The coronavirus disease 2019 (COVID-19) pandemic has caused severe disruption of the health, social and economic systems around the world for almost two years. Despite the rapid development and rollout of vaccines against the causative severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the subsequent emergence of a number of variants of concern (VOCs) has challenged the ability to control the spread of SARS-CoV-2.
Taken together, it remains important to understand how SARS-CoV-2 causes disease and how its treatment affects humans. A new study under consideration at the journal Microbiome and posted to the preprint server Research Square* discusses changes in the microbiota of various parts of the body with SARS-CoV-2 infection.
Study: Assessment of Microbiota in The Gut and Upper Respiratory Tract Associated With SARS-Cov-2 Infection. Image Credit: ART-ur / Shutterstock.com `

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Introduction
SARS-CoV-2 binds the host cell angiotensin-converting enzyme 2 (ACE2) to subsequently enter cells to cause productive infection. The major target of SARS-CoV-2 is the respiratory tract, and COVID-19 pneumonia is a frequent occurrence.
However, many other organs are also known to be involved in SARS-CoV-2 infection, as ACE2 receptors are present in multiple types of organs and tissues. In fact, SARS-CoV-2 particles and nucleic acids have been identified in bronchoalveolar lavage fluid (BALF) samples, sputum, swabs from the nose and throat, blood, feces, and saliva.
Respiratory viruses can disrupt the microbial community on the mucosa of the gut and respiratory tract. This can trigger secondary bacterial infection, thereby increasing the severity of the illness and worsening the outcome.
To explore this, the researchers used a ribonucleic acid sequencing (RNA-seq) library construction strategy called MINERVA, which accelerates the time required for sequencing SARS-CoV-2 and metatranscriptomics from different parts of the body in COVID-19 patients. The types of changing patterns in the gut were compared to those in the respiratory tract in order to elucidate their relationship with SARS-CoV-2 load.
Study findings
The current study included almost 540 samples obtained from over 200 COVID-19 patients. All samples were taken within two weeks of symptom onset.
The researchers found that respiratory tract species were less diverse in patients as compared to controls, though species were equally rich in all pharyngeal, feces, and sputum samples. Despite the absence of any obvious difference in alpha diversity in the gut microbiome, there is a definite change in the flora in these patients.
The most severe illness was linked to the greatest reduction in diversity in all types of samples. With COVID-19, respiratory samples showed a reliable reduction in Shannon diversity; however, in feces samples, though the alpha diversity remained unchanged, the microbial composition was altered. This was due to an increase in SARS-CoV-2, along with antibiotic and antiviral therapeutics.
The greatest increase in microbial abundance was in Streptococcus that was present in respiratory samples, both normal and opportunistic species. In fecal samples, there was no difference in the most abundant species, of which included Bacteroides. However, Fecalibacterium showed a reduction, whereas Acinetobacter, Citrobacter, and Pseudomonas increased in feces samples.
Interestingly, many of the species that were reduced were involved in the production of short-chain fatty acids, engaged in the regulation of gut health, and participated in the gut inflammatory response.
Some species that showed increased numbers in COVID-19 were not typical of human feces. One example is Pseudomonas fluorescens, which is a low pathogenic bacterium, and Halomonas, which is usually an inhabitant of marine environments that is usually a commensal organism that is sometimes responsible for hospital infections.
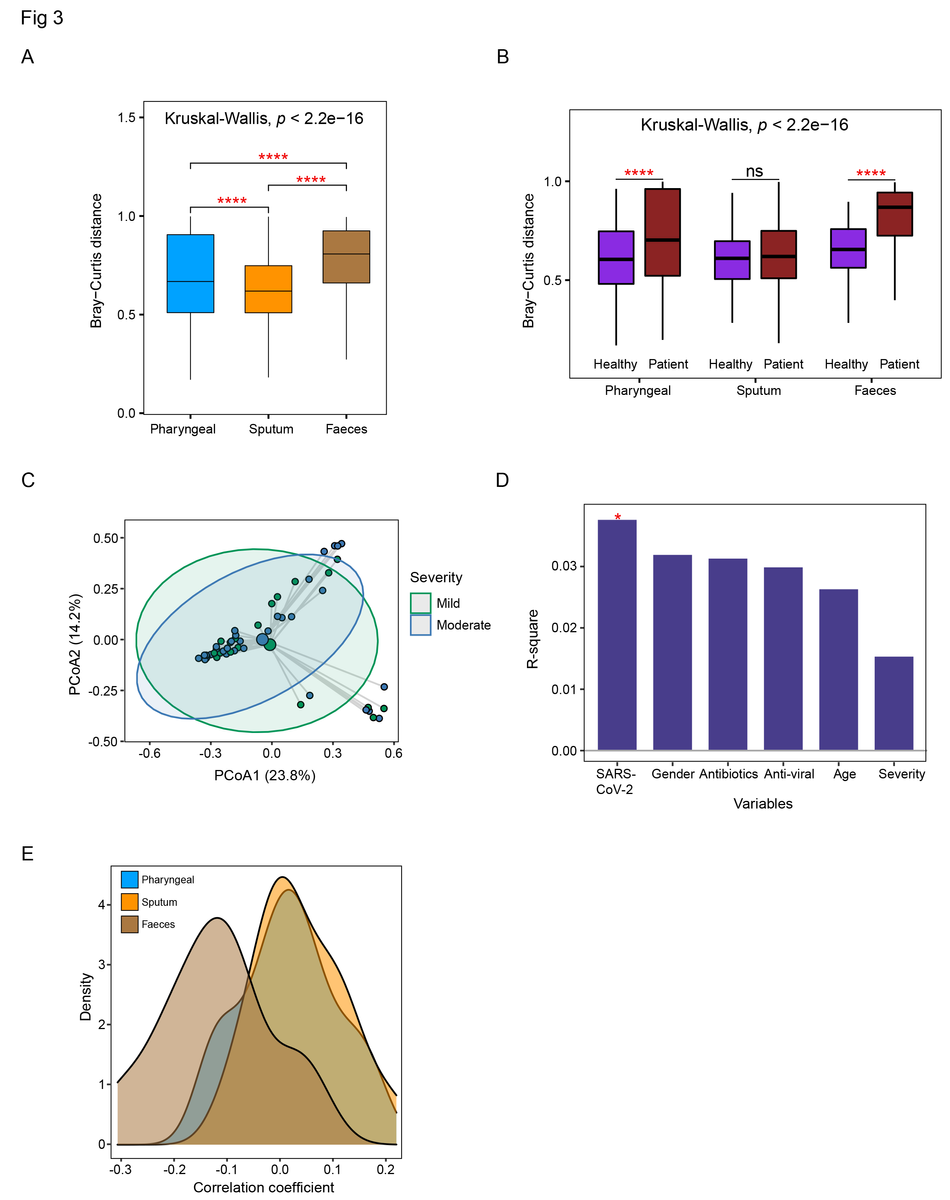
Comparison of the dysbiosis patterns of microbial composition in respiratory tract and gut samples. (A) Comparison of Bray-Curtis distance between patients and healthy controls in three sample types. (B) Comparison of the within-patient and within-healthy control Bray-Cutis distance in three sample types. (C) PCoA analysis of the microbial composition in faeces samples from mild and moderate patients. (D) PERMANOVA test to identify meta factors potentially associated with the microbial composition in faeces samples from mild and moderate patients. (E) Distribution of the correlation coefficients between SARS-CoV-2 and representative species in three sample types. Kruskal-Wallis test was used for multiple-group comparison and Wilcox rank-sum test was used for post-hoc two-group comparison.
Gut microbiota was more severely affected by SARS-CoV-2 infection as compared to respiratory samples, thus demonstrating the impact of disease severity and antibiotic treatment. Overall, the composition of the microbiota remained relatively stable during the study period.
The dysbiosis profile in the gut varied in functional alterations from that in the respiratory tract. In the gut, carbohydrate metabolism was affected along with single-chain fatty acid (SCFA) production, with a higher expression of genes related to the stress response. Conversely, stress genes and toxin genes predominated in the respiratory tract microbiome.
In pharyngeal samples, the transportation of many different molecules appeared to be a focus of gene upregulation in patient samples, as demonstrated by the higher level of expression of iron, zinc, manganese, or copper transport molecules. Cellular components that may indicate a higher risk of antibiotic resistance, such as multidrug efflux pump SatA and SatB, were enriched in severe COVID-19. Notably, both SatA and SatB participate in norfloxacin and ciprofloxacin ABC transport.
In sputum, transport genes, stress response genes, and virulence genes were similarly abundant in patients as compared to healthy controls.
Many genes in the respiratory tract microbiome also focused on the stress response, thereby indicating that the microbes were experiencing stress, whether due to SARS-CoV-2 infection or treatment. Fecal samples in healthy controls showed more genes responsible for fatty acid metabolism, including SCFA. However, in COVID-19 patients, genes expressed at higher levels were primarily responsible for amino acid metabolism and the stress response.
Implications
The current study shows that with COVID-19, and depending on the severity of illness, different patterns of alterations in the microbiomes of the gut and upper respiratory tract are observed. Both the composition and function of the microbiomes of these regions were affected.
Upper respiratory tract samples showed less diversity of the microbiome, perhaps because normal species were lost and Streptococcus species expanded. Notably, in the gut samples, beneficial species that generate SCFA were lost. This activity is key to gut mucosal integrity, regulation of metabolism, and homeostasis of the immune system.
Along with the depletion of normal species in both the gut and respiratory microbiomes, gut bacteria responded more readily to the abundance of SARS-CoV-2 and showed a wider range of responses among patients. This included loss of carbohydrate and fatty acid metabolism, including the production of SCFA.
Comparatively, the functional changes in the respiratory microbiome included a major increase in stress response and toxin response genes. These changes, which affect microbiome homeostasis, could persist for months after recovery, according to earlier research.
Taken together, the microbiome changes observed in this study support the evolution of antibiotic resistance during the phase of acute illness. This could prolong the recovery of the normal microflora at each location and cause delayed effects of the viral illness.
While the prophylactic application of antibiotics is sometimes essential for prevention and treatment of secondary infections, close monitoring and strategies for more precise use of antibiotics is urgently required in the ongoing SARS-CoV-2 pandemic.”

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Li, J., Jing, Q., Li, J., et al. (2022). Assessment of Microbiota in The Gut and Upper Respiratory Tract Associated With SARS-Cov-2 Infection. Research Square. doi:10.21203/rs.3.rs-1371786/v1. https://www.researchsquare.com/article/rs-1371786/v1.
- Peer reviewed and published scientific report.
Li, Jiarui, Qiuyu Jing, Jie Li, Mingxi Hua, Lin Di, Chuan Song, Yanyi Huang, Jianbin Wang, Chen Chen, and Angela Ruohao Wu. 2023. “Assessment of Microbiota in the Gut and Upper Respiratory Tract Associated with SARS-CoV-2 Infection.” Microbiome 11 (1). https://doi.org/10.1186/s40168-022-01447-0. https://microbiomejournal.biomedcentral.com/articles/10.1186/s40168-022-01447-0.