COVID-19 (coronavirus disease 2019) is caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), displaying high mortality rates. Responsible for over 3.34 million deaths the world over, the virus continues to rage on in many countries. Due to the rise of variants of concern (VoC) and the long-term threat of other emerging coronaviruses in the near future, a comprehensive and effective pandemic response strategy is required. However, despite the unprecedented development of vaccines and their administration across the globe, the prevalence of epidemics and the immune escape of viral variants, various studies are underway to develop a pan-coronavirus vaccine.
In the pre-clinical assessment of four categories of S-domain ferritin nanoparticles, including stabilized S-trimer-ferritin nanoparticles (SpFN), RBD-ferritin nanoparticles (RFN), S1-ferritin nanoparticles, and RBD-NTD-ferritin nanoparticles, the researchers found substantial neutralizing antibody titers against SARS-CoV-2 and the related VoC.
The researchers tested the antibodies in the K18-hACE2 mouse (transgenic mice expressing human ACE2) and found that the antibody levels provide robust protective immunity against SARS-CoV-2 challenge in this model. Importantly, they also demonstrated that subsequent immunizations increased not only the neutralization titer for SARS-CoV-2 but also the neutralization breadth and titer against the heterologous SARS-CoV-1 virus.
“Iterative structure-based design for viral glycoproteins stabilizing neutralizing epitopes or epitope-based vaccine design show that rational vaccine design can lead to the elicitation of broad immune responses.”
The coronaviruses are decorated with the spike (S) protein on the surface; it mediates virus entry, is immunogenic, and encodes multiple neutralizing epitopes. Therefore, the S protein is the primary target for natural and vaccine-induced CoV humoral immunity and vaccine design.
Spike is a glycoprotein, consisting of an S1 attachment subunit and S2 fusion subunit. In the S1 subunit, there is an N-terminal domain (NTD), and a C-terminal domain (CTD) that includes the receptor-binding domain (RBD). The RBD binds to the human angiotensin-converting enzyme 2 (hACE2), facilitating cell entry.
In this study, the researchers designed four categories of S-domain ferritin-fusion recombinant proteins as immunogens for expression as nanoparticles based on the major antigenic domains of the S ectodomain: (i) S ectodomain (residues 12-1158) (ii) RBD (residues 331-527), (iii) RBD linked in tandem to the NTD (residues 12-303), and (iv) S1 (residues 12-696). The Helicobacter pylori ferritin molecule was genetically linked to the C-terminal region of these S antigens.
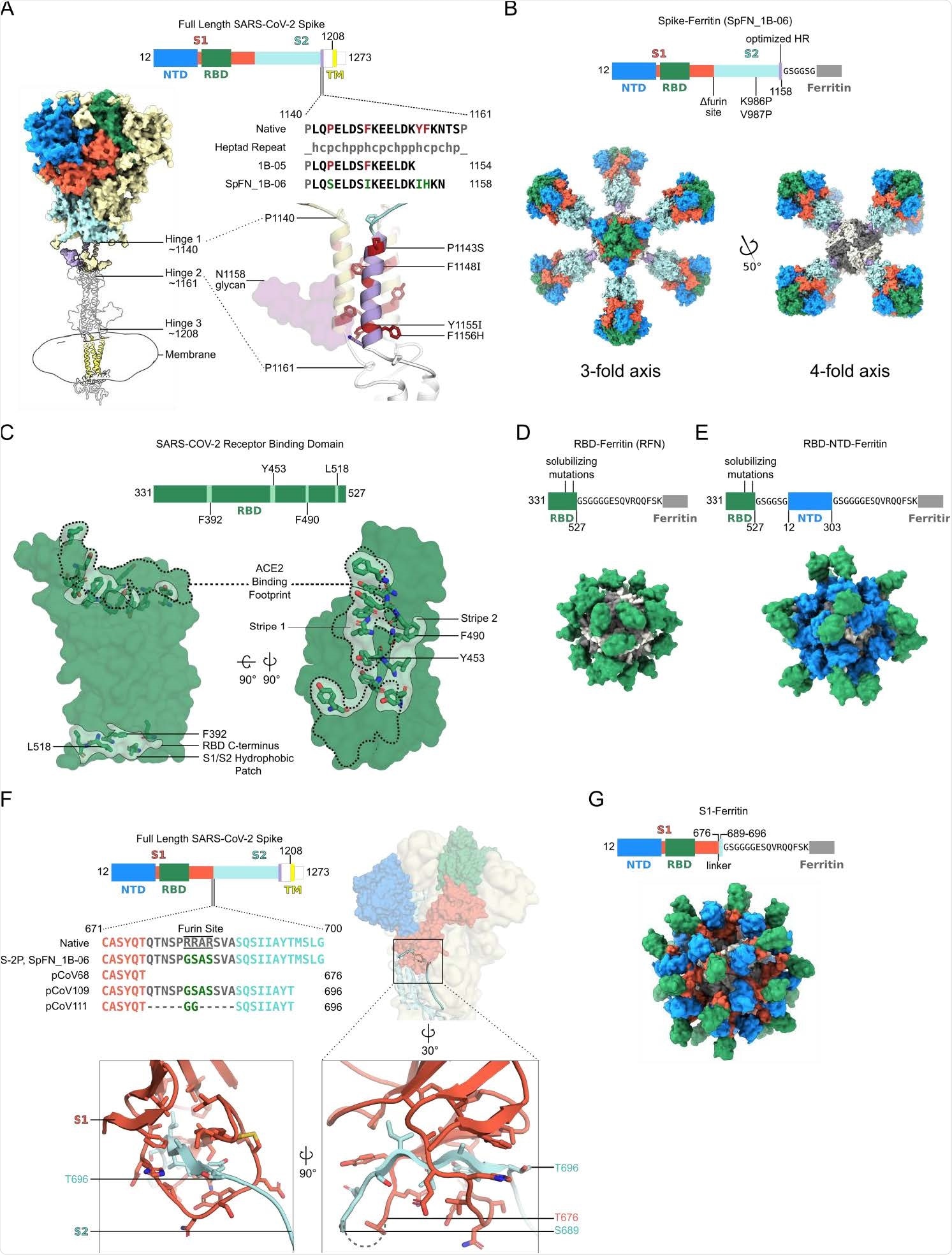
Structure-based design of SARS-CoV-2 S-based ferritin nanoparticle immunogens (A) Full length SARS-CoV-2 S schematic and 3D-structure. S hinges identified by molecular dynamics simulations and electron cryotomography are labeled on the 3D- model ((Turonova et al., 2020). The structured trimeric ectodomain is colored according to the schematic with the N-terminal domain (NTD) and Receptor-Binding Domain (RBD) of the S1 polypeptide and the C-terminal coiled coil N-terminal to hinge 1 colored blue, green, and purple, respectively. The remaining portions of the S1 and S2 polypeptides are colored in red and cyan with regions membrane-proximal from hinge 2 colored in white. The transmembrane domain of all chains is depicted in yellow. To design a Spike-Ferritin molecule, the C-terminal heptad repeat (residues 1140 to 1161) between Hinge 1 and 2 were aligned to an ideal heptad repeat sequence. Residues in the native S sequence which break this pattern are highlighted in red. These residues are also labeled and highlighted in red on the 3D-structure. Two engineered designs (1B-05 and 1B-06) are shown, with S end residue used to link to Ferritin, and heptad-repeat mutations colored green. (B) Schematic and 3D model of Spike Ferritin nanoparticle (SpFN). Differences between the native S sequence and the engineered nanoparticle are indicated on the schematic. A 3D811 model of SpFN displaying eight trimeric Spikes was created using PDB ID 6VXX and 3EGM with the ferritin molecule shown in alternating grey and white. The nanoparticle is depicted along the 4-fold and the 3-fold symmetry axes of the ferritin. (C) RBD–Ferritin nanoparticle design and optimization. The RBD of SARS-CoV-2 (PDB ID:6MOJ) is shown in surface representation, with the ACE2 binding site outlined in dashed lines. Three hydrophobic regions of the RBD which were mutated for nanoparticle immunogen design are shown in light green surface, with residues in stick representation. The ACE2 binding site contains two of these regions, while a third hydrophobic patch near the C-terminus of the RBD is typically buried by S2 and part of S1 in the context of the trimer molecule. (D) Schematic and 3D model of an RBD–Ferritin nanoparticle. A modeled 24-mer nanoparticle displaying the RBD domain is depicted at the 3-fold symmetry axis of ferritin and colored green. Truncation points, linkers, and alterations made to the RBD sequence are indicated on the schematic. (E) Schematic and 3D model of an RBD–NTD–Ferritin nanoparticle. A modeled nanoparticle displaying RBD and NTD epitopes is depicted and colored according to the schematic. Truncation points, linkers, and alterations made to the native S sequence are indicated on the primary structure. (F) S1-Ferritin immunogen design. The SARS-CoV-2 S1 forms a hydrophobic collar around the N-terminal β-sheet of S2 (residues 689-676). S1-ferritin immunogen design required inclusion of this short stretch of S2 (colored cyan) attached by a linker. Terminal residues of the structured portions of S1 and S2 are labeled. (G) Schematic and 3D model of an S1–Ferritin nanoparticle. A modeled nanoparticle displaying RBD and NTD domains is depicted and colored according to the S1–ferritin schematic with truncation points and domain linkers indicated.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Two adjuvants used in this study are ALFQ (a liposome-based adjuvant containing the saponin QS-21, and synthetic Monophosphoryl Lipid A (3D-PHAD), and Alhydrogel (contains aluminum hydroxide gel). The researchers observed that ALFQ was superior to Alhydrogel as an adjuvant to elicit binding
and neutralizing responses. Also, Alhydrogel led to a skewed antibody isotype immune response that was TH2 in nature, as opposed to the balanced immune response seen with ALFQ adjuvanted animals, the researchers noted.
The researchers stated that based on the results described in this study and other data from associated non-human primate experiments, an S-Ferritin immunogen with a liposomal adjuvant, ALFQ is currently being assessed in phase I clinical trial (NCT04784767).
The researchers characterized and assessed the nanoparticles using a variety of biophysical, structural and antigenic assays combined with animal immunogenicity testing. The overall objective of the study was to determine the best immunogen for further development by evaluating the immunogen structure, antigenicity, and immunogenicity of the different immunogens.
“Utilizing naturally occurring nanoparticle molecules such as bacterial ferritin, antigens are fused to the ferritin molecule to recapitulate complex trimeric class I glycoproteins, and to increase the immune response for weakly immunogenic targets.”
Through the use of nanoparticle vaccine technology and new adjuvants, researchers have engineered candidates that are more likely to induce effective and broader immune responses.
In conclusion, the researchers reported the structure-based design and pre-clinical assessment of four categories of S-domain ferritin nanoparticles, including stabilized S-trimer-ferritin nanoparticles (SpFN), RBD-ferritin nanoparticles (RFN), S1-ferritin nanoparticles, and RBD146 NTD-ferritin nanoparticles. They demonstrated the robust protective immunity of these nanoparticles against the SARS-CoV-2, and related VoC, and also showed the expanded neutralization breadth and titer against the heterologous SARS-CoV-1 virus.
The highlights in this study are: 1) Iterative structure-based design of four Spike-domain Ferritin nanoparticle classes of immunogens; 2) SpFN-ALFQ and RFN-ALFQ immunization elicits potent neutralizing activity against SARS-CoV-2, variants of concern, and SARS-CoV-1; 3) Passively transferred IgG from immunized C57BL/6 mice protects K18-hACE2 mice from lethal SARS-CoV-2 challenge.
Each of these different designs and the underlying development processes provides a greater understanding and framework for ongoing and future pan-coronavirus vaccine design and development, the researchers write.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
SARS-CoV-2 ferritin nanoparticle vaccines elicit broad SARS coronavirus immunogenicity, M. Gordon Joyce, Wei-Hung Chen, Rajeshwer S. Sankhala, Agnes Hajduczki, Paul V. Thomas, Misook Choe, William Chang, Caroline E. Peterson, Elizabeth Martinez, Elaine B. Morrison, Clayton Smith, Aslaa Ahmed, Lindsay Wieczorek, Alexander Anderson, Rita E. Chen, James Brett Case, Yifan Li, Therese Oertel, Lorean Rosado, Akshaya Ganesh, Connor Whalen, Joshua M. Carmen, Letzibeth Mendez-Rivera, Christopher Karch, Neelakshi Gohain, Zuzana Villar, David McCurdy, Zoltan Beck, Jiae Kim, Shikha Shrivastava, Ousman Jobe, Vincent Dussupt, Sebastian Molnar, Ursula Tran, Chandrika B. Kannadka, Michelle Zemil, Htet Khanh, Weimin Wu, Matthew A. Cole, Debra K. Duso, Larry W. Kummer, Tricia J. Lang, Shania E. Muncil, Jeffrey R. Currier, Shelly J. Krebs, Victoria R. Polonis, Saravanan Rajan, Patrick M. McTamney, Mark T. Esser, William W. Reiley, Morgane Rolland, Natalia de Val, Michael S. Diamond, Gregory D. Gromowski, Gary R. Matyas, Mangala Rao, Nelson L. Michael, Kayvon Modjarrad, bioRxiv 2021.05.09.443331; doi: https://doi.org/10.1101/2021.05.09.443331, https://www.biorxiv.org/content/10.1101/2021.05.09.443331v1
- Peer reviewed and published scientific report.
Joyce, M. G., Chen, W.-H., Sankhala, R. S., Hajduczki, A., Thomas, P. V., Choe, M., Martinez, E. J., Chang, W. C., Peterson, C. E., Morrison, E. B., Smith, C., Chen, R. E., Ahmed, A., Wieczorek, L., Anderson, A., Case, J. B., Li, Y., Oertel, T., Rosado, L., & Ganesh, A. (2021). SARS-CoV-2 ferritin nanoparticle vaccines elicit broad SARS coronavirus immunogenicity. Cell Reports, 37(12), 110143. https://doi.org/10.1016/j.celrep.2021.110143. https://www.cell.com/cell-reports/fulltext/S2211-1247(21)01639-9.