A team of US-based scientists from the University of Chicago, NCH Healthcare System and the University of South Florida has identified a specific set of genes associated with a poor prognosis of coronavirus disease 2019 (COVID-19). The study is currently available on the bioRxiv* preprint server.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative pathogen of COVID-19, has high infectivity and pathogenicity. Although a substantial proportion of SARS-CoV-2-infected individuals remain asymptomatic or mildly symptomatic, certain groups of people, such as elderly people or people with comorbidities, are highly susceptible to developing severe COVID-19.
Since the emergence of the COVID-19 pandemic, many studies have been conducted to identify molecular and genetic risk factors that are associated with disease severity. In this context, elevated blood levels and certain pro-inflammatory biomarkers, such as ferritin, C-reactive proteins and procalcitonin, have been shown to predispose COVID-19 patients to develop critical illness that requires mechanical ventilation. However, more studies are required to identify and characterize reliable biomarkers that can prioritize the best treatment options according to disease severity.
Current study design
According to the available literature, COVID-19 survivors are susceptible to developing pulmonary fibrosis, which is a pulmonary disease caused by damage and scarring of the lung tissue. Given this fact, the scientists had repurposed a 52-gene signature, which they had developed previously to predict the mortality risk of pulmonary fibrosis patients and to predict the clinical consequences of SARS-CoV-2 infection.
Of a set of 52 genes, they analyzed the expression of 50 genes using peripheral blood samples collected from SARS-CoV-2 patients. Moreover, they conducted single-cell expression analysis to identify the cellular sources of these genes. They excluded two non-coding RNAs as they were unavailable in some of the datasets.
Important observations
Of 50 studied genes, 7 showed increased expression, and 43 showed reduced expression. Using a classification algorithm, the scientists identified low-risk and high-risk genomic profiles. They observed that people with a high-risk genomic profile were significantly older and had a higher risk of developing severe COVID-19. Moreover, the need for ICU admission or mechanical ventilation was significantly higher among them. The blood levels of lactate, C-reactive protein, and d-dimer were also significantly higher in people with a high-risk genomic profile compared to those with a low-risk genomic profile.
To identify the cellular sources of 50 genes, they conducted patient-level analysis using the classification algorithm, which had identified five low-risk and three high-risk genomic profiles. By comparing the proportions of different cell types between low-risk and high-risk profiles, they observed that people with high-risk profiles displayed higher proportions of red blood cells, monocytes, neutrophils, and dendritic cells. In contrast, higher proportions of T and B lymphocytes, natural killer cells, and antibody-producing plasmablasts were found in people with low-risk genomic profiles.
To further validate the data, they conducted cell-type-specific analysis using the classification algorithm, which had identified 46 high-risk and 109 low-risk genomic profiles. The findings revealed that red blood cells, neutrophils, monocytes, eosinophils, and dendritic cells comprised the high-risk profiles, whereas T and B lymphocytes, natural killer cells, and plasmablasts comprised the low-risk profiles. These findings indicate that T cell- and B cell-mediated robust immune responses may reduce the risk of developing a severe form of COVID-19.
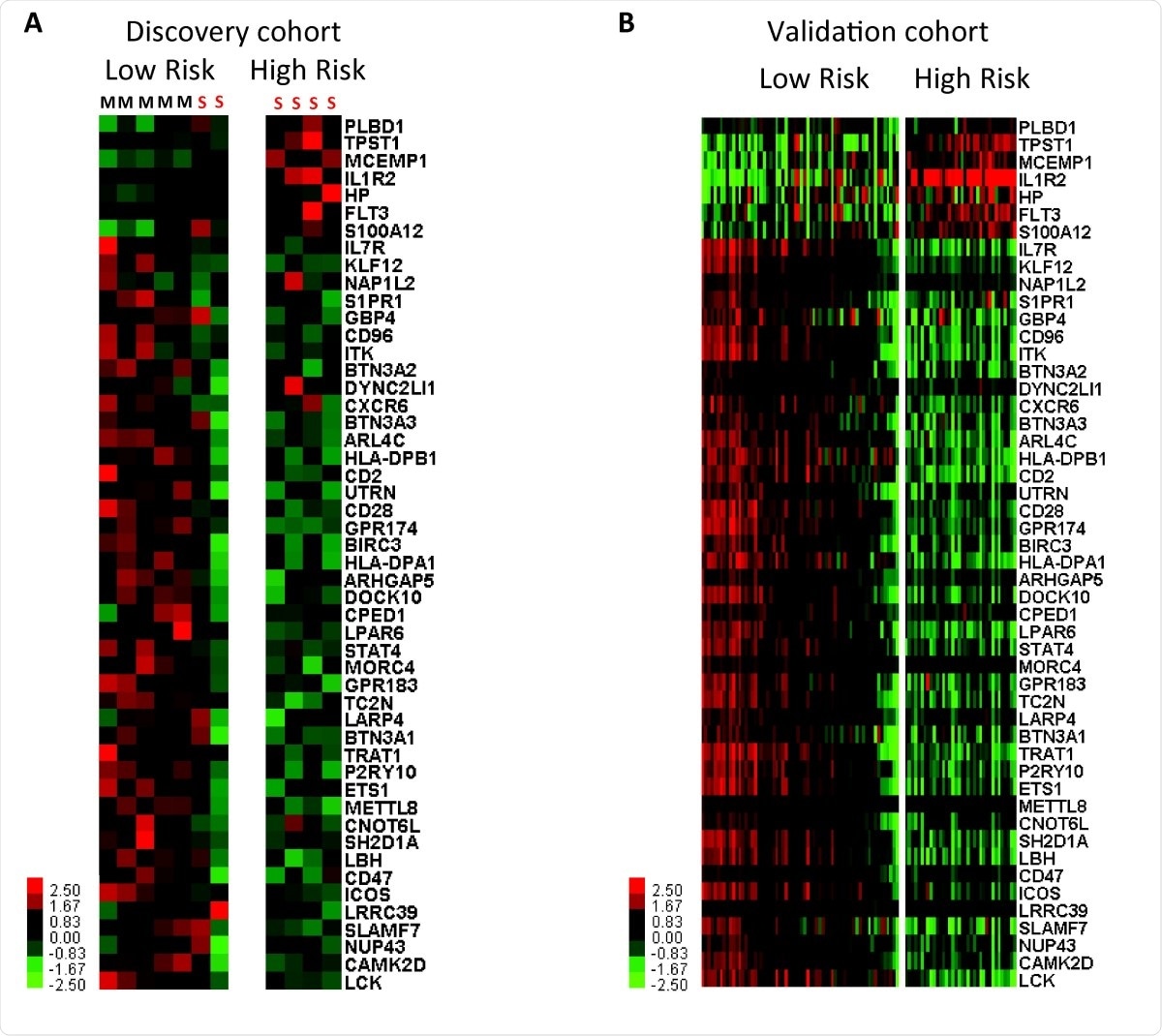
Genomic risk profiles based on the 50-gene signature are predictive of poor outcomes in SARSCoV- 2. Clustering of SARS-CoV-2 infected individuals based on genomic risk profiles (high vs low) derived from the 50-gene signature using SAMS in discovery (A) and validation cohorts (B). Every column represents a subject and every row represents a gene. Log-based two color scale is shown next to heatmaps; red denotes increase expression over the geometric mean of samples and green, decrease. S=SARS-CoV-2 severe disease. M=SARS-CoV-2 mild disease.
Study significance
The study findings indicate that the expression profiles of 50 genes in the peripheral blood can predict the clinical outcome of COVID-19 patients. The study also identifies the cellular sources of these genes. Taken together, important biomarkers identified in the study can be used to categorize COVID-19 patients based on their disease severity, which in turn can facilitate proper utilization of medical resources, including therapeutics and life-saving equipment. This can also reduce the risk and cost of unnecessary hospitalization.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Juan-Guardela B. M. (2020) A repurposed, blood gene signature is associated with poor outcomes in SARS-CoV-2. bioRxiv preprint server. doi: https://doi.org/10.1101/2020.11.21.392670,
https://www.biorxiv.org/content/10.1101/2020.11.21.392670v1
- Peer reviewed and published scientific report.
Juan Guardela, Brenda M., Jiehuan Sun, Tong Zhang, Bing Xu, Joseph Balnis, Yong Huang, Shwu-Fan Ma, et al. 2021. “50-Gene Risk Profiles in Peripheral Blood Predict COVID-19 Outcomes: A Retrospective, Multicenter Cohort Study.” EBioMedicine 69 (July): 103439. https://doi.org/10.1016/j.ebiom.2021.103439. https://www.thelancet.com/journals/ebiom/article/PIIS2352-3964(21)00232-2/fulltext.