The COVID-19 pandemic has spread throughout the whole world within six months, causing millions of infections and hundreds of thousands of deaths. Yet there is no effective vaccine or antiviral to deal with it.
A new study by researchers at the Fred Hutch Cancer Research Center and the University of Washington and published on the preprint server bioRxiv* in June 2020 reports on the interactive visualization and open analysis pipeline connecting mutations of the viral receptor-binding domain (RBD) with changes in the viral biochemical phenotype. This could be a first step in monitoring the genetic diversity of the virus over time.
Genotype-Phenotype Linkage
Thousands of genomes have been sequenced from all over the world, and are available in international genomic databases. These have proved invaluable in interpreting the evolution of the virus and its spread patterns. However, there is a severe lack of understanding when it comes to linking the variations in the genomes with corresponding changes in the phenotype.
SARS-CoV-2 is a betacoronavirus, and as such, binds to host cell ACE2 receptors through the viral spike protein, which bears a high-affinity receptor-binding domain (RBD). The RBD is, therefore, essential for viral transmission across species and adaptation. It is also the target of neutralizing antibodies, with many vaccines being newly developed based on this protein alone.
Mutations in the RBD
The RBD is among the genomic regions with the highest variability, because of the many selection pressures that cause different mutations to become prevalent. Many such mutations have already become apparent, but little is known about how these affect receptor binding.
The current study seeks to map every amino acid change in the RBD with its effect on the way the resulting protein is folded and the affinity of the protein for ACE2, which are both vital factors in determining viral fitness. The researchers used a yeast-display platform to evaluate the expression of the folded RBD protein as well as its binding to the ACE2 receptor. The advantage of using yeast cells is the occurrence of N-glycans at the same locations on the RBD as in humans.
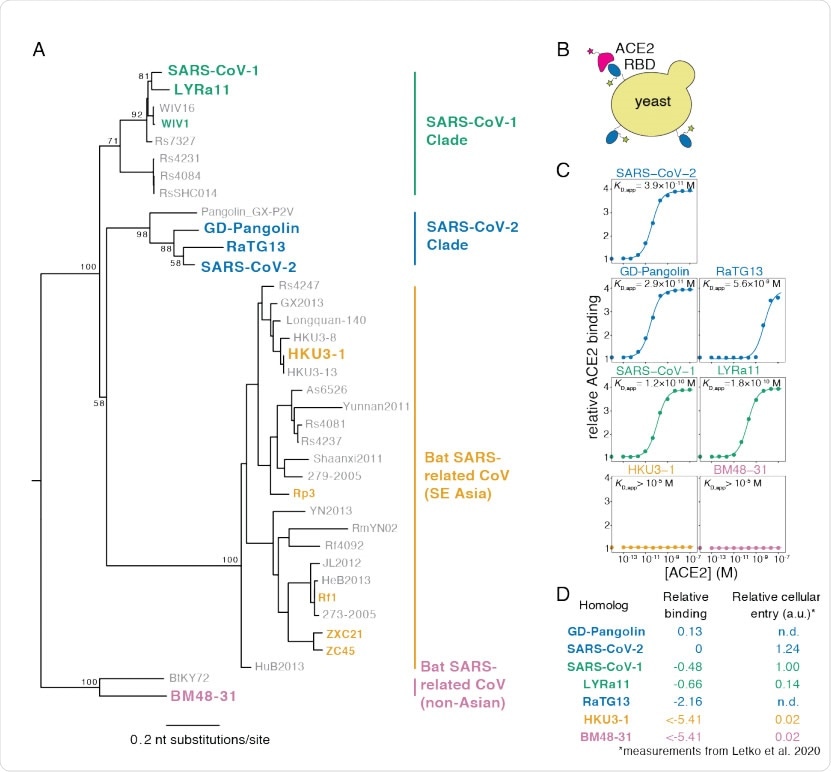
Yeast display of RBDs from SARS-CoV-2 and related sarbecoviruses. (A) Maximum likelihood phylogenetic tree of sarbecovirus RBDs. RBDs included in the present study are in bold text colored by RBD clade. Node labels indicate bootstrap support. (B) RBD yeast surface display enables fluorescent detection of RBD surface expression and ACE2 binding. (C) Yeast displaying the indicated RBD were incubated with varying concentrations of human ACE2, and binding was measured via flow cytometry. Binding constants are reported as KD,app from the illustrated titration curve fits. (D) Comparison of yeast display binding with previous measurements of the capacity of viral particles to enter ACE2-expressing cells. Relative binding is Δlog10(KD,app) measured in the current study; relative cellular entry is infection of ACE2-expressing cells by VSV pseudotyped with spike containing the indicated RBD, reported by Letko et al. (Letko et al., 2020) in arbitrary luciferase units relative to SARS-CoV-1 RBD; n.d. indicates not determined by Letko et al.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Mutational Tolerance
These maps show that the RBD is mutationally tolerant, but there is a tremendous difference in the degree of mutational constraint across the regions. Most mutations of the RBD are either neutral or have a slightly harmful effect on either or both of these biochemical expressions. There is a single mutation, V367F, which increases the expression of the folded protein. However, multiple mutations boost affinity at the RBD sites.
The map also shows that ACE2 binding positions can tolerate many mutations even though many residues that are considered essential for virus-ACE2 binding are different. Such mutations Despite the emergence of several mutations, none have shown a notable tendency to be selected over any other even if it produces a marked increase in ACE2 binding affinity.
Limitations of the Study
The current study maps only the biochemical phenotypes and not viral fitness. This distinction is important because the RBD protein displayed on yeast cells is not directly related to viral fitness for several reasons. For one, the glycan structures on yeast and human cells are different even though they have the same function of stabilizing the RBD. Secondly, the viral spike protein comprises several components, of which the RBD is one. And thirdly, viral entry via the spike protein is only a single part of viral fitness, since multiple interrelated factors affect the efficiency of viral spread.
It is also true that despite the limitations of basic biochemical research like the current study, the study does show a good correlation with the results of viral entry studies that use pseudotyped viruses carrying the spike protein with sarbecovirus RBD homologs or single mutations of the SARS-CoV-2 RBD. Another advantage is that fitness is the result of a combination of the activity of multiple biochemical phenotypes, which are the expression of the underlying genotype. Thus, studies like this one provide an initial movement towards understanding how mutation affects viral function.
Examining Antigenic Drift Potential
One especially relevant area of the current study is its ability to understand the chances of antigenic drift with the SARS-CoV-2 by fixing the mutations at the sites used for antibody binding. This is because the RBD is a prime target for neutralizing antibody production, which means that any antigenic drift is constrained, or limited, by the fact that its function must be preserved or enhanced.
The researchers found that most RBD mutations allow both protein folding and ACE2 binding to occur normally. The ACE2 binding site is constrained still more tightly than the RBD site, and therefore antibodies to the antigens at this interface are more challenging to escape from via antigenic change.
As a result of these observations, the scientists suggest that among all the antibodies that have been studied in detail so far, none have comparably constrained epitopes as the RBD surface that attaches to the ACE2 molecule. Thus, it should be possible to focus on epitopes that cannot be changed without decreasing viral fitness.
Secondly, many mutations of the RBD increase ACE2 binding affinity, which could mean that many amino acid substitutions can overcome the effects of harmful mutations in this region. This is supported by the existence of escape mutations occurring in multiple steps in other viruses. As the method is used to achieve a direct map of immune-escape mutations, this will increase the state of current knowledge about the potential for antigenic drift.
Applications of the Mutational Map
Many teams are striving to produce vaccines targeting the RBD antigens. In this approach, too, the current genotype-phenotype mapping of the RBD can shape the efforts to create such vaccines. The first step would be to make use of the mutations identified in the current study that increase RBD expression, which makes the vaccine more potent. Secondly, the maps can help select the mutations that are safe for the RBD in terms of preserving its biochemical function, which could lead to engineering the required antigen so as to encourage the production of antibodies targeting specific epitopes.
A final area of help is to understand the most constrained areas of the RBD, to target these areas for the production of neutralizing antibodies in all sarbecoviruses, using structure-guided vaccines.
The study also has theoretical benefits, advancing the understanding of how the sarbecoviruses are adapting and spilling over into the human host. The vast array of RBD sequences and phenotypes must be narrowed down to those which can bind human receptors efficiently. This study thus shows, through the mapping, how SARS-CoV-2 mutations affect ACE2 binding and could be extended to understand further how these viruses mutate to become human viruses.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources