The study findings indicate that endothelial cells are resistant to SARS-CoV-2 infection and that vascular complications observed in COVID-19 patients may be caused by systemic pro-inflammatory mediators and other cellular factors. The study is currently available on the bioRxiv* preprint server.
To curb the ever-growing COVID-19 trajectory, the development of effective therapeutics or vaccines has become the need of the hour. Although several potential vaccines are in the pipeline, it is still uncertain if these vaccines can provide long-term protection. Medicines that are currently in use to treat COVID-19 patients are mostly repurposed drugs that are known to target viral replication in general. Therefore, a proper understanding of the disease’s pathophysiology is important for developing therapeutics that can specifically target SARS-CoV-2 and ameliorate disease outcomes in those infected with SARS-CoV-2.
Primarily, SARS-CoV-2 infection occurs in the respiratory epithelium, where the interaction between viral spike protein and host cell angiotensin-converting enzyme 2 (ACE2) receptor leads to viral entry. An alternative pathway may also exist, wherein SARS-CoV-2 enters host cells via a transmembrane glycoprotein called basigin (BSG). Some recent evidence suggests that people with severe COVID-19 develop persistent cardiovascular complications, including coagulopathy or bleeding disorder, even after recovery. This indicates that SARS-CoV-2 may affect the endothelium lining of blood vessels, in addition to respiratory epithelium.
The current study has been carried out to thoroughly investigate the ability of SARS-CoV-2 to infect endothelial cells.
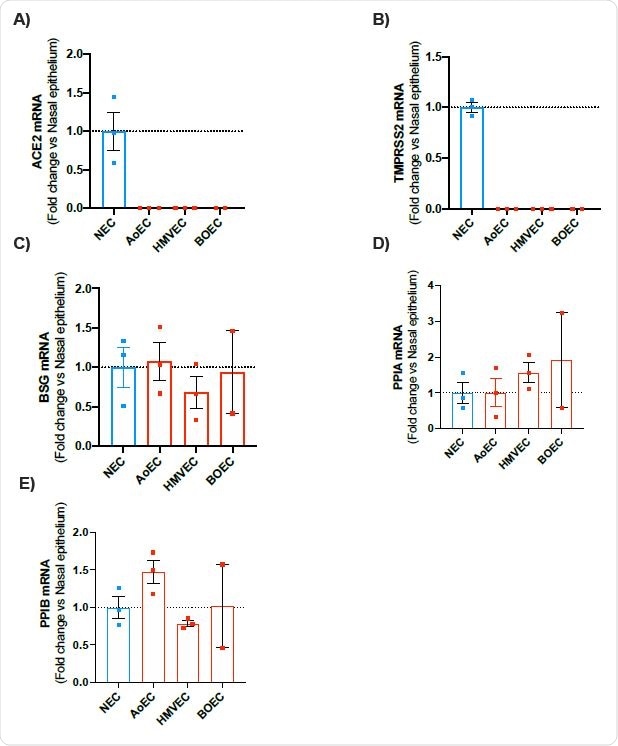
mRNA expression of ACE2, TMPRSS2, BSG, PPIA and PPIB in human nasal epithelial cells (NEC) and endothelial cells (aortic, microvascular and blood outgrowth). Expression levels for the genes ACE2, TMPRSS2, BSG, PPIA and PPIB were obtained from aortic (AoEC), microvascular (HMVEC) and blood outgrowth (BOEC) endothelial cells and nasal epithelial cells (NEC). Data for each donor were normalized using the average of the housekeepers (18S and Gapdh) and analyzed using a comparative Ct method (2DDCt). Data are shown as the mean +/- SEM fold change compared to nasal epithelium (n=3 wells using cells from 2 donors) for AoEC (n=3 wells using cells of 3 separate donors), HMVEC ((n=3 wells using cells of 3 separate donors and BOECs (n=2 wells using cells of 2 separate donors).
Current study design
The scientists conducted a series of in vitro experiments using primary endothelial cell lines that were infected with either live SARS-CoV-2 or a pseudovirus expressing the viral spike protein. They used three types of cell lines, including blood outgrowth endothelial cells, lung microvascular endothelial cells, and aortic endothelial cells. They chose three cell lines, including nasal epithelial cells, primate kidney fibroblast cells, and human kidney embryonic cells, as positive controls to compare with endothelial cell findings.
Important observations
The scientists observed that the expressions of ACE2 and TMPRSS2 (a cell surface protease required for viral entry) were significantly lower in endothelial cells compared to that in nasal epithelial cells. However, they noticed a comparable expression of proteins of the alternative viral entry pathway, such as BSG, PPIA, and PPIB, in endothelial and epithelial cells. These findings suggest an ACE2-independent pathway (BSG pathway) may be responsible for SARS-CoV-2 entry into endothelial cells.
By incubating both endothelial cells and primate kidney fibroblast cells with live SARS-CoV-2, they observed that the virus successfully entered the fibroblasts cells and started replicating. However, they were unable to detect SARS-CoV-2 infection in endothelial cells. To check if endothelial cells are resistant to SARS-CoV-2 infection at the viral entry-level or viral replication level, they infected the cells with a pseudovirus expressing the spike protein. Similar to previous findings, they could not detect spike protein-mediated entry of SARS-CoV-2 into endothelial cells. These findings suggest that endothelial cells are not susceptible to SARS-CoV-2 infection even with a high level of BSG expression.
To further verify the involvement of ACE2- and BSG-mediated pathways in viral entry, they incubated human kidney embryonic cells transiently expressing ACE2 or BSG with pseudovirus expressing the spike protein. As expected, they observed that only ACE2 expression, and not BSG expression, facilitated the viral entry into cells.
Because an active viral infection is associated with a pro-inflammatory cellular environment, the scientists thought of checking if inflammatory mediators can facilitate the viral entry into endothelial cells. For this, they primed the cells with interleukin-1β (IL-1β), a leukocytic pyrogen, which led to increased production of other pro-inflammatory cytokines, such as IL-6 and IL-8. However, even in this inflammatory milieu, endothelial cells remained resistant to SARS-CoV-2 infection.
Given the high resistance of endothelial cells to SARS-CoV-2 infection, the scientists believe that molecules released by nearby epithelial cells or systemic pro-inflammatory mediators are mainly responsible for cardiovascular complications related to COVID-19. The pro-inflammatory cellular environment created by SARS-CoV-2 infection may be responsible for the loss of barrier function, facilitating the viral entry into blood vessels and its transmission to multiple organs via the bloodstream.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.