In a recent cutting-edge study, US researchers unveiled an anti-staphylococcal enterotoxin B monoclonal antibody that can be repurposed to target the cleavage site of the SARS-CoV-2 spike glycoprotein and, in turn, inhibit viral infection. Their findings are currently available on bioRxiv* preprint server.
SARS-CoV-2, a causative agent of coronavirus disease (COVID-19), can give rise to severe interstitial pneumonia with hyperinflammation, as well as a plethora of extrapulmonary manifestations. Since the COVID-19 pandemic is still ongoing, there is a pressing need for effective therapeutics that inhibit viral infection and suppress the hyperinflammatory cytokine storm in severe forms of the disease.
A novel multisystem inflammatory syndrome (MIS), which is reported in both children and adults, has been described in patients that either had a positive test result or epidemiological links to COVID-19. In children, MIS manifests as persevering fever and hyperinflammation with the involvement of many organs in the body.
Some striking similarities between MIS in children or severe COVID-19 and the toxic shock syndrome (TSS) that arises due to the action of bacterial superantigens (SAg) led to the hypothesis that SARS-CoV-2 might possess a SAg-like motif that can trigger hyperinflammation processes.
Recently, a superantigen-like motif akin to staphylococcal enterotoxin B (SEB) was discovered near the S1/S2 cleavage site of the SARS-CoV-2 spike protein, which might explain the MIS in children and cytokine storm in patients with severe COVID-19.
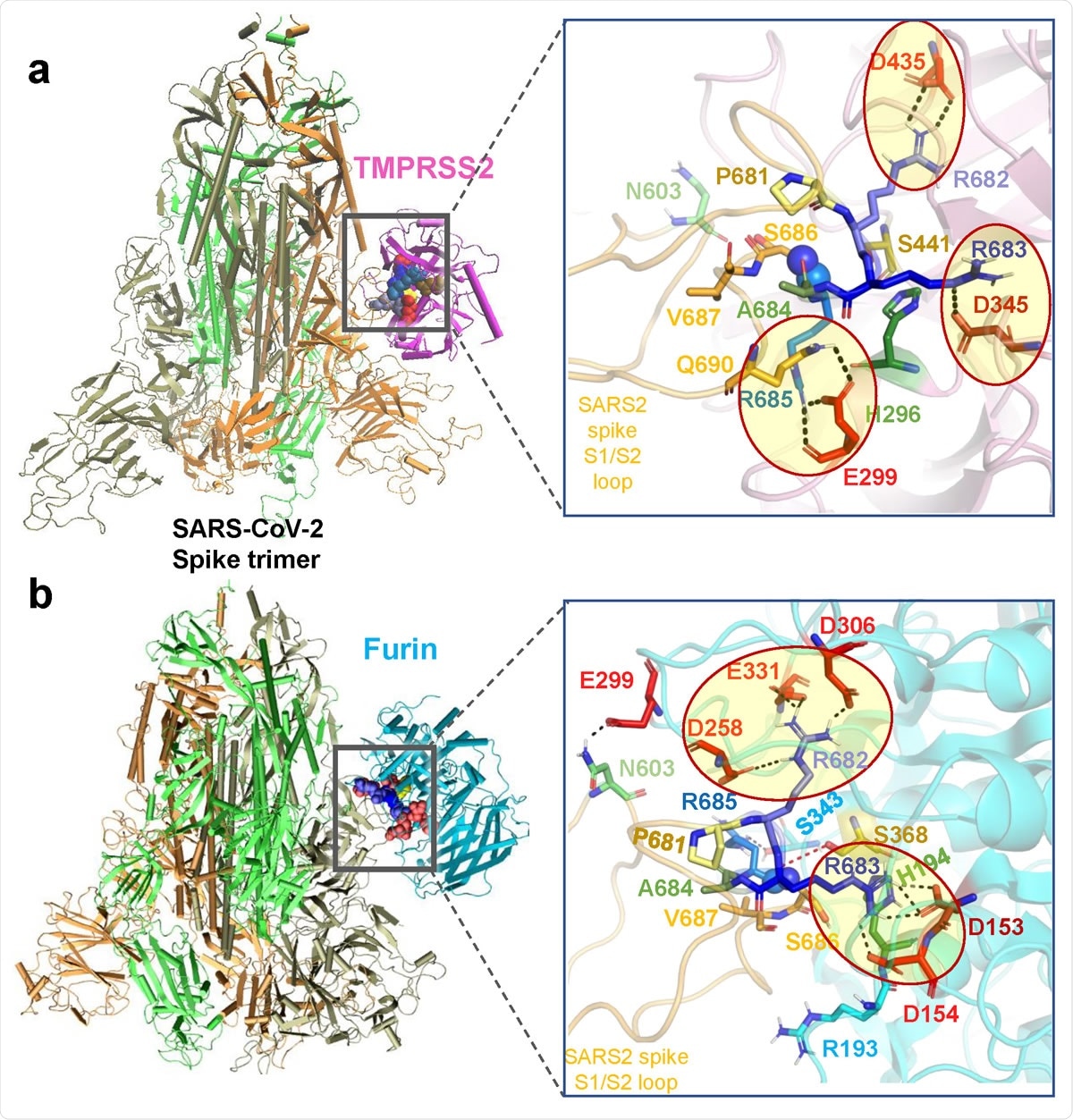
Binding poses of human proteases TMPRSS2 and furin to SARS-CoV-2 S protein. (a-b) Structural models for the SARS-CoV-2 S protein complexed with TMPRSS2 (a), and furin (b), obtained from docking simulations followed by refinements. An overview (left) and a zoomed in view (right) are shown in each case. The arginines in the S1/S2 loop P681RRARS686 are shown in different shades of blue, and their interaction partner (acidic residues) in the proteases are shown in red. Spheres (right panels) highlight the peptide bond that would be cleaved (between R685 and S686). TMPRSS2 catalytic triad residues are in yellow for S441, green for H296, and dark red for D345. Their counterparts in furin are S368, H194 and D153. Note the short distance between the carbonyl carbon from R685 and the hydroxyl oxygen of the catalytic serine S441 of TMPRSS2 (3.5 Å) or S368 of furin (3.1 Å). Black dashed lines show the interfacial polar contacts and salt bridges, and those including the S1/S2 loop arginines are highlighted by ellipses.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Superantigen-like region in SARS-CoV-2
This specific SAg-like region overlaps with the S1/S2 cleavage site on the surface of SARS-CoV-2 spike glycoprotein, but even more interesting finding in that area is a unique insertion sequence 681PRRA684 (or PRRA in abbreviated form).
The aforementioned insert also takes part in the SAg-like motif, and SARS-CoV-2 is actually the only beta-coronavirus that harbors such sequence, despite striking similarities with other members of this viral genus.
Furthermore, its role in recognizing and binding transmembrane protease serine 2 (TMPRSS2) and furin found in the host is well known, while its cleavage activity is closely linked to spike protein priming that facilitates SARS-CoV-2 entry and increases its infectivity after binding to the host cell co-receptor neuropilin-1.
In view of the sequence and structure similarities between the PRRA-insert-enclosing SAg-like motif and the bacterial enterotoxin SEB, a research group led by Dr. Mary Hongying Cheng from the University of Pittsburgh hypothesized that previously generated anti-SEB monoclonal antibodies (mAbs) may bind the viral SAg-like motif (particularly PRRA) and subsequently block access to the S1/S2 cleavage site.
From the hypothesis to the experimental model
As a result, in this paper researchers have focused on this polybasic site – which is basically unique to SARS-CoV-2 among SARS-family of betacoronaviruses – as a target for binding of mAbs. Hence, they have appraised possible interactions of known antiSEB mAbs with SARS-CoV-2 spike gylcoprotein.
The researchers have also generated structural models to explain the interaction of the spike protein with TMPRSS2 and furin in order to demonstrate that the 6D3 binding site overlaps with those found in these proteases, indicating that 6D3 could interfere with viral entry.
Furthermore, they have conducted detailed in silico screening of SARS-CoV-2 neutralizing mAbs in order to probe their propensity to bind near the PRRA insert, or the furin-like cleavage site. Live viruses were also used in some of the experiments.
Successfully inhibiting viral entry
This study showed rather high affinity of SEB-specific mAb 6D3 for binding to the S1/S2 site of SARS-CoV-2, while the computational analysis additionally predicted that the 6D3 mAb has a greater affinity for binding this polybasic region spike SAg motif when compared to TMPRSS2, and an affinity corresponding to that of furin.
Moreover, the experiments with live viruses revealed that 6D3 indeed successfully inhibited viral entry to the host cell. Since this specific site does not overlap with those observed to bind antibodies, the study has shown that 6D3 might actually be used together with other neutralizing antibodies that target receptor-binding domain or other non-overlapping sites in an attempt to increase the efficacy of mAbs.
"Our analysis revealed that 4A8 might also bind the same site and as a result obstruct the S1/S2 site", say authors of the study. "Therefore, 4A8 and 6D3 may potentially serve as scaffold for designing wide-spectrum Abs for reducing the infectivity of SARS-CoV-2 and even other human coronaviruses that harbor a furin-like cleavage site", they add.
A promising therapeutic pathway
In a nutshell, the affinity of 6D3 and 4A8 for this site unveils their potential usefulness for treating COVID-19, MIS in children, or even common cold caused by human coronaviruses that harbor a furin-like cleavage site.
"Our work may lead to an improved understanding of coronavirus immunity, facilitating future studies to understand mechanisms of antibody recognition and neutralization, and help screen SARS-CoV-2 antibodies for the treatment of COVID-19", conclude study authors in this bioRxiv paper.
These findings also raise exciting possibilities of combination antibody treatments in our battle against severe COVID-19 and MIS. Likewise, cocktail options (which combine furin-like cleavage site-targeting antibodies with RBD-targeting neutralization antibodies) may result in complementary protection against the dire pathogenesis of SARS-CoV-2.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Cheng, M.H. et al. (2020). A monoclonal antibody against staphylococcal enterotoxin B superantigen inhibits SARS-CoV-2 entry in vitro. bioRxiv. https://doi.org/10.1101/2020.11.24.395079, https://www.biorxiv.org/content/10.1101/2020.11.24.395079v1
- Peer reviewed and published scientific report.
Cheng, Mary Hongying, Rebecca A. Porritt, Magali Noval Rivas, James M. Krieger, Asli Beyza Ozdemir, Gustavo Garcia, Vaithilingaraja Arumugaswami, Bettina C. Fries, Moshe Arditi, and Ivet Bahar. 2021. “A Monoclonal Antibody against Staphylococcal Enterotoxin B Superantigen Inhibits SARS-CoV-2 Entry in Vitro.” Structure 29 (9): 951-962.e3. https://doi.org/10.1016/j.str.2021.04.005. https://www.cell.com/structure/fulltext/S0969-2126(21)00121-0?.