A recent computational study from India implicated a dysregulated vitamin D pathway in the pathobiology of the infection with the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) – and opened the door for experimental validation of these observations. The paper is currently available on the bioRxiv* preprint server.
A major public health calamity due to the coronavirus disease (COVID-19), which is a life-threatening disease caused by SARS-CoV-2, mobilized the global research community in deciphering fundamental mechanisms and cellular pathways that are responsible for symptom variety, disease severity and progression, as well as recovery or death.
Vitamin D is a steroid hormone with manifold cellular functions; albeit a defective vitamin D pathway has been linked to SARS-CoV-2 pathobiology, the status of vitamin D-modulated genes in lung cells of infected patients remains undetermined.
In short, cellular effects of vitamin D stem from its nongenomic/cytoplasmic effects, but also its genomic/nuclear effects. Consequently, a myriad of receptors, enzymes and factors are implicated in how it modulates inflammation and feedback loops.
This is why the researchers, led by Dr. Bijesh George from the Rajiv Gandhi Centre for Biotechnology (Trivandrum) and Manipal Academy of Higher Education in India, conducted a study to appraise the exact status of the vitamin D pathway in SARS-CoV-2-infected patients by using public transcriptomic datasets and stringent computational approaches.
Cell lines, transcriptomic approaches and network analysis
The researchers initially assessed the expression levels of core components of the vitamin D pathway in different models of viral infection by utilizing the Signaling Pathways Project Datasets (SPPD) – an assemblage of transcriptomic datasets biocurated by the Nuclear Receptor Signaling Atlas (NURSA) Organization.
Subsequently, the lung cells of COVID-19 patients were analyzed to evaluate core molecules of the vitamin D pathway in three separate RNA sequencing-based transcriptomic datasets of bronchoalveolar lavage fluid (BALF) cells, but also in A549, Calu3 and NHBE human lung cell lines expressing SARS-CoV-2.
Finally, the network analysis of various sets of genes has been performed with the use of the Network Analyst 3.0 tool. Moreover, functional enrichment analysis has been done by using Reactome, a freely accessible online pathway database with intuitive bioinformatics tools.
Reduced expression of vitamin D pathway components
In short, the results of this study support the notion of a purported association between SARS-CoV-2 infection and reduced expression of different components of the vitamin D pathway. More specifically, there was a reduction of vitamin D and retinoid X receptors, as well as CYP27A1 (belonging to cytochrome P450 gene family) in BALF cells of patients infected with the virus.
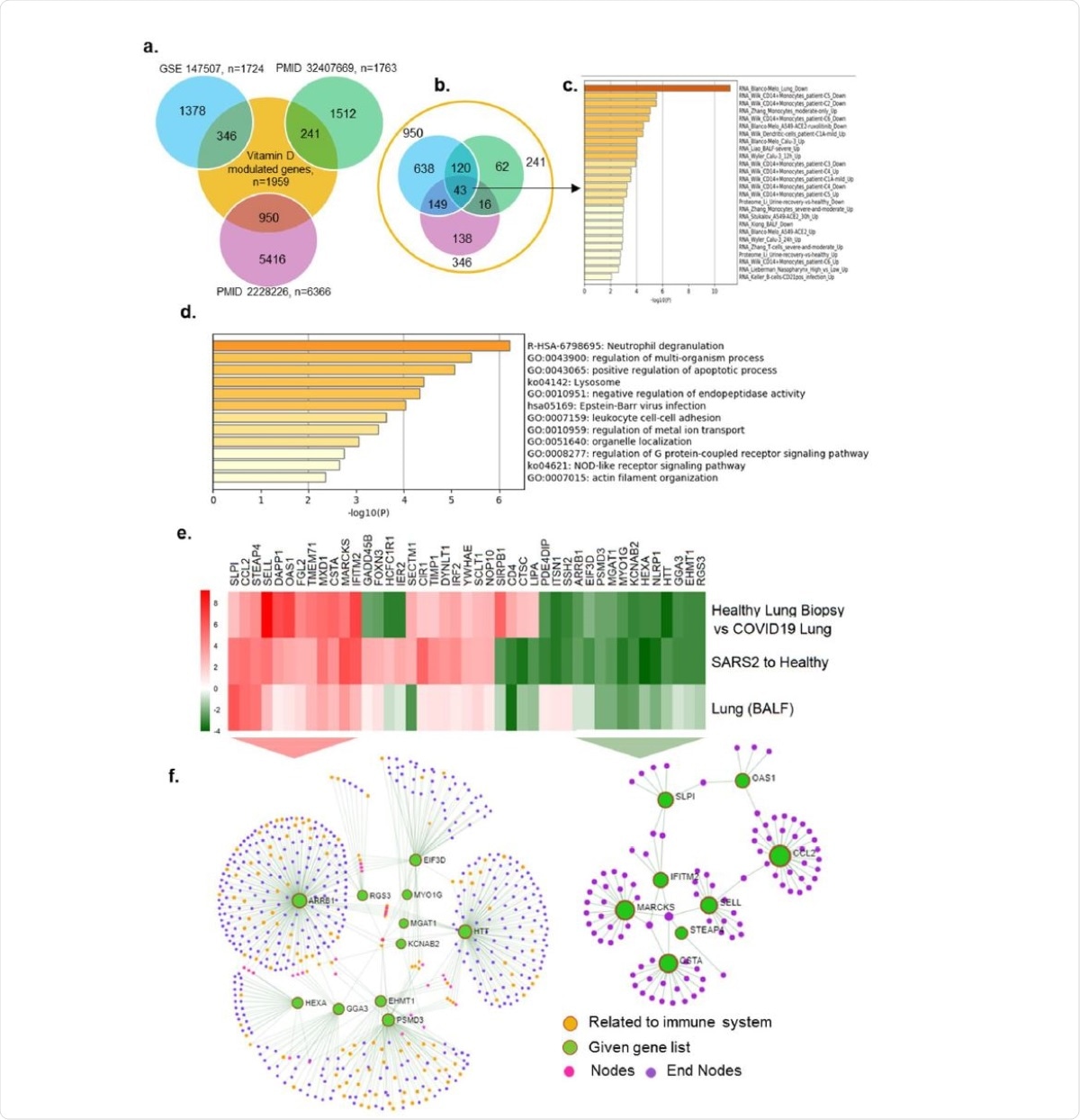
Expression of Vitamin D regulated genes in COVID19 patients. a) Number of genes found overlapped with Vitamin D regulated genes and three SARS-COV-2 datasets are denoted. b) 43 genes were identified which are common for 3 SARS-COV-2 datasets and also regulated by Vitamin D, c) Bar diagram shows the summary of enrichment analysis in SARS-COV-2 from Metascape. d) Bar diagram shows the summary of functional enrichment analysis in from Metascape. e) The expression status of these 43 genes in SARS-COV-2 patient data is plotted as heat map. f) Network analysis of 12 genes found up-regulated in all three patient dataset and also regulated by Vitamin D is given. The functional enrichment analysis of 12 genes using Reactome identified “Activation of the pre-replicative complex” as primary enrichment (rank 1, hits 9/32 pavlue 3.5e-11), Network analysis of 12 genes downregulated in all three SARS-COV-2 patient transcriptome datasets. Genes are found functionally related to immune system (rank 57, 131/1140 pvalue 1.31e-21) are remarked.
Furthermore, by identifying 107 differentially expressed and chiefly down-regulated genes modulated by vitamin D in transcriptomic datasets from patients’ cells, the role of direct and indirect mechanisms of gene expression by the dysregulated vitamin D has been established.
A detailed network analysis of such differentially expressed genes revealed immune system pathways, NF-kB/cytokine signaling, and cell cycle regulation as prime predictive events that may be affected in the cells of such patients.
Another novel notable observation is the upregulation of PAK1 expression (but not another family of survival AKT kinases) in the same SARS-CoV-2 sample sets with a reduced expression of vitamin D pathway components, suggesting a potential correlative relationship between these two phenomena", say the study authors.
A step towards the experimental validation
Everything said, it is clear how this study provides important insights about a potentially causal association between a compromised vitamin D pathway on many layers with SARS-CoV-2 infections in patients, resulting in a dysregulation of pathways downstream to vitamin D.
This dysregulation is characterized by both down-regulation and up-regulation of vitamin D cellular genes, which entails molecules that are involved in proinflammatory Th1 response and immune regulation.
These preliminary results now set the stage for experimental validation of the observations and postulations made using computational approaches," conclude study authors based on the results of their study.
In any case, the stage is set by this research endeavor for larger studies that will confirm whether a compromised vitamin D pathway can indeed influence the susceptibility of lung cells to SARS-CoV-2, and elucidate its exact role in the protean manifestations of COVID-19.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources