Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the causative agent of coronavirus disease 2019 (COVID-19). SARS-CoV-2 originated in Wuhan, Hubei province, China, in late 2019 and spread to over 200 nations causing the current pandemic that has resulted in the most significant global health crisis in one hundred years.
While many infected people experience only mild symptoms, individuals with comorbidities such as cardiovascular diseases or diabetes, and the elderly are more likely to have severe COVID-19 and need hospital care.
Genetic variations in the patients may explain why some people are prone to severe COVID-19 outcomes, while others promptly recover from mild symptoms. This makes it clear that a better understanding of the role of genetic variants is critical to improving our knowledge of COVID-19 pathogenesis.
However, the role played by genetics in COVID-19 outcomes is poorly understood. Despite extensive research efforts by scientists worldwide, the genes and pathways that drive the disease are still unclear.
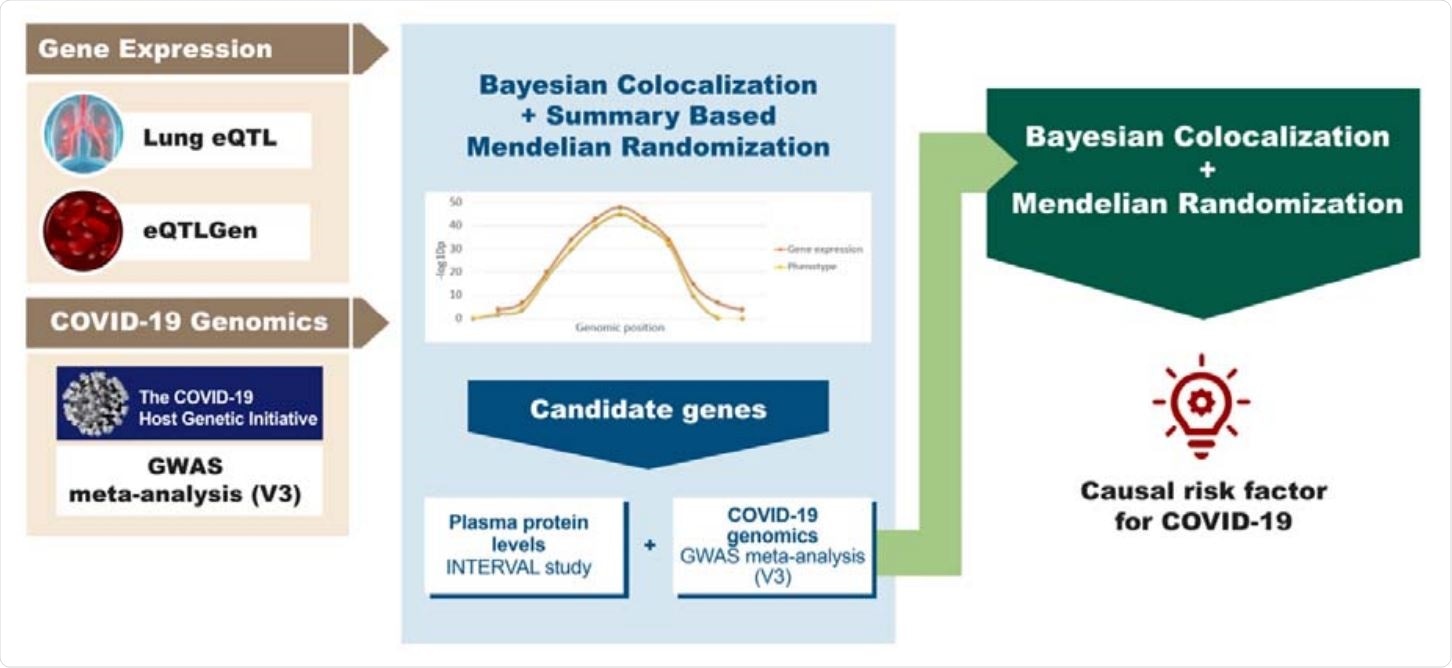
Study overview. The diagram summarizes the genomics datasets and analytic pipeline of the study. First, publicly available -omics datasets were obtained, which were later processed using integrative genomics (IG) methods (Bayesian Colocalization and Summary-based Mendelian Randomization) to identify potential candidate genes for COVID-19 phenotypes. Lastly, using a Bayesian Colocalization and Mendelian Randomization approach we explored the causal association between the plasma protein levels of the most promising candidate gene and COVID-19.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Studies exploring the genes associated with COVID-19 susceptibility
A recent pooled genome-wide association study showed a susceptibility loci, including 3p21.31 and 9q34.2, for severe COVID-19 leading to respiratory failure. However, these loci comprised several genes, and the specific genes associated with COVID-19 were not explored in the study.
Recently, researchers from the University of British Columbia, University of Sydney, University of Groningen, Merck Research Laboratories, Université Laval, Québec City, and Icahn School of Medicine at Mount Sinai, NY, used an integrative genomics approach to find the genes responsible for COVID-19 and its severity. Their work is published on the preprint server, medRxiv.*
The team used Bayesian colocalization (COLOC) and summary-based Mendelian randomization to combine gene expression quantitative trait loci (eQTLs) from the lung eQTL (n=1,038) and eQTLGen (n=31,784) studies using published data from the COVID-19 genome-wide association study by the COVID-19 Host Genetics Initiative.
The researchers also used COLOC to integrate plasma protein quantitative trait loci (pQTL) from the INTERVAL study (n=3,301) with COVID-19-linked loci. Lastly, they determined causal associations between COVID-19 and plasma proteins with the help of multi-variable, 2-sample Mendelian randomization.
Gene expression in blood and lung tissues linked to COVID-19
The team found that the expression of 31 genes in blood and 20 genes in the lung was associated with COVID-19. Out of these genes, only 3 – LZTFL1, SLC6A20, and ABO – were previously linked to COVID-19 in GWAS.
The newly found loci included genes involved in interferon pathways - IL10RB, IFNAR2, and OAS1. More importantly, plasma ABO, a protein associated with human blood type, was shown to have a significant relationship with COVID-19 in MR analysis. An increase in plasma levels was associated with an increased COVID-19 risk as well as the risk of severe COVID-19.
“Although the chromosome 3p21 region has been previously associated with severe COVID-19, this locus encompasses six genes, making it hard to identify the precise gene responsible for the association with COVID-19.”
Using a multi-omics approach, the team identified many candidate genes that may be involved in COVID-19 pathogenesis. Their study revealed close links between COVID-19 genomics and gene expression in blood and lung tissues, and their approach identified specific genes of the COVID-19 loci implicated in previous studies, and also found new genes associated with severe COVID-19 risk.
According to the team, their study identified several genes associated with COVID-19 that can be prioritized by scientists for future research work. They also claim that this is the first study demonstrating a causal association between plasma ABO protein and severe COVID-19 risk.
“Importantly, our analysis suggests that the ABO protein is a causal risk factor for severe COVID-19 and COVID-19 susceptibility.”
Limitations of the study
This study has several limitations. The analysis of the research group was limited by the number of genetic variants that overlapped between the datasets they used and the COVID-19 HG meta-analysis, which means they might not have tested some genes of importance.
The lung and blood genes data used in the study show the transcriptomic and proteomic profile under normal conditions. Still, these associations may change during actual viral infection or acute inflammation.
Also, the team could not replicate their study results in an independent dataset because the COVID-19 GWAS data was not available at the time. So first-time associations identified by their analyses need to be considered with caution.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Multi-omics highlights ABO plasma protein as a causal risk factor for COVID-19 Ana I Hernandez Cordero, Xuan Li, Stephen Milne, Chen Xi Yang, Yohan Bosse, Philippe Joubert, Wim Timens, Maarten van den Berge, David Nickle, Ke Hao, Don D Sin medRxiv 2020.10.05.20207118; doi: https://doi.org/10.1101/2020.10.05.20207118, https://www.medrxiv.org/content/10.1101/2020.10.05.20207118v1
- Peer reviewed and published scientific report.
Hernández Cordero, Ana I., Xuan Li, Stephen Milne, Chen Xi Yang, Yohan Bossé, Philippe Joubert, Wim Timens, et al. 2021. “Multi-Omics Highlights ABO Plasma Protein as a Causal Risk Factor for COVID-19.” Human Genetics 140 (6): 969–79. https://doi.org/10.1007/s00439-021-02264-5. https://link.springer.com/article/10.1007/s00439-021-02264-5.