Researchers in the Pirbright Institute, UK, have experimentally proved that virus-related double-stranded RNA (dsRNA) and the sites of viral RNA synthesis are associated with one another. In addition, they confirm the location of viral synthesis that exists ‘inside’ the membrane-bound compartments within the cell.
Replication organelles and their significance
Replication organelles (ROs) are specialized membranous structures that serve as the sites of robust virus replication. All positive sense (+) RNA viruses, including coronaviruses, remodel host cell intracellular membranes into these specialized structures, that efficiently coordinate the steps of virus replication cycle as well as protect the virus particles from host cell defenses. The RO structures may vary between virus families, from convoluted membranes, double membrane vesicles (DMVs), to spherules, but they all share structural similarities.
Owing to the possibility of antiviral therapies being able to target the ROs, extensive studies have been carried out in recent years to locate the sites of coronavirus RNA synthesis. SARS-CoV RNA synthesis has been shown to take place inside membrane-bound compartments, using biochemical methods. More recently, DMVs are shown to be the sites of viral RNA synthesis during MERS-CoV and IBV replication. However, the methods used in the study could not conclusively show the location of RNA synthesis, i.e., whether on the interior or cytoplasmic face of DMV membranes.
A subsequent study showed that DMV membranes are porous, providing a route by which RNA synthesized inside DMVs could exit to cytoplasm for translation or assembly. Despite this, the location of viral RNA synthesis either inside or on the outside of DMVs remained to be confirmed.
IBV, a gammacoronavirus, is an economically important virus to the poultry industry causing highly contagious respiratory disease in chickens and other poultry birds, has been shown to induce the formation of DMVs, zER (zippered endoplasmic reticulum membranes) and spherules. In the current study, using IBV, bromouridine (BrU)- labels viral nascent RNA, and different cell permeabilization methods, the team investigated the localization of viral RNA synthesis and the link with replication organelles in host cells.
Background
dsRNA is considered to be a replication intermediate, formed during the replication of the viral genome and its formation is a well conserved step across all +RNA virus families. To understand whether dsRNA is protected within membrane-bound compartments, the team utilized IBV infected cells and two permeabilization agents, Digitonin and Triton X-100 (TX100).
Precisely, Digitonin is a weak permeabilizing agent, which at low concentrations selectively permeabilize plasma membrane, but not intracellular membranes, while TX100 permeabilizes all cellular membranes. IBV infected chicken fibroblast DF1 cells were permeabilized with either TX100 or digitonin, then labeled with antibodies specific for non-structural protein 12 (nsp12) and dsRNA. The team had shown previously that nsp12, the viral RNA-dependent RNA polymerase (RdRp), does not colocalize with dsRNA over the course of the IBV life cycle.
In the current study, cells permeabilized with TX100 clearly showed both nsp12 (green) and dsRNA (red) labeling inside cytoplasm. dsRNA labeling increasing markedly over the course of infection. However, when cells were permeabilized with digitonin, the weak permeabilizing agent, dsRNA was no longer visible in the cells, which indicated that it was held within an intracellular membrane and, therefore, became inaccessible to the antibody. nsp12 staining, however, remained unaffected on permeabilizing with digitonin, indicating that it is possibly existing free in the cytoplasm or on the cytoplasmic face of a membrane.
To understand the association of dsRNA with sites of viral RNA synthesis, the team incorporated uridine analog bromouridine (BrU) into the nascent viral RNA. Transcription inhibitor, Actinomycin D (ActD) was used to selectively inhibit cellular transcription to allow visualization of sites of active viral RNA synthesis. Using super-resolution microscopy, the team confirmed that dsRNA and BrU signal (signifies site of active viral RNA synthesis) were in close association with each other. However, over the course of IBV infection, nsp12 did not colocalize with BrU.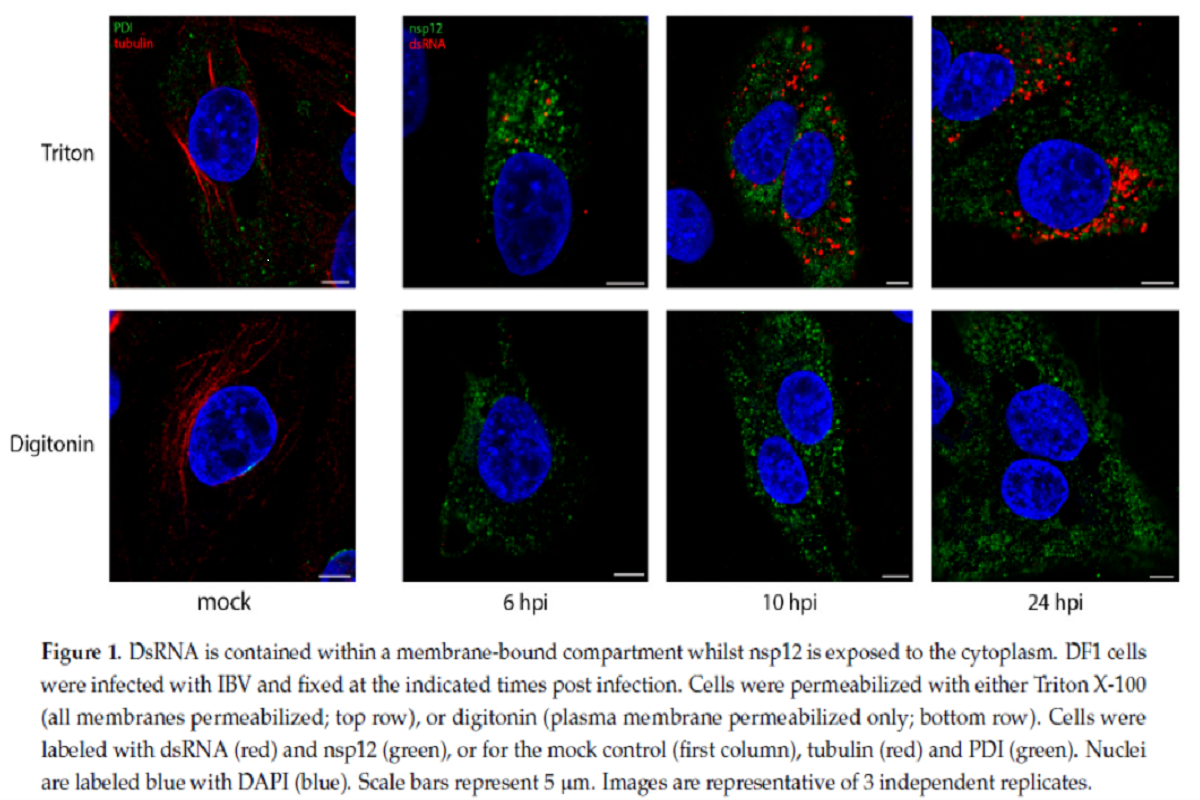
Location of viral RNA synthesis sites
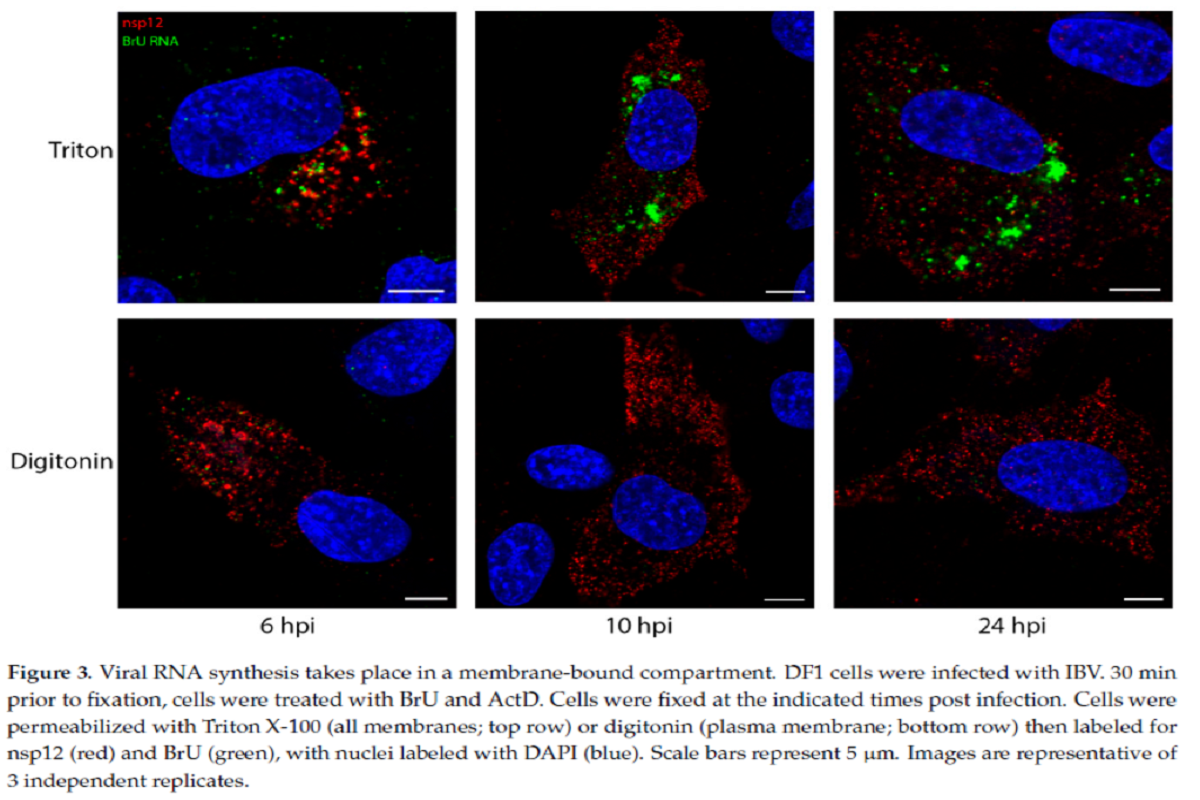
To figure whether the sites of viral RNA synthesis are located on the inside of these membranous structure or outside, the team did another experiment where DF1 cells were infected and treated with BrU and ActD, followed by immunofluorescent (IF) labeling using either TX100 or digitonin permeabilization.
The newly synthesized viral RNA labelling pattern increased through the course of infection from smaller puncta to larger foci in TX100-permeabilized cells. Noticeably, following digitonin permeabilization, most of the BrU signal at different timepoints was not detectable. Some of the newly synthesized viral RNA was found in the cytoplasm, however this observation indicated that the larger proportion of newly synthesized viral RNA was enclosed within a membranous structure.
Overall viral RNA labeling followed a very similar staining pattern to dsRNA and suggested the same membrane-bound compartmental location for both the molecules.
A pulse-chase experimental approach with bromouridine (pulse) and unlabeled uridine (chase), demonstrated that by 24 hours post infection (hpi), viral RNA that was produced between 7–8 hpi showed two separate pools. The first pool remained in large membrane-bound foci, possibly containing positive and negative sense RNA templates. The second pool, possibly containing newly synthesized positive sense RNAs, got exported to the cytoplasm and was associated with structural proteins, perhaps in the process of assembly into new virions.
Like IBV (betacoronavirus), the team analyzed one representative virus from each of the rest three other genera of coronaviruses (HCoV 229E [alphacoronavirus], SARS-CoV-2 [betacoronavirus], and PDCoV [deltacoronavirus]) for this experiment. Location of nascent viral RNA labeled with BrU was assessed using TX100 and digitonin as before. Consistent with the observations for IBV, the experiment demonstrated a conserved mechanism for viral RNA synthesis to be held within a membrane-bound compartment across the whole coronavirus family.
Implications
A previous study has reported the existence of transient pores in the DMV membrane. The reported diameter of pore is 2–3 nm at their narrowest point, which is not large enough to allow entry of an antibody complex of ~30 nm. The team therefore suggests it is likely that the virus aims to shield the dsRNA from detection intracellular pattern recognition receptors (PRRs) within these membrane-bound compartments.
Regarding the fact that activation of interferon (IFN) signaling is delayed following IBV infection, it was suggested that the IFN response later in IBV infection could be happening due to dsRNA “leaking” from DMVs. However, based on the current finding that dsRNA is sealed within the membrane compartment at 24 hpi, a timepoint after which IFN signaling is activated, the team points towards the existence of another mechanism to allow for activation of IFN signaling later in IBV infection, and these remain to be elucidated.
Knowing that all CoVs synthesize viral RNA within a membrane-bound compartment is a significant step in understanding the replication of this important virus family” concludes the team.