The immune system appears to be rapidly evolving in response to the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants circulating worldwide. A new preprint study posted to the bioRxiv* server led by Ali H. Ellebedy from the Washington University School of Medicine found that the monoclonal antibody 2C08 produced by the Pfizer vaccine could help neutralize COVID-19 variants.
Previous work investigated germinal center (GC) B cells after vaccination and found they produced antibodies specific to the receptor-binding domain to the spike protein of the D614G SARS-CoV-2 strain.
The current findings provide further evidence that vaccination is necessary for delivering lasting immunity to SARS-CoV-2 variants.
Monoclonal antibody binding to variants
The researchers used 13 human monoclonal antibodies that were effective in binding to the D614G strain. Using an ELISA, they further assessed these antibodies towards three variants, including B.1.1.7, B.1.351, and B.1.1.248.
Only one monoclonal antibody, 1H09, decreased binding to the receptor-binding domain of the B.1.1.7 variant. Four monoclonal antibodies showed significant reductions in binding activity when bound to the receptor-binding domain of the B.1.351 and B.1.1.248 variants. About eight monoclonal antibodies showed equivalent binding to all tested strains.
They next looked at the neutralizing activity of 13 antibodies when exposed to the D614G strain. About five antibodies — 2C08, 1H09, 1B12, 2B06, and 3A11 — were able to neutralize 80% of the D614G strain. When exposed to the B.1.1.7, B.1.351, and B.1.1.248 variants, 1H09 failed to neutralize any emerging variants. Monoclonal antibodies 1B12, 2B06, and 3A11 neutralized the B.1.1.7 variant but not the B.1.351 and B.1.1.248 variants.
Interestingly one antibody created from the spike-specific GC B cells developed the 2C08 antibody that neutralized D614G and several SARS-CoV-2 variants. This suggests it recognizes other residues on the receptor-binding domain that remain unchanged amongst variants.
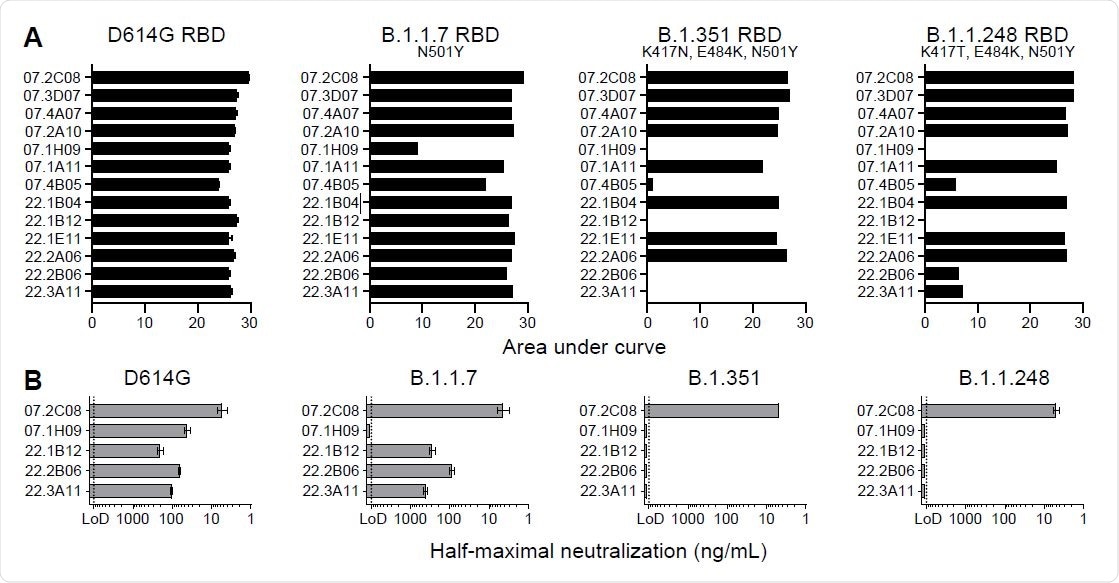
mAb 2C08 potently neutralizes diverse SARS-CoV-2 strains. (A and B) ELISA binding to recombinant RBD from (A) and neutralizing activity in Vero-TMPRSS2 cells against (B) indicated SARS-CoV-2 strains by the indicated mAbs. ELISA binding to D614G RBD previously reported in (29). Baseline for area under the curve was set to the mean + three times the standard deviation of background binding to bovine serum albumin. Dotted lines indicate limit of detection. Bars indicate mean ± SEM. Results are from one experiment performed in duplicate (panel A, D614G) or in singlet (panel A, B.1.1.7, B.1.351, and B.1.1.248), or two experiments performed in duplicate (panel B).

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Antibody 2C08 protected against coronavirus infection
The researchers used 4-6 week Syrian hamsters treated with 2C08 and challenged with either the D614G strain or the B.1.351 variant.
The 2C08 prevented severe complications from being infected with the D614G SARS-CoV-2 and the B.1.351 variant. Hamsters showed reduced inflammation by decreasing the number of proinflammatory cytokines in the lungs. They also showed lower viral RNA levels in the lungs by more than 10,000-fold with D614G and a 1,000-fold reduction in the B.1.351 variant. A significant decrease in host gene-expression was observed for Ccl3, CcL5, Ifit3, Ifit6, Ip10, Irf7, and Rig-I in the lungs of hamsters given the 2C08 antibody.
Hamsters treated with 2C08 also did not lose weight compared to hamsters given the control antibody. In fact, they gained more weight throughout the study than the control group.
Low possibility of a mutation escaping 2C08 neutralization
The team used chimeric viruses using the spike protein from the D614G strain to find variants that can evade neutralization from 2C08. They identified the S escape mutations G476D, G476S, G485D, F486P, F486V, and N487D, which are in the receptor-binding domain and play a role in ACE2 binding.
Using genomic surveillance, they screened 829,162 genomes from the GISAID public database to see if any of the identified 2C08 escape mutations were associated with any SARS-CoV-2 variants.
Of the 6, 4 escape mutants were found in circulating SARS-CoV-2 strains — although the frequency of finding one was rare. Results showed a 0.008% chance of finding an escaped mutant in a virus. It was more likely to find the D614G substitution, which appeared in 49% of sequenced samples.
2C08 binds to the receptor-binding motif
Because the antibody S2E12 has a 95% amino acid identity to 2C08 and shares similar heavy amino acid residues that engage the receptor-binding domain, the researchers suggest its mechanism of action is similar 2C08. S2E12’s can detect a receptor-binding domain epitope that partially overlaps with the ACE2 receptor footprint known as the receptor-binding motif.
More evidence of this mechanism of action is observed with 253H55L and COV2-2196, who have similar interactions and share genetic features to 2C08.
The researchers also uncovered 2C08-like clonotypes with 20 more antibodies that share the same genetic attributes of 2C08, S2E12, 253H55L, and COV2-2196 isolated by different patients. Thus, 2C08 is labeled as a ‘public’ B cell clonotype as it is clonally related to other antibodies characterized around the world.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Schmitz AJ, et al. A public vaccine-induced human antibody protects against SARS-CoV-2 and emerging variants. bioRxiv, 2021. doi: https://doi.org/10.1101/2021.03.24.436864, https://www.biorxiv.org/content/10.1101/2021.03.24.436864v1
- Peer reviewed and published scientific report.
Schmitz, Aaron J., Jackson S. Turner, Zhuoming Liu, Julian Q. Zhou, Ishmael D. Aziati, Rita E. Chen, Astha Joshi, et al. 2021. “A Vaccine-Induced Public Antibody Protects against SARS-CoV-2 and Emerging Variants.” Immunity 54 (9): 2159-2166.e6. https://doi.org/10.1016/j.immuni.2021.08.013. https://www.cell.com/immunity/fulltext/S1074-7613(21)00345-9?.