Study: Comprehensive mapping of mutations to the SARS-CoV-2 receptor-binding domain that affect recognition by polyclonal human serum antibodies. Image Credit: NIAID
Such antigenic evolution is prompted by positive selection for mutations in the viral spike glycoprotein – most notably in regions that are implicated in receptor binding. Hence, in order to trace similar occurrences in SARS-CoV-2, it is pivotal to appraise which viral mutations have the biggest impact on human polyclonal antibody immunity.
However, studies conducted thus far provided an incomplete view of all possible mutations, so we are basically left without the complete description of the possible effects of viral mutations on recognition by polyclonal antibodies in the human sera.
In this new study, a research group led by Dr. Allison J. Greaney from the University of Washington in Seattle decided to comprehensively map how all amino-acid mutations to the RBD of the SARS-CoV-2 spike glycoprotein affect binding by the plasma antibodies in individuals that had COVID-19.
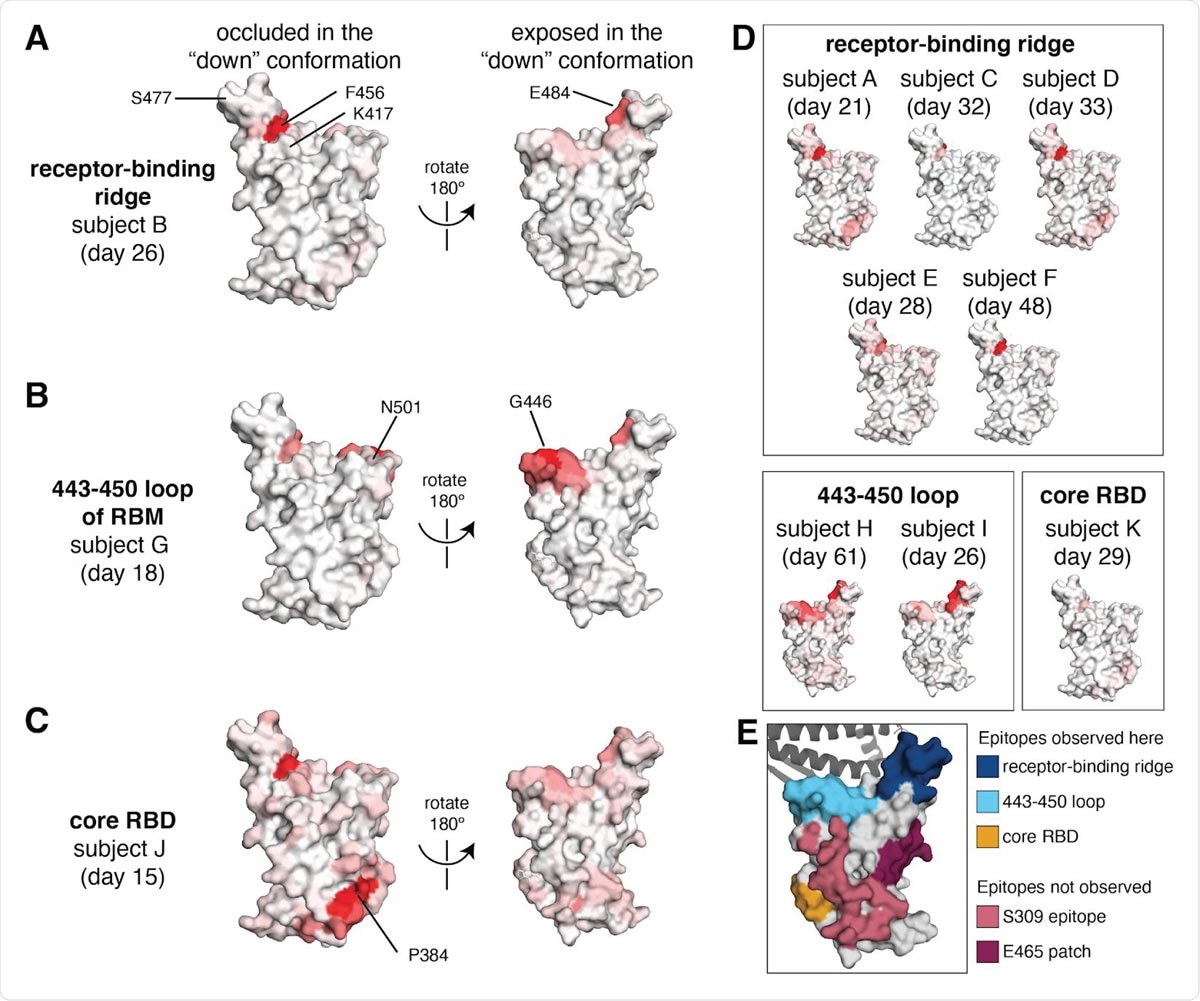
Regions of the RBD where mutations strongly reduced binding by the antibodies in sera collected from 11 individuals. The total effect of mutations at each site (sum of escape fractions) are projected onto the structure of the RBD (PDB 6M0J), with white indicating no effect of mutations at that site and red indicating a large reduction in antibody binding. Two views of the RBD are shown: the surface of the RBD that is buried in the “down” conformation, and the surface that is always exposed and accessible (Walls et al., 2020; Wrapp et al., 2020) . (A) For some individuals (typified by subject B), antibody binding is predominantly reduced by mutations in the receptor-binding ridge, particularly at sites F456 and E484. (B) For some individuals (typified by subject G), antibody binding is strongly reduced by mutations in the 443–450 loop of the RBM in addition to the receptor-binding ridge. (C) For a few individuals (typified by subject J), antibody binding is affected by mutations in the core RBD epitope around site P384. (D) Samples from the other eight individuals fall in one of the three classes detailed in panels (A) to (C) . For panels (A) to (D) , the white-to-red coloring scale is set to span the same range as the y-axis limits for that serum in Figure 2 . (E) Mutations in two major surface regions (the S309 epitope and the sites near E465) do not strongly affect serum antibody binding for any of the subjects. Shown is a surface representation of the RBD, with the 3 polyclonal serum epitopes colored as in Figure 2 . The S309 epitope and region near E465 (“E465 patch”) are shown in pink and maroon. ACE2 is shown in a dark gray cartoon representation.
Comprehensive binding studies
In a nutshell, the researchers characterized serum antibodies from 35 plasma samples, which were longitudinally collected from 17 different SARS-CoV-2-infected individuals approximately 1 to 3 months after their symptom onset. RBD-conjugated beads were used to deplete RBD-binding antibodies, and the neutralizing activity was compared before and after such depletion.
Then they have appraised how depletion of RBD-binding antibodies affected total serum binding to the ectodomain of spike glycoprotein but also measured how it affected the neutralization of lentiviral particles pseudotyped with the G614 variant of the SARS-CoV-2 spike glycoprotein.
Furthermore, to thoroughly map RBD mutations that have the propensity to reduce binding by polyclonal serum antibodies, the researchers have expanded a deep-mutational scanning technique previously utilized to pinpoint mutations that elude binding by monoclonal antibodies.
For the assays with spike-pseudotyped lentiviral particles, mutations with substantial effects on serum antibody binding (according to mapping) were selected, while mutations present in circulating strains of SARS-CoV-2 were prioritized.
Mutations in three major epitopes
This study has shown that binding by polyclonal serum antibodies is affected by mutations arising in three main epitopes in the RBD of the spike glycoprotein; however, a substantial variation in the impact of mutations was noted, both among individuals and within the same person over a period time.
But despite such heterogeneity, the mutations that substantially reduce antibody binding were most often found at just a few sites in the RBD's receptor-binding motif. The paramount site was E484, where neutralization by certain sera was reduced more than 10-fold by several mutations – including the one in emerging viral lineages in Brazil and South Africa.
Another major epitope was basically centered on the loop formed by the amino acid residues 443–450 in the RBD's receptor-binding motif, with mutations in this epitope at times strongly affecting serum antibody neutralization. A third epitope, distal from the receptor-binding motif, had much smaller effects.
Implications for surveillance and vaccinology
"The strong contribution of RBD-binding antibodies to serum neutralization demonstrates that mapping mutations that escape these antibodies is crucial for understanding the potential for SARS-CoV-2 antigenic evolution", say study authors in this bioRxiv paper.
The comprehensive nature of this type of mapping endeavor opens the door for the ongoing assessment (i.e., surveillance) of which circulating RBD mutations will expectedly have the greatest impact on human immunity, but also elucidates broader features of antibody immunity relevant to SARS-CoV-2 evolution.
All of this represents an important area for future research, as we need to understand how viral mutations impact vaccine-elicited immunity. Subsequently, this knowledge may be used to tailor vaccines that are rather robust to viral antigenic evolution.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Greaney, Allison J., Andrea N. Loes, Katharine H.D. Crawford, Tyler N. Starr, Keara D. Malone, Helen Y. Chu, Jesse D. Bloom. 2021. “Comprehensive Mapping of Mutations in the SARS-CoV-2 Receptor-Binding Domain That Affect Recognition by Polyclonal Human Plasma Antibodies.” https://www.biorxiv.org/content/10.1101/2020.12.31.425021v1.
- Peer reviewed and published scientific report.
Greaney, Allison J., Andrea N. Loes, Katharine H.D. Crawford, Tyler N. Starr, Keara D. Malone, Helen Y. Chu, Jesse D. Bloom. 2021. “Comprehensive Mapping of Mutations in the SARS-CoV-2 Receptor-Binding Domain That Affect Recognition by Polyclonal Human Plasma Antibodies.” Cell Host & Microbe, February. https://doi.org/10.1016/j.chom.2021.02.003. https://www.cell.com/cell-host-microbe/fulltext/S1931-3128(21)00082-2.