Common symptoms of coronavirus disease 2019 (COVID-19) are cough, fever, fatigue, and loss of smell. Many COVID-19 patients also have gastrointestinal symptoms. This correlates with the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection of the intestinal epithelium in the COVID-19 patients. The SARS-CoV-2 virus has so far claimed over 1.16 million lives worldwide since the initial cases were discovered in Wuhan, China, in late December 2019.
However, little is known about the importance of the enteric phase of SARS-CoV-2 - both for the viral lifecycle and for the development of COVID-19-associated pathologies. It is not clear if the symptoms seen in patients are associated with the direct replication of SARS-CoV-2 in the gastrointestinal (GI) tract or are a consequence of the strong proinflammatory response.
Megan L. Stanifer et al. study the immune response in the human gut upon SARS-CoV-2 infection. The study published as a bioRxiv* preprint reveals a robust proinflammatory response: a cell type-specific or a tissue-specific regulation of interferon-mediated signaling during SARS-CoV-2 infection.
In this study, the researchers explore if the innate immune response triggered in the human gut to combat viral infection is similar or distinct compared to the one triggered in other organs. They also compare the responses in infected and bystander cells. They use human ileum- and colon-derived organoids as a non-transformed culture model. The intestinal "mini-gut" organoids are an excellent model to study SARS-CoV-2 infection of the gastrointestinal tract.
It is understood that SARS-CoV-2 replicates in human intestinal epithelial cells; however, the specific cell-types that become infected are not well defined. Using single-cell transcriptomics approaches (scRNAseq) and targeted scRNAseq, the authors identify a subpopulation of enterocytes (namely, immature enterocytes 2) as the most susceptible to SARS-CoV-2 infection.
An important observation in this study is the lack of correlation between ACE2 expression level and the copy numbers of the SARS-CoV-2 genome in the cell. This implies that ACE2 expression can not be used for conjectures on infectibility.
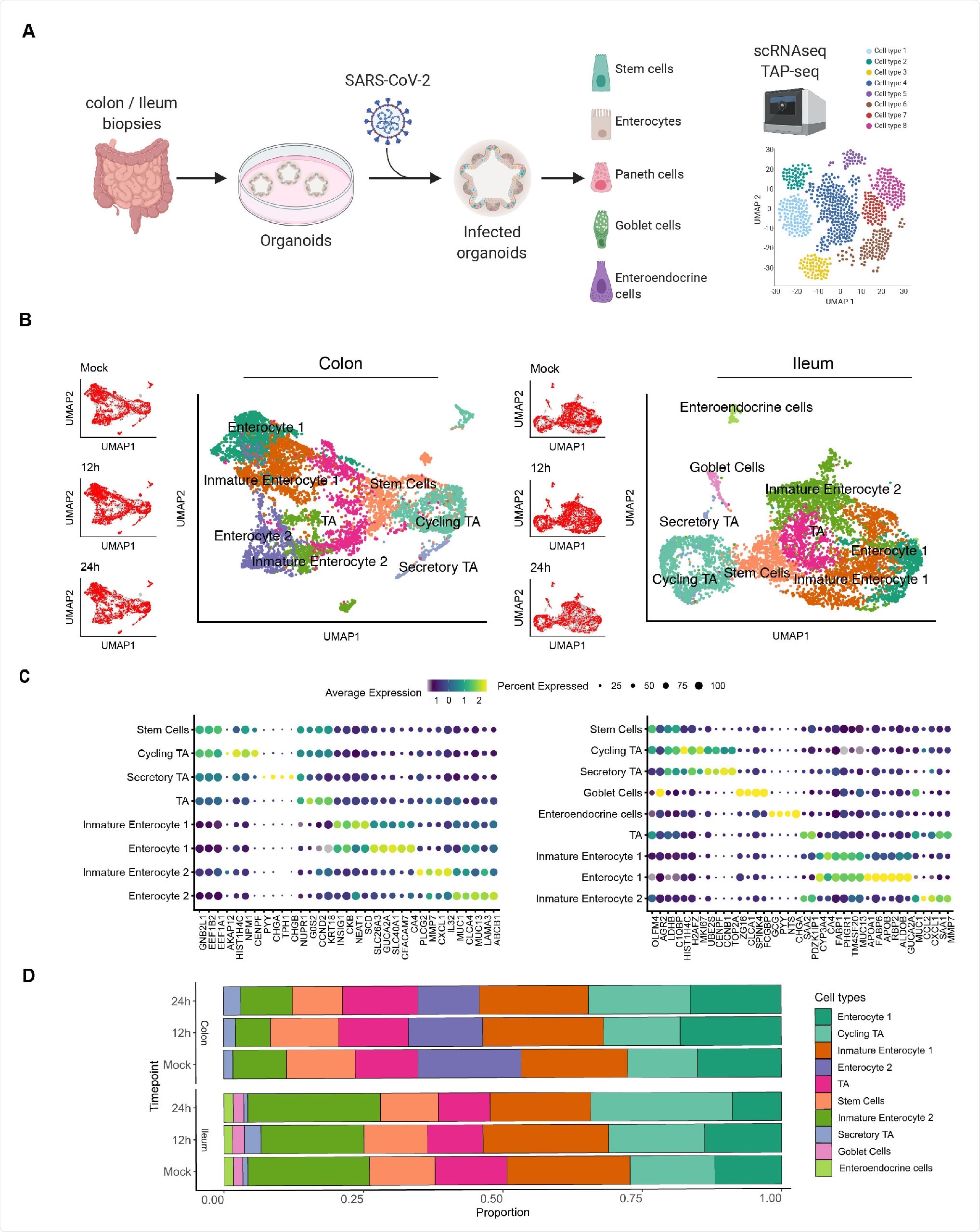
Single cell sequencing of SARS-CoV-2 infected colon- and ileum-derived human organoids. A. Schematic representation of the experimental workflow. 625 B. Uniform manifold approximation and projection (UMAP) embedding of single-cell RNA-Seq data from mock and SARS-CoV-2 infected colon-derived (left panels) and ileum-derived (right panels) organoids colored according to the cell type. Small insets represent the UMAP for mock and infected organoids at 12 and 24 hpi. C. Dot plot of the top marker genes for each cell type for (left) colon and (right) ileum-derived organoids. The dot size represents the percentage of cells expressing the gene; the color represents the average expression across the cell type. D. Bar plot displaying the proportion of each cell type in mock and infected organoids (12 and 24 hpi).

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Therefore, the authors emphasize that the investigation of SARS-CoV-2 tropism must be carried based on biological validation of infection and should not be done solely on the analysis of transcriptional profiles of individual cells or tissues.
Since they found that TMPRSS2 (type II transmembrane serine protease 2) expression levels were associated with the SARS-CoV-2 genome copy numbers in human intestinal epithelial cells, they speculate TMPRSS2 plays an essential role in the SARS-CoV-2 cell tropism, more so than the role of ACE2.
They also find that upon infection, ACE2 levels decrease both in infected and bystander hIECs (human intestinal epithelial cells). These show that the observed regulation of ACE2 upon infection might be tissue-specific and time-dependent.
When the researchers treated the cells with UV-inactivated SARS-CoV-2 - interferon (IFN), an interferon-stimulated gene (ISG) production did not occur. This implies that active virus replication is required to induce an IFN-mediated response.
The proinflammatory response occurs by upregulation of the NFκB and TNF pathways - this is clearly observed in the infected intestinal cells. The bystander cells do not activate the proinflammatory response, as observed in other similar studies. The inflammatory response in SARS-CoV-2 infected human intestinal epithelial cells is discussed in detail by the researchers. A moderated upregulation of IFN expression in infected cells in the current model is observed, while a strong ISG upregulation is observed in bystander cells.
The researchers also compare the immune response generated by organoids derived from different GI tract sections: 1) ileum organoids were more immunoresponsive than colon organoids, 2) a uniform up-regulation of proinflammatory genes across, and 3) ileum organoids, particularly bystander cells, produced more ISGs comparatively.
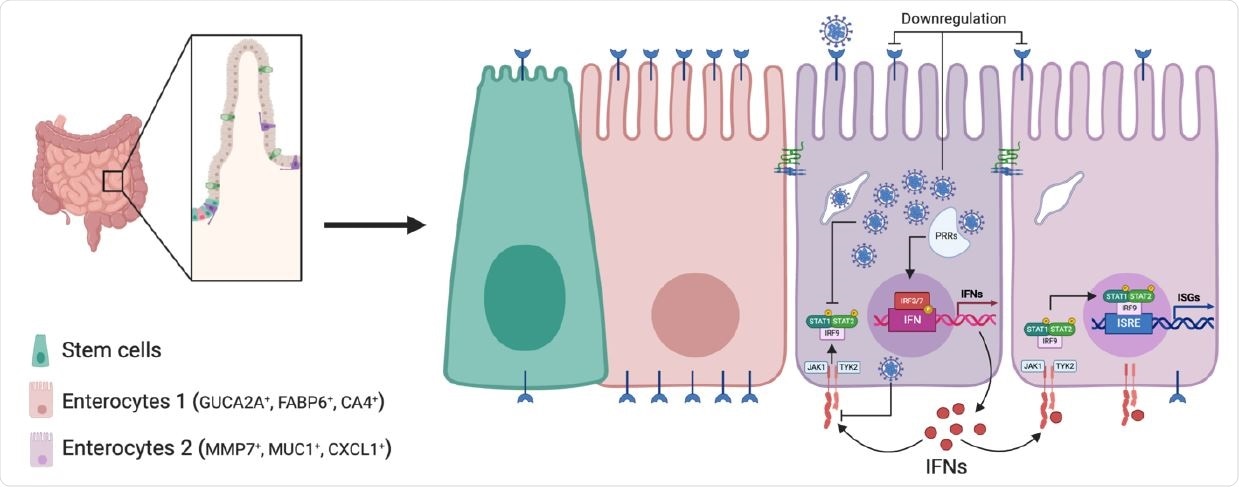
Schematic of SARS-CoV-2 infection of human intestinal epithelial cells. SARS CoV-2 infects a subpopulation of enterocytes. Upon infection, enterocytes mount a pro inflammatory response characterized by the upregulation of NFκB and TNF. Bystander cells respond to secreted IFN and upregulate the expression of ISGs. SARS-CoV-2 infection induces the downregulation of ACE2 expression and interferes with IFN-mediated signaling in infected cells.
This agrees with the previously known fact that different sections of the GI tract respond differently to microbial challenges.
This study's results reveal that the production of ISGs is mostly restricted to the bystander while the production of IFN is detected mostly in infected cells. The latter also fails to produce ISGs and become refractory to IFN stimulus. This implies that the SARS-CoV-2 has developed proviral mechanisms to shutdown IFN-mediated signaling and the subsequent production of ISGs in infected cells.
This reveals a replication advantage for de novo virus production by curtailing the signaling of antiviral pathways.
This is an important study where the cellular and the molecular players are identified in the SARS-CoV-2 infection of the human intestinal model. This study takes us forward in understanding the complete pathogenesis during the interference of SARS-CoV-2 in the intestines of COVID-19 patients.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Single-cell analyses reveal SARS-CoV-2 interference with intrinsic immune response in the human gut, Sergio Triana, Camila Metz, Zumaran, Carlos Ramirez, Carmon Kee, Patricio Doldan, Mohammed Shahraz, Daniel Schraivogel, Andreas R. Gschwind, Lars M. Steinmetz, Carl Herrmann, Theodore Alexandrov, Steeve Boulant, Megan, Lynn Stanifer, bioRxiv 2020.10.21.348854; doi: https://doi.org/10.1101/2020.10.21.348854, https://www.biorxiv.org/content/10.1101/2020.10.21.348854v1
- Peer reviewed and published scientific report.
Triana, Sergio, Camila Metz‐Zumaran, Carlos Ramirez, Carmon Kee, Patricio Doldan, Mohammed Shahraz, Daniel Schraivogel, et al. 2021. “Single‐Cell Analyses Reveal SARS‐CoV‐2 Interference with Intrinsic Immune Response in the Human Gut.” Molecular Systems Biology 17 (4). https://doi.org/10.15252/msb.202110232. https://www.embopress.org/doi/full/10.15252/msb.202110232