A new study shows the role that viral genomic sequencing can play in the current COVID-19 pandemic.
The virus severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic has spread rapidly across the globe, infecting over 2.6 million and killing over 183,000. The only intervention currently possible is in the form of non-pharmacological measures, chiefly contact tracing, quarantine measures, social distancing, and lockdowns.
How well do these work? In the absence of universal testing, it is difficult to tell because many cases are asymptomatic. This is where sequencing can play a significant role.
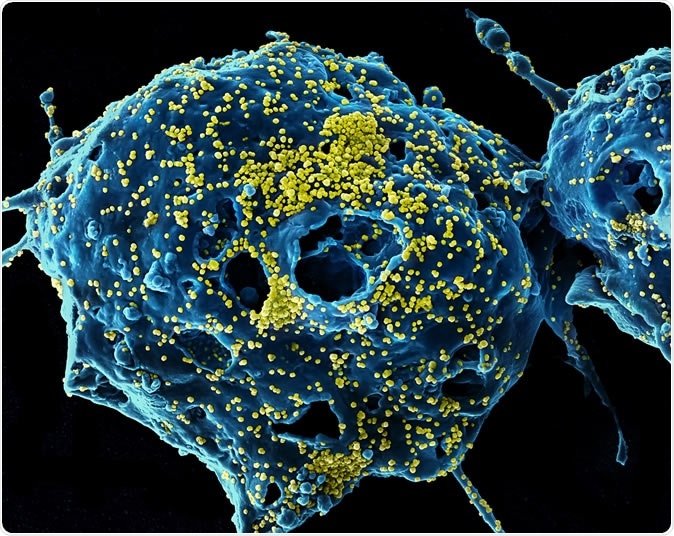
MERS Virus Particles Colorized scanning electron micrograph of Middle East Respiratory Syndrome virus particles (yellow) attached to the surface of an infected VERO E6 cell (blue). Image captured and color-enhanced at the NIAID Integrated Research Facility in Fort Detrick, Maryland. Credit: NIAID
The beginning of genomic surveillance
In China, scientists rapidly sequenced the viral genome and identified it as a betacoronavirus similar to the virus that caused the 2002 SARS epidemic. Throughout the spread of the virus, genomic sequencing has played a role in epidemiologic tracking of the viral strains carried all over the world. Such strains are traced back to their origin by their unique mutations and help to link remote infections.
This method has been immensely moved forward by the development of targeted sequencing protocols, the GISAID (Global Initiative on Sharing All Influenza Data) repository, which allows open real-time sharing of sequences, and Nextstrain. This analytical platform allows the mutations in a given viral strain to be traced speedily. All these have been put to good use in visualizing the likely paths of the spread of the virus.
Some unexpected findings include the finding of a genetic sequence in a patient in the Seattle area, without a history of travel to a hotspot or contact with a COVID-19 case, which was related to that found in a traveler 5 weeks before – showing that community spread had been occurring extensively though silently all that time.
Computer monitor shows DNA sequencing. D-VISIONS / Shutterstock
New York – the new epicenter
New York has witnessed an inferno of disease and death ever since the first confirmed case on March 3, 2020. This solitary case exploded into over 250,000 cases in New York State alone, which is just short of one-tenth of the total cases in the world.
However, New York City alone accounts for over half the total cases in the state with over 142,000 confirmed cases. At present, the epicenter of the NY outbreak feeds the NYU Langone Health system of hospitals. The current study aimed at recording the features of early COVID-19 spread in a large city, and to trace the strains back to their earliest known origins.
How was the study done?
The researchers wanted to see how the virus was spreading within the catchment area of the NYU Langone Health network of hospitals in Brooklyn, Manhattan, and Nassau County. They first set up an optimally designed workflow for the genomic sequencing and analysis. They randomly sampled a population of confirmed cases tested between March 12 and April 1, 2020.
The RNA was sequenced, and a robotic library was constructed. Shotgun sequencing methods were used to generate high-quality sequences directly. Analysis of the viral genomes was done with 156 sequences that passed al the quality tests. The medical records were analyzed to identify exposures.
What did the study show?
The researchers found that the samples came from throughout the area mentioned above, but most were from Brooklyn and Manhattan. Most of these were located in the catchment area of the hospitals in the New York metropolitan area.
However, one region, Westchester County, located to the north of the city, and outside the area served by the hospital, was not represented, though it saw the first regional outbreak.
Over 50% of cases had no recorded history of exposure. The researchers, therefore, carried out sequencing and phylogenetic analysis of the sample sequences to find out how closely they were related to each other. If cases spread from one or a few index cases, the viruses should be almost identical.
They colored each sequence according to where the patient lived. This showed that there were multiple strains of the coronavirus circulating within the region of sampling dating from the first week in March.
They then compared the genomes to over 7,600 globally collected sequences using the GISAID EpiCov repository. After tracing the phylogeny, they colored each sequence according to the most similar sequence from another region. Surprisingly, the researchers found that over 41% of the samples were most closely related to Europe, while 46% were from the US or Canada.
When they looked at the date of collection of the latest common strain, stratified by region, they found that approximately 66% were similar to European samples from as early as the last week of February 2020.
Little data is available regarding the early spread of the pandemic in January and February particularly, because few sequences have been collected from the route of transmission. As a result, say the investigators, finding a link to European samples alone doesn’t mean that they suspect a specific transmission event or have found a timeline.
Overall, they found almost 190 nucleotide variants and 97 altered amino acids. More continue to be found as more sample sequencing is performed. This means that surveillance must be persistent, at local, regional, and national levels and even international, if the pandemic is to be adequately monitored.
How is the study important?
Mutations in the influenza virus that causes seasonal flu are essential for virulence, enabling the virus to escape the immune system even after vaccination. It also enables the virus to develop resistance to the antiviral drug oseltamivir. However, this is not the case with most of the mutations in the SARS-CoV-2 virus, which are expected to be nonfunctional. These are the result of genetic drift, say the researchers.
Again, genomic tracing can help track viral spread independently of medical history. This is how the current sampling from the NY area showed a much broader spectrum of transmission had occurred than was thought to be the case back in Seattle. This kind of analysis may need to be done retrospectively, to find out how the spread is occurring invisibly in the community within each region.
This will also throw light on the effects of public health policies, and behavioral changes such as social distancing and quarantine. It will also help to shape the prospective management of current outbreaks.
But such surveillance is not easy to set up instantaneously. Thus, in view of these critical functions, the scientists say, “Given the logistical and regulatory hurdles to establishing such surveillance, it is critical to have this infrastructure already in place for future waves of COVID-19.”
Journal reference:
Maurano, M. T., Ramaswami, S., Westby, G., Zappile, P., et al. (2020). Sequencing analysis of the spread of SARS-CoV2 in the Greater New York City Region. medRxiv preprint doi: https://doi.org/10.1101/2020.04.15.20064931. https://www.medrxiv.org/content/10.1101/2020.04.15.20064931v1