The coronavirus disease 2019 (COVID-19) pandemic is not showing any signs of letting up. Despite the initiation of vaccination efforts, scientists continue to work on effective antivirals to prevent and treat this disease, which requires a better understanding of how and why it spreads. A new preprint research paper published on the bioRxiv* server describes the role of a host protein called tetherin in the spread of the novel coronavirus.
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) gains entry to the host cell via the spike protein, which binds to the host cell receptor, the angiotensin-converting enzyme 2 (ACE2). This virus differs from earlier pathogenic coronaviruses in possessing a novel polybasic furin cleavage site that allows the spike to split into its two subunits when exposed to the host protease, furin.
The two spike fragments, S1 and S2, remain covalently bound, and the latter undergoes cleavage by another host protease, TMPRSS2. The S1 now binds to the host receptor neuropilin-1, allowing the virus to enter the cell. The virus bound at the cell's surface is enveloped within a complete fold of membrane, the endosome, that fuses with the viral envelope to liberate the viral capsid into the cytoplasm. Finally, viral RNA hijacks the cell organelles to replicate itself, synthesize other viral components, and assemble the new virions within the ERGIC (endoplasmic reticulum-Golgi intermediate compartment) organelles. The viral replication hubs within the cell are double-membrane vesicles (DMVs) composed of modified host organelles.
Viruses can be classified based on whether or not they have a host-derived lipid envelope. The envelope protects the viral capsid from the host immune response and thus enhances viral entry to new cells. On the other hand, the host cell can use this to incorporate antiviral factors into the new viral particles. Many enveloped viruses can tether themselves to the plasma membrane of cells once they reach the cell surface, hindering further spread and infection. This is mediated by tetherin, a host protein.
Tetherin restricts viral release
Tetherin is an integral type II membrane protein with a GPI anchor at the membrane, with a short tail inside the cytosol, and a large coiled domain outside the cell. It is expressed by many types of cells, but type I interferon increases tetherin levels. Tetherin molecules link the cell and the virus via homodimer formation, using three cysteine residues in the extracellular loop to create disulfide bonds. This holds the newly forming virions on the surface of the infected cells, allowing endosomes to hold on to them and preventing their spread.
Tetherin restricts the spread of several enveloped viruses, including HIV-1, Ebola, Kaposi's sarcoma-associated herpesvirus and human coronavirus 229E (HCoV-229E). Thus, several enveloped viruses use several mechanisms to lose tetherin and thus allow viral release. SARS-CoV downregulates tetherin, causing the increased spread of the virus, but the mechanism is unclear.
For tetherin to restrict the virus, these molecules must be built into the virus during the single enveloping step within ERGIC. However, the researchers found that while tetherin moves through these organelles, it does finally distribute at the plasma membrane and the endosomes. This further supports the hypothesis that tetherin restricts viral release when the endosomes or other DMVs fuse with the plasma membrane.
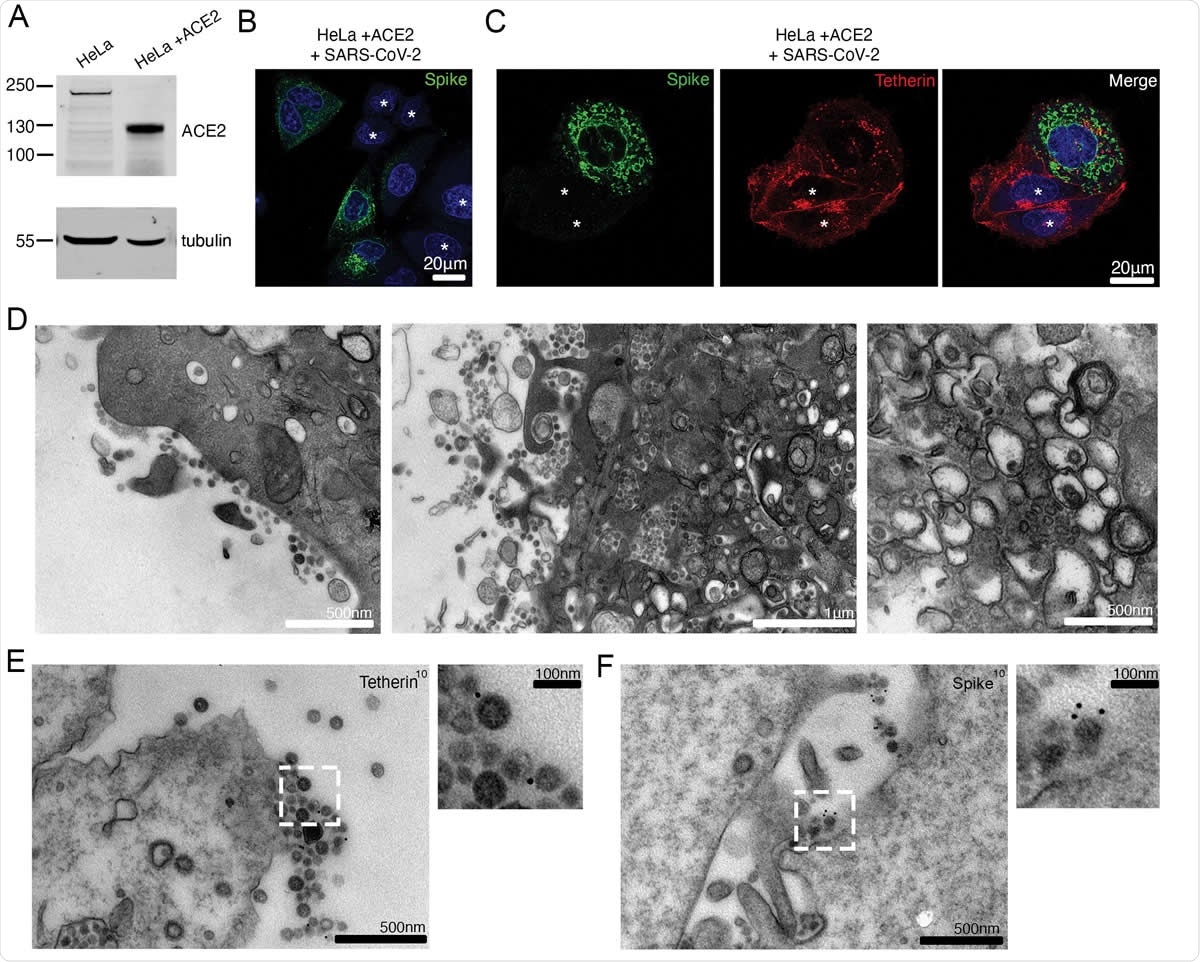
SARS-CoV-2 infection downregulates tetherin in HeLaWT +ACE2 cells. (A) HeLa cells were transduced with ACE2 lentivirus to generate stable cell lines. Mock and ACE2 transduced cells were lysed and immunoblotted for ACE2. Tubulin was used as a loading control. (B) HeLaWT +ACE2 cells were infected with SARS-CoV-2 (MOI 0.5). Cells were fixed at 24 hpi and stained for the spike (green) and DAPI (blue). (C) HeLaWT +ACE2 cells were infected with SARS-CoV-2 (MOI 0.5). Cells were fixed at 24 hpi and stained for the spike (green), tetherin (red), and DAPI (blue). Uninfected cells are shown with an asterisk. (D) Electron micrographs showing plasma membrane-associated SARS-CoV-2 virions and virus filled intracellular organelles. SARS-CoV-2 infected HeLaWT +ACE2 cells (MOI 0.5) were fixed at 24 hpi and processed for TEM. Left micrograph – plasma membrane-associated virus, middle micrograph – virus filled tubulovesicular compartments are directed towards the plasma membrane, right micrograph – virions within DMVs. (E) Surface immunogold electron microscopy of SARS-CoV-2 infected HeLaWT +ACE2 cells. Cells were infected with SARS-CoV-2 (MOI 0.5), fixed at 24 hpi and immunogold labeled with antibodies against tetherin. (F) As (E) but labelled with antibodies against SARS-CoV-2 spike.
Interferon regulates tetherin concentration
Type I IFN responses are attenuated in COVID-19 as in other coronavirus infections. Interferons have been tried out to treat this condition, perhaps by increasing the concentration of tetherin and thus hindering tetherin release. However, this is not the only mechanism by which tetherin is downregulated.
The current study demonstrates that with SARS-CoV-2, the virions are bound to the infected cell surfaces by tetherin. The infection of these cells results in a steep decline in the level of expression of tetherin on the cell surface. They found that the viral spike protein mediates the downregulation of tetherin, while the ORF7a causes fragmentation of the Golgi.
SARS-CoV-2 disrupts tetherin synthesis
The researchers found that uninfected human cells display tetherin on the plasma membrane and within the perinuclear compartments. After infection, tetherin was lost from the plasma membrane, and staining points appeared within the cell at higher concentrations. Some areas of the cell membrane appeared to be stained, indicating areas where the virus was tethered to the cell surface.
Using transmission electron microscopy (TEM), they found that the viruses were clustered on the membrane and not uniformly distributed. Viruses within ERGICs were found near such sites. DMVs were also seen. However, Golgi cisternae were absent, indicating disruption of the biosynthesis of proteins.
However, the researchers found that surface protein synthesis was not universally suppressed. Instead, specific proteins are affected. They discovered reduced tetherin concentrations. When tetherin expression is reduced, they found a dramatic decline in surface-associated virions, though virions inside DMVs were found in the perinuclear region in infected cells.
When tetherin expression was decreased, the infected cells released many more viral particles, indicating their inability to restrain the new virions during assembly.
The researchers concluded, "Tetherin exerts a broad restriction against numerous enveloped viruses, regardless of whether budding occurs at the plasma membrane or within intracellular compartments."
Role of viral proteins in tetherin downregulation
Next, they examined the role of the viral ORF7a protein, which showed that it was associated with the fragmentation of the Golgi apparatus. This interrupted protein biosynthesis within the cell. This does not appear to make it critical in human infection, however. A higher expression of viral ORF3a leads to the accumulation of tetherin within the cell. This could reflect the fact that tetherin is trafficked to the lysosomes but degraded slower than usual.
On the other hand, viral spike protein leads to tetherin loss from the cell surfaces. This is in addition to its known role in engaging to the host cell surface and causing cell to cell fusion following infection.
What are the complications?
The researchers comment, "The convergent evolution displayed by different families of enveloped viruses to downregulate tetherin highlights how important overcoming cellular restriction is to the success of enveloped viral pathogens." Since tetherin loss is associated with increased viral titer, this finding may facilitate the development of novel therapeutics that target tetherin inhibition or downregulation to offer an effective way of managing many such disease conditions.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.