The currently circulating severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has taken an enormous toll on human life, health and economic activity all over the world. Despite the recorded mortality of over 2.4 million, the pathogenesis of this virus remains unclear.
A new study by researchers at the University of Connecticut Health Center, USA, reports the crucial nature of the pathogen pattern recognition receptor (PRRs), especially the two RLRs, retinoic acid-inducible gene I (RIG-I) and melanoma differentiation-associated protein 5 (MDA5), in triggering host cell immune responses within the cell. The RLRs are viral RNA sensors.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
The team released the findings of their study on the bioRxiv* preprint server.
The importance of RLRs
The innate immune response to a viral infection is initiated by pathogen pattern recognition receptors (PRR), including C-type lectins, Toll-like receptors (TLR), retinoic acid-inducible gene I (RIG-I) like receptors (RLR), and nucleotide-binding oligomerization domain (NOD)-like receptors (NLR).
RLRs in the cytoplasm bind to viral RNA and subsequently undergo translocation. This complex binds to mitochondrial antiviral signaling protein (MAVS), activating a signaling pathway that eventually results in the transcription of host immune genes, including the key interferon (IFN) genes that are the rapid responders in this situation.
With RNA viruses, RIG-1 and MDA5 RLRs are the pivotal PRRs, but show differential preferences with different RNA viruses. The role they play in SARS-CoV-2 infection is unclear, and is the focus of the current preprint.
Parallel infection by SARS-CoV-2 in cell lines expressing RIG-1, MDA5 and MAVS, as well as in knockout cells where each of these was precisely silenced, was achieved, and the IFN responses evaluated.
Viral load
The researchers found that the viral RNA load within the knockout cells was higher by 2-3.5-fold in all cases, by 24 hours following infection. By 72 hours, the viral RNA load was higher by 5-12-fold. Extracellular viral titers were also much higher in the media surrounding knockout cells than the wildtype cells, of two different cell lines.
Antiviral immune responses
The study also showed a steady increase in the mRNA encoding IFNB1 and IL29 (a type III IFN), as well as the interferon-stimulated gene (ISG15), over time in infected wildtype cells, but not to the same extent in MDA5 and MAVS knockout cells.
The supernatants from these cells also showed lower levels of IFN-β and C-X-C motif chemokine ligand 10 (CXCL10) proteins relative to wildtype cells. Interestingly, type II and type III IFNs, as well as ISG15, were expressed at higher levels in RIG-1 knockout cells. This may suggest that the presence of these receptors is correlated with an independent antiviral mechanism, unlike the IFN-dependent mechanism operating in the other cases.
Similarly, the host cell receptor for the virus, angiotensin-converting enzyme 2 (ACE2), showed a constant upregulation over the course of infection in the wildtype cells. Even so, RIG-1 knockout cells had 2.5-fold higher mRNA levels compared to other knockout cells or in the wildtype cells. This may mean that this system of receptors suppresses the expression of ACE2.
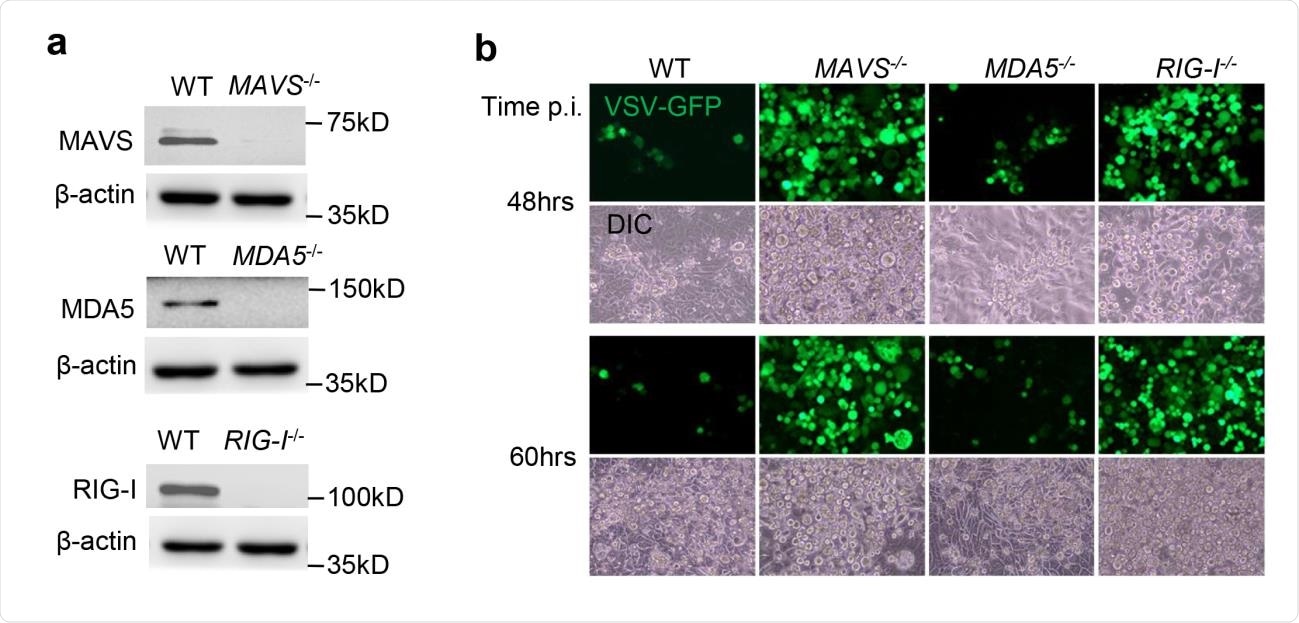
Functional validation of gene knockouts by CRISPR-Cas9. a) The immunoblots show gene knockout efficiency in Calu-3 cells. ß-actin is a house keeping gene and serves as a protein loading control. b) Fluorescent microscopic images of VSV-GFP at several time points post infection (p.i.). Magnification: 100 x. DIC: differential interference contrast. The results are representative two reproducible independent experiments.
RLRs in lung epithelium
The investigators then attempted to understand the major PRR pathways in the lung epithelium, being the primary target of SARS-CoV-2, and thus the first location for host antiviral responses. It is noteworthy that RLRs function in all cells and tissues, unlike the TLRs, which perform the same viral RNA-sensing function but only within immune cells.
They found that in lung epithelium, MDA5 is the major RLR signaling system, but its knockout, and that of MAVS, did not completely prevent the induction of IFNs. Thus, other PRRs may also play a part in antiviral signaling.
RIG-1 knockout left IFN responses intact in SARS-CoV-2 infection, but failed to impact rising viral replication rates. This supports the independent activity of RIG-1 against viral infection, rather than via either MAVS or IFNs.
Here again, ACE2 expression was 2.5 times higher in RIG-1 knockout cells at one hour post-infection, relative to either wildtype and other RLR-knockout cells.
Possible mechanisms of RIG-1
This suggests the possibility that the antiviral activity of RIG-1 may comprise inhibition of viral entry into the host cell by preventing the expression of its receptor, ACE2.
Another explanation could be that RIG-1 has a broad spectrum of antiviral functions in the absence of normal IFN responses, such as initiating antiviral effector expression, via a signaling pathway independent of IFN, Janus kinase (JAK) and signal transducer and activator of transcription (STAT) activity.
Such mechanisms include preventing the normal binding of hepatitis B polymerase with viral RNA, which in turn prevents viral replication. With influenza A, the binding of RIG-1 to the nucleocapsid may suppress replication by a direct mechanism.
What are the implications?
With the current virus, both RIG-1 and MDA5 may displace already bound viral proteins from their locations on double-stranded (ds) DNA, thus proving their antiviral activity to be broadly inhibitory against many viruses, while being independent of either MAVS or IFNs.
While a recent study has shown that RIG-1 may not be vital in suppressing SARS-CoV-2 infection, the above findings are discordant, perhaps due to the different methods used.
Given that MDA5 appears to be central in sensing viral RNA and activating the immune response, and that RIG-1 seems to have IFN-independent antiviral activity, the researchers suggest exploring MDA5 mimetics for therapeutic efficacy against the SARS-CoV-2 infection.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources