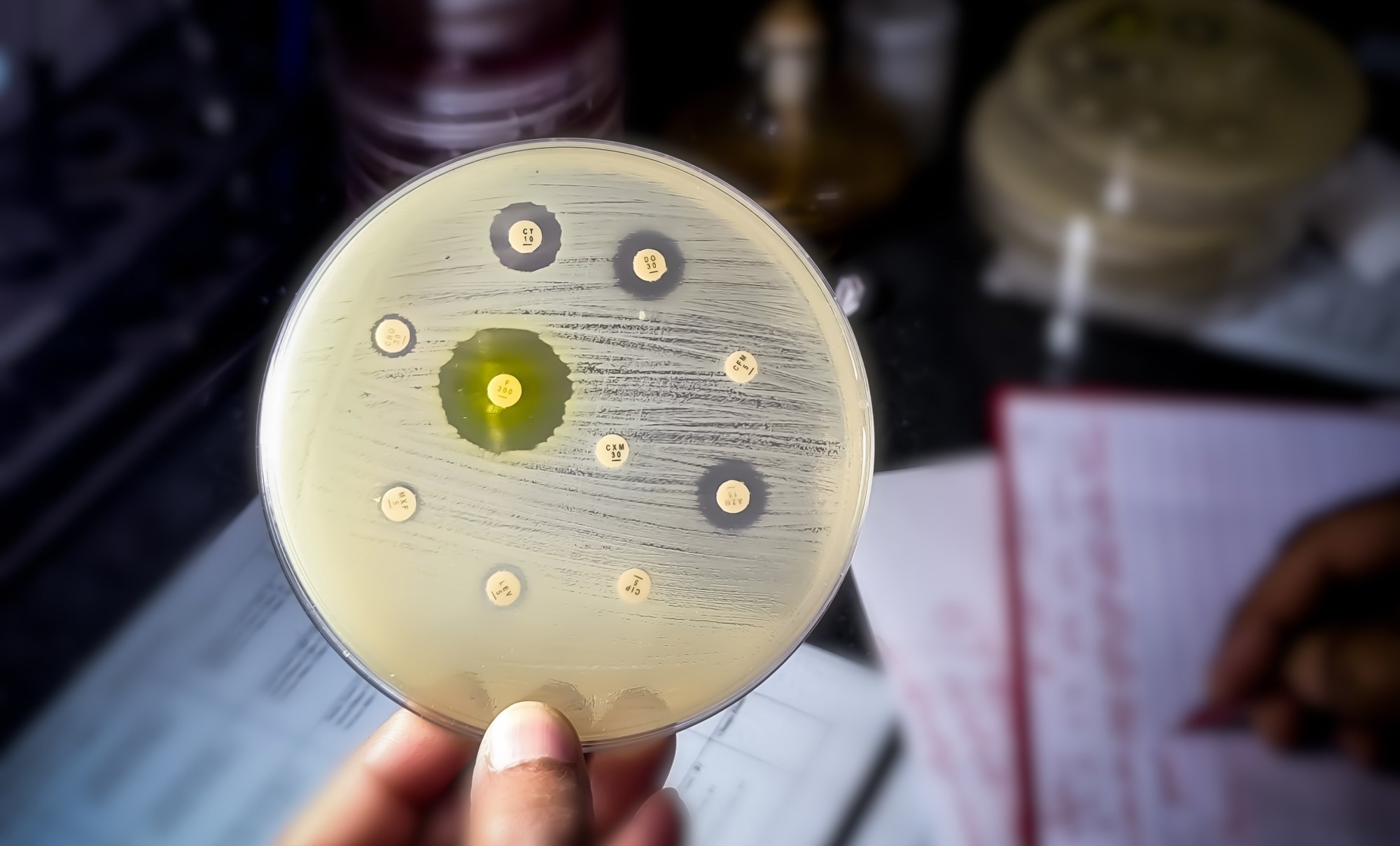 Image Credit: Saiful52 / Shutterstock.com
Image Credit: Saiful52 / Shutterstock.com
Introduction
Antimicrobial resistance (AMR) is a major global health threat caused by antibiotic overuse, genetic adaptability, and the spread of resistance genes. This article discusses the role of ESKAPE pathogens in AMR, mechanisms of drug evasion, clinical impact, and emerging strategies to combat these high-risk hospital-associated infections. Because these organisms are opportunistic pathogens, they are especially problematic in immunocompromised patients and in people exposed to invasive procedures or medical devices.1
ESKAPE pathogens
ESKAPE pathogens comprise six clinically important opportunistic bacteria that are often multidrug-resistant or associated with multidrug-resistant strains:1,2,3 Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter (ESKAPE) species.1,2 Some recent One Health literature also uses the expanded term ESKAPEE to include Escherichia coli, particularly when discussing food, water, livestock, and environmental reservoirs of resistance.4 Collectively, these bacteria utilize mechanisms including enzymatic degradation of antibiotics, efflux pumps, and reduced outer membrane permeability to reduce antibiotic efficacy.3
Infections with ESKAPE pathogens are associated with high mortality rates and other severe outcomes like prolonged hospitalization, higher treatment failure rates, and increased risk of intensive care unit (ICU) admission.1,3,4 Hospital-acquired infections like catheter-associated urinary tract infections (UTIs), ventilator-associated pneumonia, and surgical site infections are often caused by ESKAPE pathogens, with the food chain serving as another source of resistance genes that transfer to microorganisms within the gut microbiota. These pathogens can also persist on hospital surfaces, complicating infection prevention and facilitating transmission within healthcare environments.1
 Image Credit: Coronea Borealis Studio / Shutterstock.com
Image Credit: Coronea Borealis Studio / Shutterstock.com
Mechanisms of antibiotic resistance
AMR can arise through innate or intrinsic resistance, which refers to the inherent insensitivity of a bacterial strain to an antibiotic or antibiotic class3. Bacteria can also acquire resistance to an antibiotic to which they were previously sensitive through mutations or the exchange of genetic material between bacteria.
Common acquired resistance mechanisms include overexpression of efflux pumps, modifications to bacterial cell permeability or antibiotic target sites3,4, and enzymatic inactivation of antibiotics. Acquired mechanisms can arise following exposure to selective pressure or the incorporation of new genetic elements, such as plasmids, transposons, or integrative elements, into the bacterial genome via horizontal transfer. Key resistance pathways in ESKAPE pathogens are often grouped into four broad categories: reduced drug uptake, target-site modification, enzyme-mediated drug inactivation, and active drug efflux.3
ESKAPE bacteria first adhere to host tissues, medical devices, or environmental surfaces and then form microcolonies that mature into biofilms. During infection, they may release proteins, enzymes, siderophores, and toxins that support colonization, tissue invasion, and immune evasion. Species-specific factors such as capsules, fimbriae, adhesins, secretion systems, and quorum-sensing signals further shape virulence and biofilm behavior. Bacterial adhesion ultimately results in the formation of biofilms, a physical barrier that protects pathogens from antibiotics and other environmental stress factors.2 The ability to form biofilms also allows ESKAPE pathogens to persist on environmental surfaces and resist killing by disinfectants. Biofilms may further increase antimicrobial tolerance by restricting drug penetration, altering bacterial metabolism, and promoting horizontal gene transfer within densely packed microbial communities.2,4
Enzymatic drug inactivation
One of the most common classes of antibiotics is β-lactams, which target penicillin-binding proteins (PBPs) and carboxypeptidases involved in peptidoglycan synthesis, thereby inducing bacterial death. ESKAPE pathogens can resist β-lactams through mechanisms that vary by species, including β-lactamase production in many Gram-negative members and altered PBPs in methicillin-resistant S. aureus. In β-lactamase-producing strains, these enzymes hydrolyze the β-lactam ring, thereby inactivating the antibiotic and limiting effective PBP inhibition.3,4
Other enzymes that inactivate or degrade antibiotics include aminoglycoside-modifying enzymes, carbapenemases, and chloramphenicol acetyltransferases (CATs). Specifically, CATs transfer an acetyl group from acetyl-CoA to chloramphenicol, thereby preventing antibiotic binding to the 50S subunit of the bacterial ribosome.4 In some resistant strains, carbapenemases are particularly concerning because they can compromise carbapenems, which are often reserved for severe infections caused by multidrug-resistant Gram-negative bacteria.3,4
Efflux pumps
Another mechanism adopted by ESKAPE pathogens is the use of membrane transporters that act as efflux pumps to expel antimicrobial agents from the bacterial cell, necessitating higher antibiotic concentrations to achieve clinically significant outcomes.2,3 In Pseudomonas aeruginosa, for example, efflux systems such as MexAB-OprM and MexCD-OprJ contribute to resistance against multiple antibiotic classes, including aminoglycosides, carbapenems, and fluoroquinolones.4
To date, six families of efflux pumps have been identified based on their energy requirements and structure, including:
- Adenosine triphosphate (ATP)-binding cassette (ABC) family
- Multidrug and toxin extrusion (MATE) group
- Major facilitator superfamily (MFS)
- Resistance-nodulation-cell division (RND) superfamily
- Small multidrug resistance (SMR) family
- Proteobacterial antimicrobial compound efflux (PACE) family
ESKAPE Pathogens: The Invisible Threat
Challenges in treatment and control
The treatment and control of ESKAPE pathogens present significant challenges due to both biological complexity and systemic limitations in healthcare. One major concern is the lack of research dedicated to discovering new antibiotics, despite the gradual decline in the efficacy of existing drugs against AMR species. Only a limited number of new antibiotic classes have reached clinical use in recent decades, increasing reliance on older or last-line agents.3
Carbapenems and cephalosporins were previously highly effective against a broad range of resistant bacterial infections. However, the emergence of resistance genes has similarly rendered these antibiotics ineffective against some rapidly evolving species. Last-line or alternative agents such as colistin, tigecycline, linezolid, daptomycin, and newer combination approaches may be used in selected infections, but toxicity, resistance emergence, pathogen specificity, and limited clinical options remain major barriers.3,4
The rapid evolution of antibiotic resistance in these pathogens has outpaced the development of many new therapies, leaving healthcare providers with increasingly limited treatment options.3,4
The rapid and accurate diagnosis of ESKAPE infections remains difficult, delaying the initiation of targeted therapy and leading to the use of broad-spectrum antibiotics that can contribute to resistance. The combination of bioinformatics with advanced technologies like next-generation sequencing (NGS) enables the accurate detection and timely characterization of resistant pathogens; however, these methods are more expensive than routine clinical tests.1,3
Addressing these challenges requires strong antimicrobial stewardship programs to optimize antibiotic use and reduce unnecessary exposure. Effective infection prevention and control measures, including hygiene practices and surveillance, are also essential to limit the spread of these high-risk pathogens within healthcare settings.1,3 From a One Health perspective, surveillance should also extend beyond hospitals to food production, animal husbandry, water, and other environmental reservoirs that can contribute to the spread of resistant bacteria and resistance genes.4
Emerging strategies to combat ESKAPE pathogens
Despite insufficient investment into the research and development of novel antibiotics, several interesting therapeutics are currently being studied for their potential to protect against ESKAPE pathogens. These include anti-virulence molecules, photodynamic light therapy, bacteriophage therapy, anti-sense oligonucleotide (ASO) therapeutics, antimicrobial peptides (AMPs), vaccines, combination therapies, and repurposed drugs.4 Other, mostly investigational, approaches include adjuvants, nanoparticles, CRISPR-Cas-based antimicrobials, live biotherapeutics, improved diagnostics, and machine-learning-guided discovery or surveillance.3
Bacteriophages, for example, are attractive therapeutics due to their ability to disrupt biofilm formation. However, the practical and large-scale implementation of this approach is associated with numerous limitations, including the inherent capacity of bacteria to rapidly acquire resistance to phage infection.3 Because ESKAPE pathogens use quorum sensing to coordinate virulence and biofilm-related behaviors, quorum-sensing inhibitors are also being explored as anti-virulence strategies that may weaken pathogenicity without directly imposing the same selective pressure as conventional bactericidal drugs.2
Advances in genomic technologies are enabling more effective surveillance and tracking of resistant strains, thereby allowing early detection of outbreaks and improved infection control measures. Together, these strategies represent a multifaceted approach to addressing the growing threat posed by ESKAPE pathogens.1,3 Animal infection models, including mice, zebrafish, Caenorhabditis elegans, Galleria mellonella, pigs, rabbits, and silkworm larvae, remain important for studying pathogenesis, host responses, and candidate therapeutics, although no single model fully reproduces human ESKAPE infections.1
References
- Yu, H., Xu, Y., Imani, S., et al. (2024). Navigating ESKAPE pathogens: considerations and caveats for the development of animal infection models. ACS Infectious Diseases 10(7); 2336-2355. DOI: 10.1021/acsinfecdis.4c00007. https://pubs.acs.org/doi/10.1021/acsinfecdis.4c00007
- Venkateswaran, P., Vasudevan, S., David, H., et al. (2023). Revisiting ESKAPE Pathogens: virulence, resistance, and combating strategies focusing on quorum sensing. Frontiers in Cellular and Infection Microbiology 13. DOI: 10.3389/fcimb.2023.1159798. https://www.frontiersin.org/journals/cellular-and-infection-microbiology/articles/10.3389/fcimb.2023.1159798/full
- Singh, A., Tanwar, M., Singh, T. P., et al. (2024). An escape from ESKAPE pathogens: A comprehensive review on current and emerging therapeutics against antibiotic resistance. International Journal of Biological Macromolecules 279. DOI: 10.1016/j.ijbiomac.2024.135253. https://www.sciencedirect.com/science/article/abs/pii/S0141813024060598
- Oyenuga, N., Cobo-Diaz, J. F., Alvarez-Ordoñez, A., & Alexa, E. (2024). Overview of Antimicrobial Resistant ESKAPEE Pathogens in Food Sources and Their Implications from a One Health Perspective. Microorganisms 12(10); 2084. DOI: 10.3390/microorganisms12102084. https://www.mdpi.com/2076-2607/12/10/2084.
Further Reading
Last Updated: Apr 30, 2026