The 2019 novel coronavirus (CoV) causes fatal pneumonia that has claimed over 1300 lives, with more than 52000 confirmed cases of infection by February 13, 2020, all in the span of just over a month. But, what is this virus? Is it a new virus altogether? Where did it come from? Scientists from top research institutes in China teamed up to answer these questions, and this pioneering study has been published in Chinese Medical Journal.

In early December, a few people in the city of Wuhan in the Hubei province of China began falling sick after going to a local seafood market. They experienced symptoms like cough, fever, and shortness of breath, and even complications related to acute respiratory distress syndrome (ARDS). The immediate diagnosis was pneumonia, but the exact cause was unexplained. What caused this new outbreak? Is it the severe acute respiratory syndrome (SARS)-CoV? Is it the Middle East respiratory syndrome (MERS)-CoV? As it turns out, scientists had undertaken a study to identify this virus in December after analyzing the first few cases. This study is now published in Chinese Medical Journal and the identity of the virus has been established--it is a completely new virus, closely related to the bat SARS-like CoV.
Dr. Jianwei Wang (Chinese Academy of Medical Sciences, Institute of Pathogen Biology), lead researcher on the study, states:
Our paper has established the identity of the bat-origin CoV that was unknown until now."
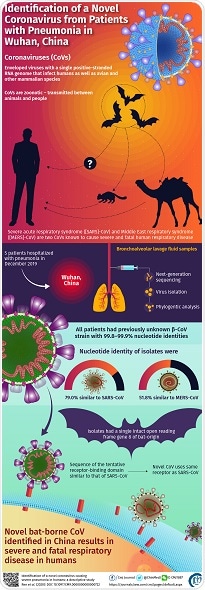
In this study, scientists from renowned research institutes in China, such as the Chinese Academy of Medical Sciences, Institute of Pathogen Biology, China-Japan Friendship Hospital, and Peking Union Medical College, jointly discovered and identified the new CoV--the main culprit of the Wuhan outbreak--by next-generation sequencing (NGS). They focused on five patients admitted to Jin Yin-tan Hospital in Wuhan, most of whom were workers in the Huanan Seafood Market in Wuhan. These patients had high fever, cough, and other symptoms, and were initially diagnosed to have pneumonia, but of an unknown cause. Some patients' condition rapidly worsened to ARDS; one even died.
Dr. Wang says:
Chest x-rays of the patients showed some hazy opacities and consolidations, which are typical of pneumonia. However, we wanted to find out what caused the pneumonia, and our subsequent experiments revealed the exact cause--a new CoV that was not known before."
For the study, the scientists used bronchoalveolar lavage (BAL) fluid samples taken from the patients (BAL is a procedure in which sterile fluid is transferred to the lungs through a bronchoscope and then collected for analysis).
First, the scientists attempted to identify the virus by genome sequencing, using NGS technology. NGS is the preferred screening method for identifying unknown pathogens because it quickly detects and rules out all known pathogenic microorganisms in the sample. Based on sequencing of the DNA/RNA from the BAL fluid samples, the scientists found that most of the viral reads belonged to the CoV family. The scientists then assembled the different "reads" that belonged to CoVs and constructed a whole genomic sequence for the new virus; these sequences were 99.8-99.9% similar among all the patients' samples, confirming that this virus was the common pathogen in all the patients. Further, using homology analysis, where a genome sequence is compared against other known genome sequences (with a preset threshold of 90% for it to be considered a "new" sequence), they confirmed that the genome sequence of this new virus is 79.0% similar to the SARS-CoV, about 51.8% similar to the MERS-CoV, and about 87.6-87.7% similar to other SARS-like CoVs from Chinese horseshoe bats (called ZC45 and ZXC21). Phylogenetic analysis showed that the sequences of the five CoV strains obtained were closest to those of bat-derived strains, but formed separate evolutionary branches. These findings clearly suggest that the virus originated from bats.
Dr. Wang states:
Because the similarities of the viral replicase gene with all other known "similar" viruses are still less than 90%, and also taking into account the phylogenetic analysis results, we consider that this is indeed a new, previously unknown CoV. This new virus is temporarily called the 2019-nCoV."
Lastly, the scientists moved to "isolating" the virus from the BAL fluid samples by checking whether the fluid samples showed cytopathic effect to cell lines in the laboratory. The cells exposed to the fluid samples were observed under an electron microscope, and the scientists found characteristic CoV-like structures. They also used immunofluorescence--a technique that uses specific antibodies tagged with fluorescent dyes. For this, they used serum from the recovering patients (which contained antibodies), which reacted with the viral particles inside the cells; this confirmed that this virus was indeed the cause of the infection.
This study paves the way for future studies to understand the virus and its sources better, especially given its rapid spread, its ability to cause fatal ARDS, and the panic caused by the outbreak. Although 4 of the 5 patients from whom this virus was identified were from a seafood market in Wuhan, the exact origin of infection is unknown. The CoV could have been transmitted to humans through an "intermediate" carrier, such as in the case of SARS-CoV (palm civet meat) or MERS-CoV (camel).
Dr. Wang concludes:
All human CoVs are zoonotic, and several human CoVs have originated from bats, including the SARS- and MERS-CoVs. Our study clearly shows the urgent need for regular monitoring of the transmission of bat-origin CoVs to humans. The emergence of this virus is a massive threat to public health, and therefore, it is of critical importance to understand the source of this virus and decide the next steps before we witness a larger scale outbreak."
Identification of a Novel Coronavirus Causing Severe Pneumonia in Humans: A Descriptive Study
This pioneering study undertakes the sequencing of the CoV's genome for the first time. Credit: Chinese Medical Journal
Source:
Journal reference:
Ren, L-L., et al. (2020) Identification of a novel coronavirus causing severe pneumonia in human a descriptive study. Chinese Medical Journal. doi.org/10.1097/CM9.0000000000000722.