New York has the first case of a new severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variant called B.1.526, picked up by genomic surveillance research from Columbia University. The findings come after reports of other variants have been reported worldwide — B.1.1.7 in the United Kingdom, B.1.351 in South Africa, and P.1/P.2 in Brazil. If confirmed, the new variant would be the second COVID variant found in the United States after B.1.427/B.1.429 was discovered in California.
Evolving new variants such as B.1.1.7 have the N501Y mutation on the spike protein, allowing it to be more contagious.
Of concern is the efficacy of the mRNA coronavirus vaccines as they were developed before the variants' emergence. The B.1.351 variant has the E484K mutation that makes it antibody-resistant.
The researchers suggest the B.1.526 variant shares similar mutations to the other reported variants, including the antibody-resistant E484K mutation.
The study "A Novel SARS-CoV-2 Variant of Concern, B.1.526, Identified in New York" is available as a preprint on the medRxiv* server, while the article undergoes peer review.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Genomic surveillance of B.1.526
The researchers performed rapid PCR-based assays to detect coronavirus variants with the N501Y and E484K mutation from 1,142 randomly selected samples of confirmed positive severe acute respiratory syndrome (SARS-CoV-2).
The samples came from the Columbia University Biobank, and patient information was collected from the COVID-Care database.
Increase in E484K-Positive Cases
Of the 1,142 samples, 83 (9%) contained the E484K mutation. The earliest SARS-CoV-2 case with the E484K mutation was collected in mid-November 2020. Since then, the researchers found an increase in E484K-positive cases.
In early November, 1.3% of E484K-positive cases were detected. This number grew to 5.3% by the middle of January 2021 and 12.3% between February 8th and 15th.
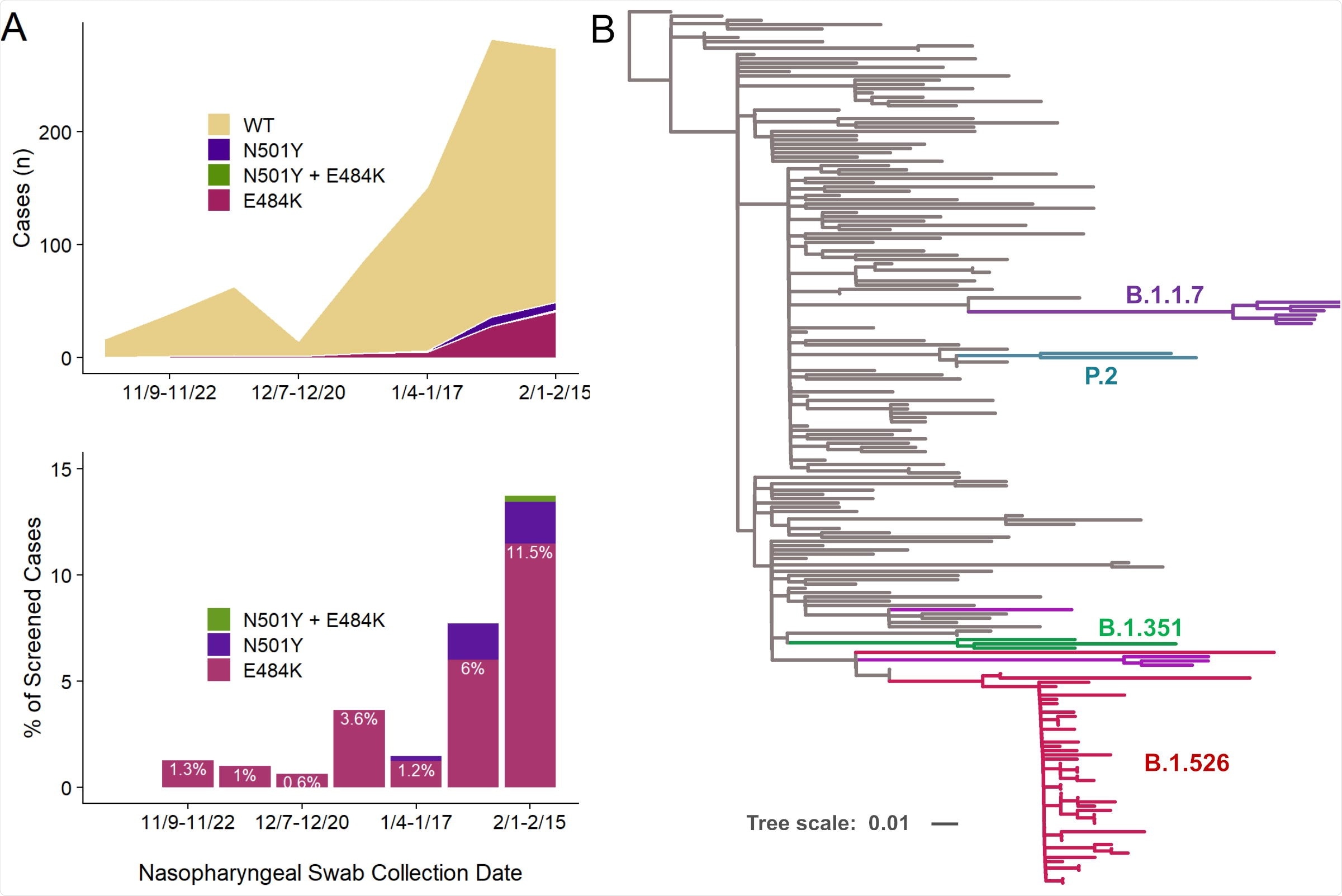
Increasing prevalence of E484K from November 2020 to February 2021 including E484K-harboring isolates within the B.1.526 lineage. (A) The earliest detected E484K variant from our study period was collected in mid-November 2020, and a total of 83 isolates with this substitution were identified between November 2020 and February 15, 2021, out of 927 total samples that showed a signal in our screening assay (upper panel). The prevalence of E484K (nE484K/(nWT + nE484K)) subsequently increased over time, from 1.3% in early November (11/09/2020-11/22/2020) to 11.5% in early February (2/1/2021-2/15/2021), as shown in the lower panel. (B) To place our E484K isolates within the context of globally circulating SARSCoV- 2 strains, we downloaded 140 genomes from GISAID randomly sampled across 50 distinct lineages. Phylogenetic analysis demonstrated that the majority of our B.1.526 isolates are tightly clustered (bottom; magenta), with the remaining E484K-harboring isolates interspersed within the parent B.1 lineage. The B.1.1.7 (UK; purple), B.1.351 (South Africa; green), and P.2 (Brazil; teal) samples identified fall within these branches, respectively, as expected, as shown. Samples from our collection which harbor E484K or N501Y but do not fall within one of these lineages are shown in magenta and purple, respectively.
Increase in N501Y-Positive Cases
A total 17 (1.8%) samples contained the N501Y mutation. Like E484K, the researchers found the number of cases with N501Y also rose over time, with the first case seen in mid-January and 2.6% of screened isolates by mid-February.
One SARS-CoV-2 positive sample had both mutations.
Detection of B.1.526 COVID Variant
The researchers performed whole-genome nanopore sequencing on 65 samples that were potentially N501Y or E484K-containing strains. They then used screening PCR assays to verify their results.
Six samples that had the N501Y mutation were from the B.1.17 variant. Two samples were of the P2 variant, which contains the E484K mutation. One sample was from the B.1351 variant, which had both mutations.
The most surprising finding was that about 75% (49 of the 65 cases) had the E484K mutation but matched closely with a different lineage called B.1.526.
The first case of the B.1.526 variant was found in November 2020 in a patient with advanced AIDs who initially had a coronavirus infection. However, this coronavirus infection did not have L5F, D253G, and E484K mutations.
The authors suggest the B.1.526 variant is the main culprit for the recently rising COVID-19 cases in New York.
Genetic composition of B.1.526
Genetic analysis of B.1.526 variants revealed several mutations on the spike protein including, L5F, T95I, D253G, E484K, D614G, and A701V.
The B.1.526 variant was tested using 4 monoclonal antibodies with emergency use authorization, 10 convalescent plasma, and 10 vaccine sera.
Results showed the B.1.526 variant also shows the ability to neutralize antibodies, making current SARS-CoV-2 treatments less effective. There was a 7.7-fold decrease in the neutralizing activity of convalescent plasma and 3.4-fold for the vaccine sera when the E484K variant was present.
Patient information
Older people were more likely to have the E484K mutation and more likely to be given general admission to the hospital or be admitted to the emergency room.
People in two diverse neighborhoods were more likely to have the E484K variants compared to people who lived in the metropolitan area
Future Concerns
When the researchers analyzed genomic data from public databases, they found approximately 140 genomes related to the B.1.526 variant spread across the Northeast. They suggest the E484K in the B.1.526 lineage is across New York state.
New York was the epicenter of the COVID outbreak in the United States. They currently rank 4th in states having over 1.6 million COVID-19 cases. America ranks first worldwide for the highest number of cases at over 28.5 million.
"In conclusion, we identified B.1.526 as a local lineage of concern due to E484K in particular, which could threaten the efficacy of current antibody therapies and vaccines. This discovery also highlights the need for a concerted national surveillance program to track and contain the spread of novel SARS-CoV-2 variants," wrote the team.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Annavajhala MK, et al. A Novel SARS-CoV-2 Variant of Concern, B.1.526, Identified in New York. medRxiv, 2021. doi: https://doi.org/10.1101/2021.02.23.21252259, https://www.medrxiv.org/content/10.1101/2021.02.23.21252259v1
- Peer reviewed and published scientific report.
Annavajhala, Medini K., Hiroshi Mohri, Pengfei Wang, Manoj Nair, Jason E. Zucker, Zizhang Sheng, Angela Gomez-Simmonds, et al. 2021. “Emergence and Expansion of SARS-CoV-2 B.1.526 after Identification in New York.” Nature 597 (7878): 703–8. https://doi.org/10.1038/s41586-021-03908-2. https://www.nature.com/articles/s41586-021-03908-2.