The ongoing coronavirus disease 2019 (COVID-19) pandemic, caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), is thought to be a disease originally occurring in animals but transmitted across the species barrier to human beings, probably through an intermediate host. Following human infection, the virus has been shown to transmit to some pet and farm animals, where it then mutates. A new preprint research paper posted to the bioRxiv* server shows that these mutations disrupt inhibition by neutralizing antibodies, both natural and therapeutic.
The American mink is a profitable business focus, farmed for its fur in the Netherlands and many other countries. The first case of SARS-CoV-2 infection in farmed mink on a Dutch farm was identified in April 2020. The application of whole-genome sequencing showed that the virus was acquired from humans.
The infection spread back into farmworkers from the infected animals and then from worker to worker. Similar data were reported from Denmark and then from other countries in Europe, involving both farmed and range-bred mink. In Denmark, this led to the culling of 17 million farmed mink.
The SARS-CoV-2 enters the host cell via its spike protein binding with the angiotensin-converting enzyme 2 (ACE2), the host cell receptor. The spike protein binds via its receptor-binding domain (RBD) to the receptor and depends on the host protease TMPRSS2 for its priming.
Study aims
The virus in farmed mink has a different genomic composition compared to that in humans, chiefly a deletion of 69/70 in the N-terminal domain of the spike protein, and a combination of Y453F in the RBD, with I692V, M1229I, and S1147L in various parts of the spike protein. The first four have been found to occur together in one variant called the cluster 3 variant.
The current study sought to show whether the Y453F mutation causes changes in the spike protein expression, altered binding to the host cell receptor, and susceptibility to neutralizing antibodies.
The researchers used SARS-CoV-2 spike-expressing vesicular stomatitis virus particles. The spike expressed was of three variants, one expressed by wildtype SARS-CoV-2 and the second with a glycine at position 614, to be the reference for spike variants with the D614G mutation that is now globally dominant. The third was spike protein bearing the mutations found in SARS-CoV-2 isolated from infected mink.
Efficient expression and processing of spike protein
The researchers found that all the mutations in the mink variant allowed the virus to incorporate and process the spike protein at the novel furin cleavage site, at the interface between the S1/S2 subunits of the spike. All three variants achieved efficient binding to the human ACE2 receptor.
All spike variants could mediate viral entry into commonly used cell lines, but the D614G variant showed more efficient entry, in agreement with earlier studies. The combination of the Y453F mutation with either D614G, or with both H69Δ, H70Δ and D614G, did not impair the efficiency of viral entry.
The combination of D614G with the four cluster 5 mutations was associated with reduced efficiency of cell entry into several cell lines, but not the human intestinal cell line Caco-2 and the lung cell line Calu-3. This indicates that the mink variants can infect human intestinal and lung cells efficiently.
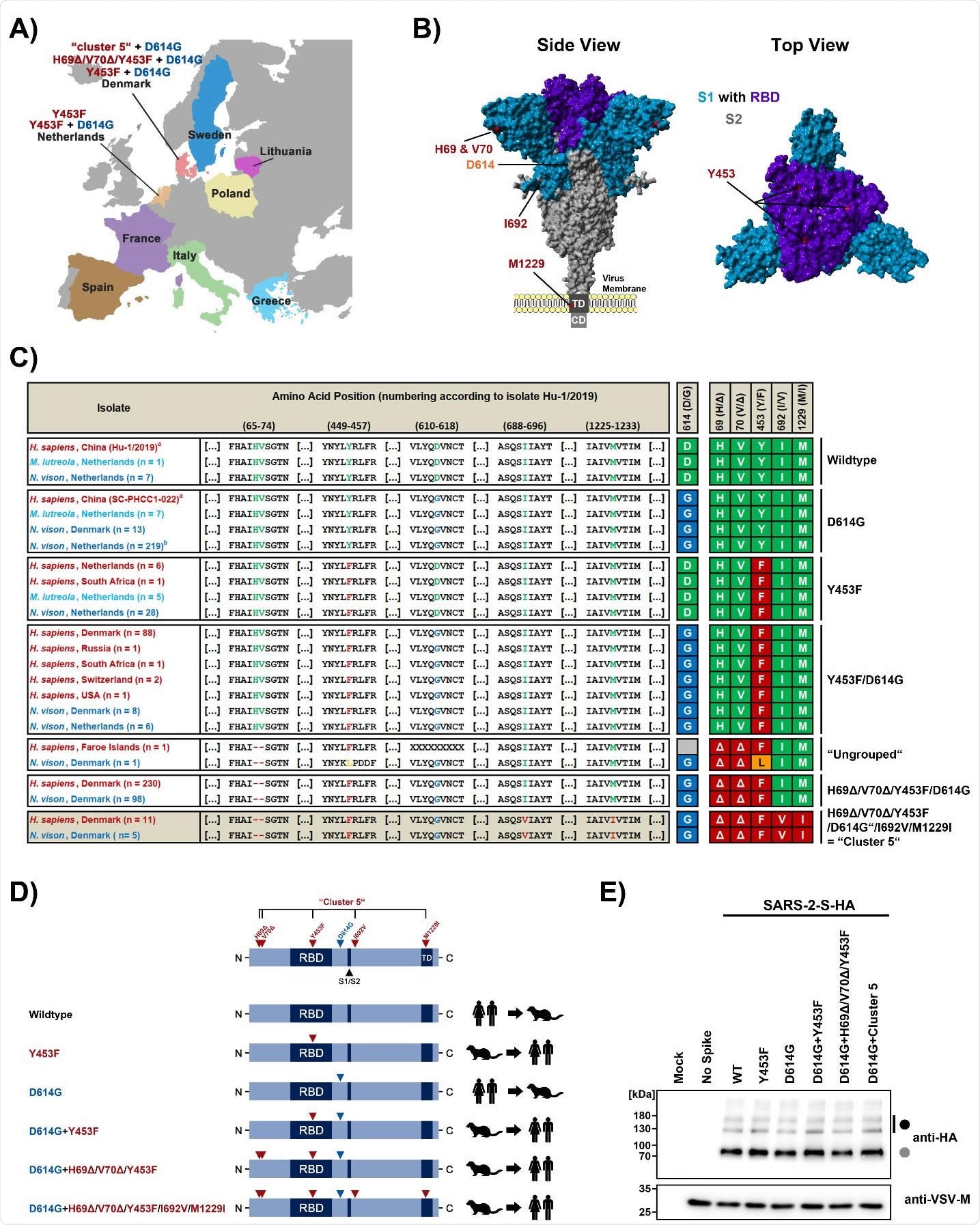
Mink-specific spike protein variants are robustly expressed, proteolytically processed and incorporated into viral particles. (A) European countries that have reported SARS-CoV-2 infection in mink. The mink-specific spike (S) protein mutations under study are highlighted. (B) Summary of mink-specific S protein mutations found in human and mink SARS-CoV-2 isolates. Sequences were retrieved from the GISAID (global initiative on sharing all influenza data) database. Legend: a = reference sequences, b = 36/219 sequences carry additional L452M mutation; Abbreviations: H. sapiens = Homo sapiens (Human), N. vison = Neovison vison (American Mink), M. lutreola = Mustela lutreola (European Mink). (C) Location of the mink-specific S protein mutations in the context of the 3-dimensional structure of the S protein. (D) Schematic illustration of the S protein variants under study and their transmission history. Abbreviations: RBD = receptor binding domain, S1/S2 = border between the S1 and S2 subunits, TD = transmembrane domain. (E) Rhabdoviral pseudotypes bearing the indicated S protein variants (equipped with a C-terminal HA-epitope tag) or no viral glycoprotein were subjected to SDS-PAGE under reducing conditions and immunoblot in order to investigate S protein processing and particle incorporation. Detection of vesicular stomatitis virus matrix protein (VSV-M) served as loading control. Black and grey circles indicate bands for unprocessed and processed (cleavage at S1/S2 site) S proteins, respectively. Similar results were obtained in four separate experiments.
Inhibition by soluble ACE2 and Camostat
The study also found that spike-bearing viruses could not infect cells in the presence of either soluble ACE2, which acts as a viral decoy, or (with Calu-3 cells) Camostat, which is a TMPRSS2 inhibitor. The most susceptible were those bearing the D614G + cluster 5 mutations, indicating that these agents could still inhibit the mink variants.
Y453F mutation affects neutralization by antibodies
The investigators also found that 13/14 sera from COVID-19 patients produced potent neutralization of spike-driven entry into the host cell. The last serum showed moderate neutralization.
With the Y453F mutation, most samples showed reduced inhibition, with the serum antibody titer required for 50% neutralization (NT50) rising to three-fold higher values. This could mean that the RBD mutation renders the SARS-CoV-2 variant resistant to neutralizing antibodies induced by viral variants in circulation at present.
The mutation also modulated inhibition by one antibody (Casirivimab/REGN10933) out of a double-antibody cocktail (the REGN-COV2 antibody cocktail) that has received emergency use authorization (EUA) for COVID-19 treatment. This supports the finding that this mutation occurs at the interface of binding of the spike protein and the antibody.
What are the implications?
This mutation reduced the efficacy of both natural and therapeutic antibodies in the mink variant. This shows that with viral entry into other species, adaptive changes are likely to arise in the viral genome to promote efficient host cell entry.
The same mutation has been shown to arise during experimental SARS-CoV-2 infection of ferrets, which may indicate the viral RBD adaptation to ferret ACE2. Another possibility is that this is an immune escape mutation, supported by the emergence of this mutation in a patient with chronic SARS-CoV-2 infection.
The cluster 5 variant mutations were associated with reduced human cell entry by the virus, which may explain the rapid extinction of this isolate in humans. It may also have had reduced ACE2 binding affinity, which was responsible for its higher vulnerability to inhibition by soluble ACE2.
The study suggests that "at least in a fraction of patients antibody responses induced upon infection and potentially also vaccination might provide only incomplete protection against infection with 200 SARS-CoV-2 amplified in mink." However, high antibody titers may provide complete protection against the mink variant as well.
The mink variant can be passed on to humans and mink in the wild, which raises the possibility of a permanent reservoir for such viruses in the wild and the emergence of new variants that may cause future outbreaks among both wild animals and humans.
Further studies will be required to understand how far specific cellular immunity against the virus is affected by this mutation and its effect on binding to mink ACE2. Nonetheless, the findings indicate that the virus's adaptation to mink infection involved the acquisition of mutations that encourage immune evasion in human infection.
This adds urgency to the efforts to prevent the transmission of human infection to mink and other animals. Secondly, the changes that occur and the possibility of amplification of the virus following such transmission to animals should be identified, as well as their effect on the crucial biological properties of SARS-CoV-2.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources