The infectious nature of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has given rise to several mutations which have led to the emergence of variants such as B.1.1.7, B.1.351, P1, and B.1.617. These pathogenic lineages were originally detected in the United Kingdom, South Africa, Brazil, and India, respectively.
The widespread transmission of these variants has also had significant impacts on the global coronavirus disease 2019 (COVID-19) pandemic, with new waves of the virus emerging because of a surge in infection rates around the world. As of June 30, 2021, COVID-19 has infected over 182 million people worldwide and has caused the deaths of almost 4 million people.
 Study: Comparative Structural Analyses of Selected Spike Protein-RBD Mutations in SARS-CoV-2 Lineages. Image Credit: Polina Tomtosova / Shutterstock.com
Study: Comparative Structural Analyses of Selected Spike Protein-RBD Mutations in SARS-CoV-2 Lineages. Image Credit: Polina Tomtosova / Shutterstock.com

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
A computational study that analyzes and explores the time-based structures of the SARS-CoV-2 variants with specific spike (S) protein mutations has recently been conducted at Clarkson University in Atlanta, Georgia. The findings, which were recently published on the preprint bioRxiv* server, will be further discussed in this article.
Computational immunoinformatics
During the development of preventative drug designs, the field of computational structural biology has become a key aspect for applied immunology, as it enables the understanding of protein structure at a basic level. During the COVID-19 pandemic, scientists have only been able to access limited studies on relatable research which could help inform them of therapeutic targets or on how to handle the infectious level of this virus. However, due to the limited availability of published studies on the subject, researchers instead relied on simulation-based tools and strategies in their attempts to study SARS-CoV-2.
Utilizing computational immunoinformatics for this purpose can assist in effectively and appropriately investigating this virus while simultaneously eliminating exposure to this infectious agent. The research discussed in the current study focuses on exploring the lineages of the SARS-CoV-2 variants of concern (VoCs). Additionally, the authors of this study looked to evaluate the implications of multiple point mutations on the spike protein receptor-binding domain (RBD), all of which are provided in Figure 1. Through measuring the structural and conformational mutations of the variants, the association between the structure and function of SARS-CoV-2 can be more effectively understood.
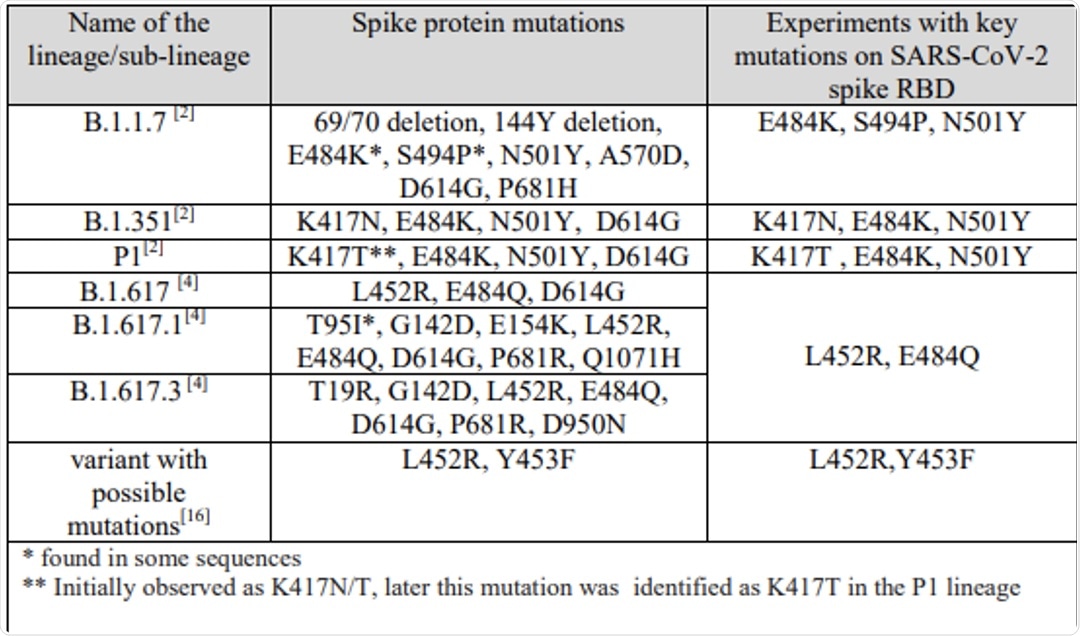 Figure 1. This table illustrates variants of concern which have been selected for investigation regarding specific mutations within the SARS-CoV-2 lineages and the experiments with key mutations on the spike protein RBD.
Figure 1. This table illustrates variants of concern which have been selected for investigation regarding specific mutations within the SARS-CoV-2 lineages and the experiments with key mutations on the spike protein RBD.
Genome of SARS-CoV-2
The genome of SARS-CoV-2 consists of both structural and non-structural proteins (NSPs). Whereas the non-structural proteins include 16 components, the structural proteins include the S, envelope (E), membrane (M), and nucleocapsid (N) proteins. The S glycoprotein of SARS-CoV-2 is the main interacting site by which the virus enters into the host cell; therefore, studies assessing the host immune response and antibody neutralization capabilities often focus on this protein.
The genome of SARS-CoV-2 is approximately 1,273 amino acids long and contains both S1 and S2 subunits of the S protein. The N-terminus signaling peptide is present at the start of this structure, which is then followed by the S1 subunit that assists with the receptor binding. The S1 subunit consists of two domains which include the N-terminal domain (NTD) and the receptor-binding domain (RBD). Mutations related to the S1 RBD are the focus of the current study, as the primary author Urmi Roy aimed to better comprehend the role of these specific mutations in SARS-CoV-2 VOCs.
In the current study, root-mean-square deviation (RMSD) plots were used to explore the SARS-CoV-2 variants with selected S1-RBD mutations. This can be used along with root-mean-square fluctuations (RMSF), as they provide structural indications on which amino acids in a protein are most likely to contribute to a molecular motion. Taken together, this information can be used to provide insight into which structural components can contribute to the mutations of the variants.
Protein stability
The study discusses the lack of consensus on whether a protein’s stability is dependent on the number of mutations that are present within the variant. By confirming this association, researchers will be able to better understand how these mutations affect the protein’s stability and, overall, the behavior of the virus. A single recorded mutation can also assist in predicting the effects of other mutations in the same amino acid site, which can further aid in combating emerging SARS-CoV-2 variants.
In the current study, the RBD mutation of both the B.1.617 and P1 variants are identified as the most stable forms of this mutation. Roy also concludes that these mutations, which include L452R, Y453F, E484Q, S494P, are also quite stable. N501Y, as well as the E484K mutations within the P1 lineage, were also found to have minimal fluctuations while also maintaining their stability in VOCs. However, certain mutations, such as K417N/T, E484K, L452R, and Y453F, are recognized as antibody neutralizing escape mutants.
The B.1.617 lineage and the L452R-Y453F mutations are characterized by minimal fluctuations, which is comparable to mutations in the B.1.351 lineage that show a high volume of fluctuations. The secondary structures of proteins, such as alpha (a) helices and beta (b) sheets, have a significant role in the protein’s stability, with mutations such as S494P in the B.1.1.7 variant being quite stable. The B.1.617 variant, which was first detected in India, was initially labeled as a ‘double mutant,’ as it consists of two mutations (E484Q, L452R) from different lineages.
Protein-level structure mutations, such as those that cause coils to transform into b sheets, are often intricate. As a result, these mutations, as well as methods that can be used for their inhibition, must be extensively studied in order to manipulate them for vaccine and therapeutic drug development purposes.
These findings are helpful when investigating infectious variants of SARS-CoV-2 as it will improve the understanding of antibody cross-reactivity and also help with producing inhibitors with specific therapeutic targets. When certain mutations have enhanced stabilities, as has been seen in the emerging variants, there can be significant consequences to the production of vaccines and therapeutic agents. Subsequently, investigating the structures of these variants at both the amino acid and protein levels can provide insight into how this can affect the transmissibility of the virus.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources