With the coronavirus disease 2019 (COVID-19) pandemic causing unprecedented morbidity and mortality, the development of an effective antiviral drug for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has become a global health priority. Although many drug candidates have been identified via in vitro and in vivo models, there is a lack of consistent evidence from clinical studies, which may be due to the imperfect trial designs. The correct design of clinical trials for antiviral drugs is not well studied, especially concerning sample size calculation methodology.
“Most clinical studies of antiviral drugs for Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) have failed to observe a statistically significant effect, which may be due to poor designs of clinical trials.”
A modeling study to investigate the reasons behind inconsistent clinical trial findings and design better clinical trials
Japanese researchers recently investigated how randomized clinical trials for antivirals must be designed, focusing on the sample size. In addition, they conducted a modeling study to understand the reasons for inconsistent clinical trial findings and to help design better clinical trials. Longitudinal viral load data for SARS-CoV-2 without antiviral treatment was first analyzed using a within-host virus dynamics model. This study has been published in the open-access journal, PLOS Medicine.
The fitted viral load was classified into three groups using a clustering approach. In comparing the estimated parameters, three distinct groups were found to be characterized by different virus decay rates. The mean decay rates were 1.17 d−1, 0.777 d−1, and 0.450 d−1 for the 3 groups, respectively. This heterogeneity in viral dynamics could be a puzzling variable if it is linked to treatment allocation in compassionate use programs or observational studies.
“We believe this is the first study investigated the study design of clinical trials for antiviral treatment using the viral dynamics model.”
They also mimicked randomized controlled trials of antivirals by adding an antiviral effect causing a 95% to 99% decrease in viral replication to the model. In order to be realistic, they assumed that randomization and treatment are started with a time lag after symptom onset.
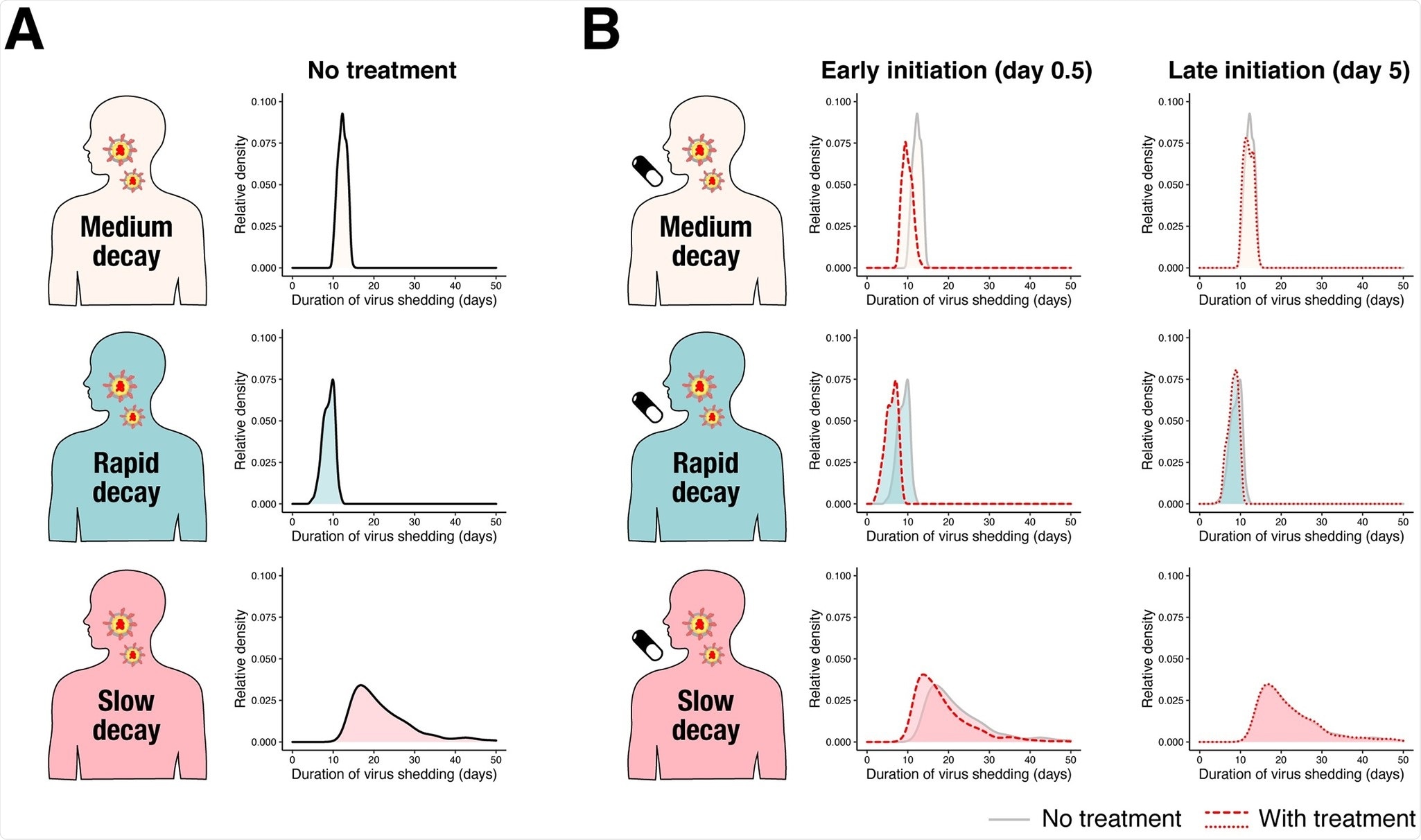
Duration of virus shedding in the 3 different groups. (A) The relative density distributions of duration of virus shedding since symptom onset for the 3 groups (light pink: medium decay group, light blue: rapid decay group, and pink: slow decay group) without treatment obtained through simulation. (B) The relative density distributions of duration of virus shedding since symptom onset for the 3 groups under antiviral treatment with different inhibition rates and different timing of treatment initiation. The left 3 panels are when antiviral treatment is initiated at 0.5 days (“Early initiation”) since symptom onset. The red dotted line corresponds to the distribution with a 99% inhibition rate. The distribution without treatment is shown in the back for comparison. The right 3 panels are when antiviral treatment is initiated at 5 days (“Late initiation”) since symptom onset. The distributions are represented as “relative density” to reflect different proportions of the 3 groups.
Results show that sample size can be dramatically reduced by enrolling patients immediately after onset of symptoms
The duration of virus shedding was used as an outcome in this study. The sample size needed to detect a statistically significant mean difference between the placebo and treatment groups was 11,670 and 13,603 when the antiviral effect was 99% and 95%, respectively, per group. The sample size was reduced to 584 and 458 for placebo and treatment groups when the antiviral effect was 95% and 99%, respectively, only when patients treated within a day of symptom onset are enrolled. The study confirmed the sample size was similarly reduced while using the cumulative viral load in log scale as an outcome.
Estimated association in observational studies can be biased due to large heterogeneity in viral dynamics across patients
According to the authors, they used a conventional virus dynamics model that may not fully reflect the detailed mechanisms of SARS-CoV-2 dynamics. The parameter settings and model structure of their model need to be calibrated to obtain a more reliable sample size calculation.
“More precise models reflecting the features of SARS-CoV-2 infection may provide more reliable sample size estimates.”
In conclusion, the researchers found that the estimated association seen in observational studies can be biased because of the considerable heterogeneity in viral dynamics across infected individuals. Also, it may be challenging to detect a statistically significant effect in randomized trials due to the small sample size. Simulation mimicking randomized controlled studies showed that the sample size would be unreasonably large at over 11,000 per group if the timing of treatment initiation is not considered. The sample size can be drastically reduced by enrolling patients immediately after symptom onset.
“Randomized controlled trials for antiviral drugs should recruit patients as early as possible after symptom onset or set inclusion criteria based on the time since symptom onset to observe statistically significant results.”