Host cells infected with giant viruses behave in a unique manner. To gain deeper insight into the infection mechanism of giant viruses, scientists at Tokyo University of Science developed a specialized algorithm that can track the movement of host cells. This method could also be used to study any other type of cells, such as cancer cells, neurons, and immune cells, serving as an efficient tool in the field of cell biology.
Viruses are probably the most peculiar “life forms” known to us, with each species having unique characteristics. One thing common to all viruses is that they attack a “host” cell and hijack its machinery, using it for its own replication. A type of large virus, called “giant virus,” has interesting ways of attacking its host organism—an amoeba. Virologists have been trying to understand what makes these viruses so unique for some time now, but to observe them, complex techniques are required.
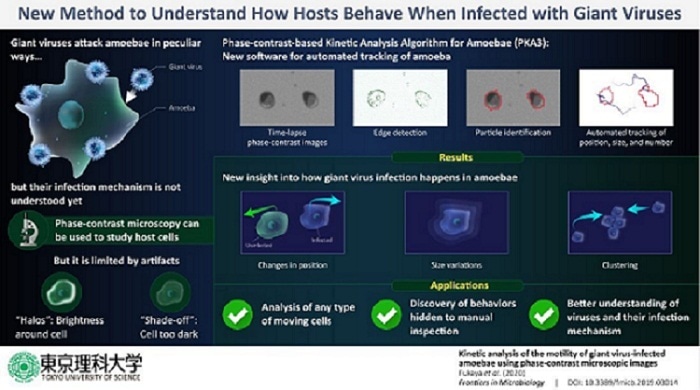
A method called phase-contrast microscopy is commonly used to study all types of cells, including amoebal cells. But, this technique relies heavily on the variation in the conditions of the cell and its background, and sometimes this leads to disruptions in the image produced—for example “halo” (in which the cell is surrounded by a bright light) and “shade-off” (in which the inside of the cell and background are of a similar intensity). This is why, to dig deeper into exactly how giant viruses infect amoebal cells, more efficient cell-tracking techniques are needed.
Just earlier this month, scientists from the Tokyo University of Science, led by Professor Masaharu Takemura, had reported the discovery of two new species of pandoravirus and mimivirus—both families of giant viruses infecting amoebae—from a riverbank in Japan. Prof Takemura says that continual discovery of viruses from soil is crucial from the standpoint of understanding the ecology of giant viruses.
More importantly, in a brand new study published in Frontiers in Microbiology, a team of scientists at Tokyo University of Science, once again led by Prof Takemura, tried to understand the behavior in which amoebal cells infected with different types of giant viruses. For this, they devised a new cell-tracking method that tackles the issues of conventional analysis methods. Prof Takemura elaborates on the motivation of the study, “Our aim was to understand how giant viruses infect amoebae in the natural environment and how this has affected to evolution of eukaryotes. For this, we wanted to develop a technique to quantitatively detect the time-dependent, sequential changes in cellular number, size, shape, and direction and distance of cell motility.”
In their study, Prof Takemura and his team focused on amoebae that were infected by a family of giant virus called “marseillevirus”. To understand the behavior of host cells infected with this particular type of virus, the scientists developed a new algorithm that can trace individual amoebal cells in a population of amoebae using time-lapse phase-contrast microscopic images. They named this algorithm as “Phase-contrast-based Kinetic Analysis Algorithm for Amoebae” or PKA3. Using PKA3, the scientists revealed new, interesting aspects of how amoebae react to a giant virus attack. For example, they quantitatively showed that giant virus-infected amoebal cells formed aggregates or “bunches.” They inferred that this could be either an antiviral strategy of the host or how the virus spreads, shedding light on the manner in which the viral infection takes place. What’s more, they managed to detect changes in the number of cells and the appearance of marseillevirus-infected amoebae much faster than by conventional methods. Interestingly, they could also analyze the time taken by amoebae to respond to a viral infection, providing useful information on the life cycle of amoebae and viruses and the relationship between them.
Prof Takemura says:
Our algorithm was successful in visualizing the movement of cells in images produced by phase-contrast microscopy, a method widely used in cell biology. Also, it allowed quantification of various parameters. This research will greatly contribute to the demonstration of amoeba behavior infected with the giant virus.”
Apart from studying giant viruses, this new algorithm could be used for various other applications, such as to study the dynamics of cancer cells, lymphocytes, macrophages, and neurons. In general, it could reveal new cellular phenomena by tracking exactly how these cells migrate.
Prof Takemura concludes by saying:
Our new analytical method could be applied to all cells that can be observed with a phase-contrast microscope and can potentially be applied to various fields including cell biology, medicine, and biotechnology.”
Indeed, a technique like this—which can potentially make cell biology research easier—was much needed!
Source:
Journal reference:
Fukaya, S., et al. (2020) Kinetic Analysis of the Motility of Giant Virus-Infected Amoebae Using Phase-Contrast Microscopic Images. Frontiers in Microbiology. doi.org/10.3389/fmicb.2019.03014.