The rapid spread of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has posed a serious threat to the global health and economy. To date, SARS-CoV-2, which is the virus responsible for the coronavirus disease 2019 (COVID-19), has claimed more than 4.26 million lives worldwide and infected more than 200 million people.
SARS-CoV-2, which belongs to the family Coronaviridae, is a highly infectious virus that mainly infects the respiratory tract of humans. The viral genome of SARS-CoV-2 is found within the nucleocapsid (N) protein, which is embedded inside phospholipid bilayers and covered by a spike (S) protein.
In addition to the N and S proteins, SARS-CoV-2 has two other two structural proteins including the membrane (M) and envelope (E) proteins, both of which are located in the viral envelope. SARS-CoV-2 also encodes for several accessory proteins.
SARS-CoV-2 infection can cause a wide range of symptoms ranging from mild to severe, the latter of which can result in serious complications and sometimes death. Scientists believe that in order to contain the spread of SARS-CoV-2, it is essential to understand the pathogenesis of this virus.
What is an inflammasome?
Severe inflammatory responses often occur when SARS-CoV-2 infects the human respiratory tract. Previous research has shown that a dysregulated innate immune response is associated with the clinical symptoms of severe COVID-19 infections.
Inflammasomes are cytosolic multiprotein oligomers of the innate immune system that are responsible for the activation of inflammatory responses. These proteins can recognize cellular stresses and infections.
There are four main types of inflammasomes, which include NLRP1, NLRP3, NLRC4, and AIM2. Among these, NLRP3 is associated with RNA virus infection.
The NRLP3 inflammasome consists of a sensor protein, adaptor protein, and effector protein. Previous research had shown that the NLRP3 protein contains three domains including the Pyrin domain (PYD), Nucleotide-binding domain, and Leucine-rich repeat domain.
Upon activation of the NLRP3 protein, the PYD interacts with the apoptosis-associated speck-like protein containing a caspase recruiting domain (ASC) and triggers the formation of ASC oligomer. This oligomer offers the platform for the activation of caspase-1, which catalyzes proteolytic processing of pro-interleukin (IL)-1β into mature IL-1β.
However, abundantly present IL-1β provokes the systemic inflammation responses, which are involved with various signaling pathways like the nuclear factor-κB (NF-κB) and c-Jun N-terminal kinase pathways. Therefore, a large number of cytokines, such as IL-6, tumor necrosis factor (TNF), interferon (IFN)-α, IFN-β, and transforming growth factor-β, are released, which can result in cytokine storms. Previous studies have revealed that SARS-CoV-2 infection elicits the production of inflammasomes that are linked with the severity of COVID-19.
Several studies have also reported that the E protein and the accessory protein ORF3a of SARS-CoV can activate NLRP3 inflammasome by altering the potassium (K+) ion permeability of the plasma membrane and by the production of mitochondrial reactive oxygen species. However, there remains a gap in research explaining the specific molecular pathway by which SARS-CoV-2 activates NLRP3 inflammasomes.
Study findings
A new study published in Nature Communications has revealed the mechanism by which the SARS-CoV-2 N protein activates NLRP3 inflammasomes to induce hyperinflammation. The authors of the current study reported that post-SARS-CoV-2 infection, the macrophages and dendritic cells (DCs) of the host trigger cytokines and chemokines. This observation was made by analyzing the GEO database (GSE155106).
Mechanistically, the mouse model revealed that the SARS-CoV-2 N protein is responsible for intensifying lung injury and causing acute inflammation in mice by facilitating the activation of NLRP3 inflammasomes. The researchers have also found that N proteins directly interact with NLRP3 to promote the assembly of the NLRP3 inflammasome.
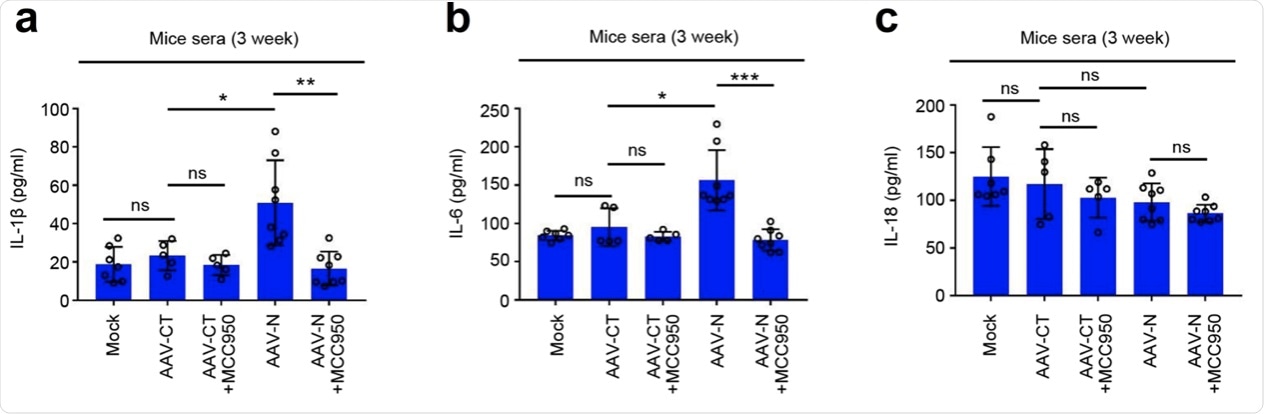
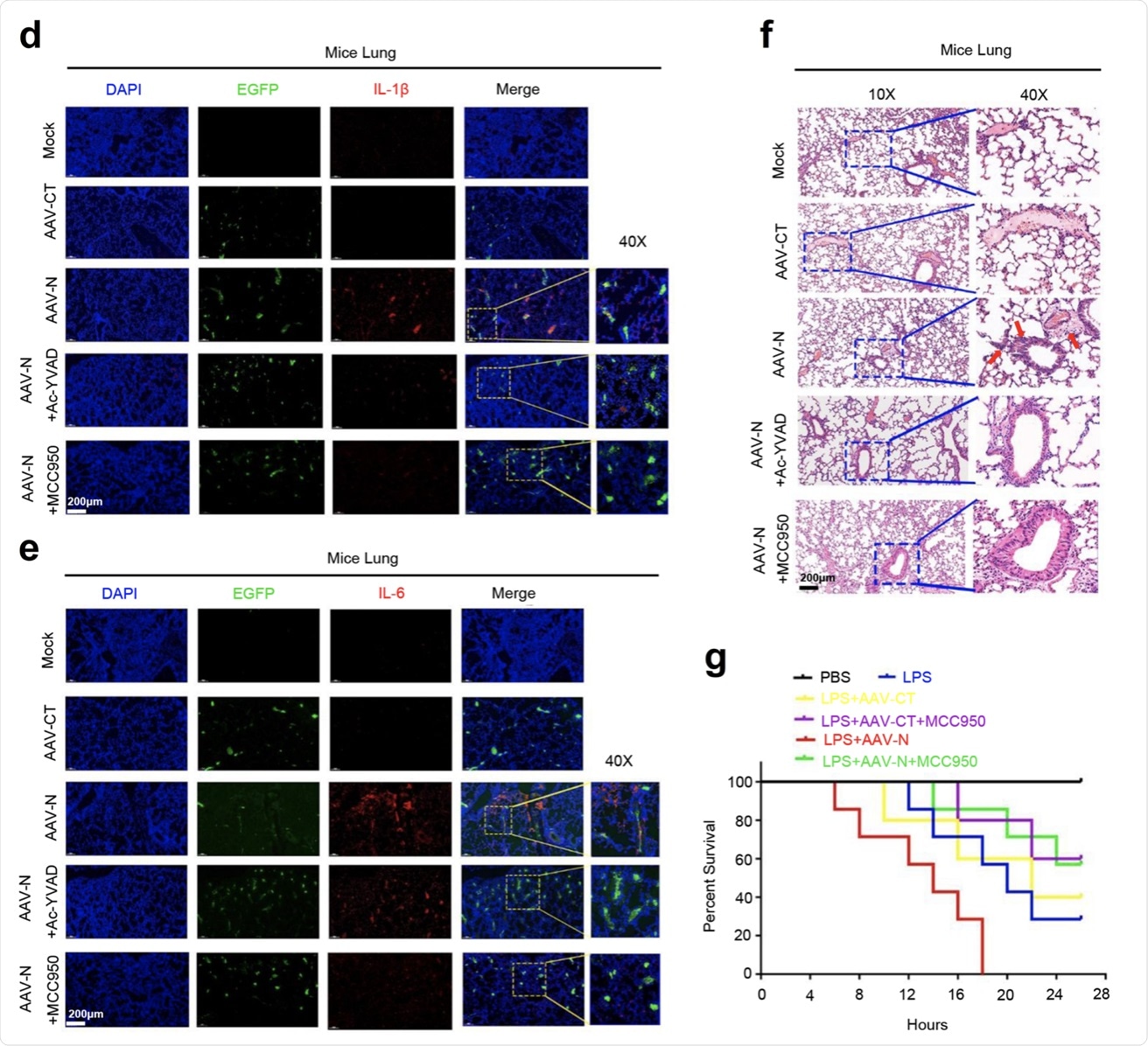 a–c C57BL/6 genetic background mice were tail vein-injected with 300 μl containing 5 × 1011 vg of AAV-Lung-EGFP (n = 10) or AAV-Lung-N (n = 16) for 2 weeks and then treated with MCC950 (50 mg/kg) by intraperitoneal injection for AAV-Lung-EGFP (n = 5) or AAV-Lung-N (n = 8) mice. Serum was collected at 3 weeks for each group from the orbit. IL-1β (a), IL-6 (b), or IL-18 (c) in the sera was measured by ELISA. Points represent the value of each serum samples. d–f At 3 weeks, mice were killed and the lungs were collected. Histoimmunofluorescence analysis of IL-1β (d, red) or IL-6 (e, red) in the lung after AAV-CT or AAV-N infection. Scale bar is 200 μm (10×) or 50 μm (40×). Histopathology analysis of lung after AAV-CT or AAV-N infection (f). Red arrows indicated the infiltrated inflammatory cells. Scale bar is 200 μm (10×) or 50 μm (40×). g After 4 weeks, pretreated AAV-Lung-N mice (n = 7) with DMSO, pretreated AAV-Lung-N (n = 7) mice with MCC950, or pretreated AAV-Lung-EGFP (n = 5) with MCC950. After 30 min, mock group (n = 7), AAV-Lung-EGFP group (n = 5), AAV-Lung-N (n = 7) group, and another AAV-Lung-N (n = 7) group were treated with LPS (30 mg/kg) by intraperitoneal injection. Another mock group (n = 3) was intraperitoneally injected with PBS. The mice survival rates were evaluated every 2 h post treatment. Mock means inject the same dose of PBS as other groups (a–h). Data are representative of two independent experiments and one representative is shown. Error bars indicate SD of each serum samples, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, two-tailed Student’s t-test (a–c). One-way ANOVA analysis (g). Source data are provided as a Source Data file.
a–c C57BL/6 genetic background mice were tail vein-injected with 300 μl containing 5 × 1011 vg of AAV-Lung-EGFP (n = 10) or AAV-Lung-N (n = 16) for 2 weeks and then treated with MCC950 (50 mg/kg) by intraperitoneal injection for AAV-Lung-EGFP (n = 5) or AAV-Lung-N (n = 8) mice. Serum was collected at 3 weeks for each group from the orbit. IL-1β (a), IL-6 (b), or IL-18 (c) in the sera was measured by ELISA. Points represent the value of each serum samples. d–f At 3 weeks, mice were killed and the lungs were collected. Histoimmunofluorescence analysis of IL-1β (d, red) or IL-6 (e, red) in the lung after AAV-CT or AAV-N infection. Scale bar is 200 μm (10×) or 50 μm (40×). Histopathology analysis of lung after AAV-CT or AAV-N infection (f). Red arrows indicated the infiltrated inflammatory cells. Scale bar is 200 μm (10×) or 50 μm (40×). g After 4 weeks, pretreated AAV-Lung-N mice (n = 7) with DMSO, pretreated AAV-Lung-N (n = 7) mice with MCC950, or pretreated AAV-Lung-EGFP (n = 5) with MCC950. After 30 min, mock group (n = 7), AAV-Lung-EGFP group (n = 5), AAV-Lung-N (n = 7) group, and another AAV-Lung-N (n = 7) group were treated with LPS (30 mg/kg) by intraperitoneal injection. Another mock group (n = 3) was intraperitoneally injected with PBS. The mice survival rates were evaluated every 2 h post treatment. Mock means inject the same dose of PBS as other groups (a–h). Data are representative of two independent experiments and one representative is shown. Error bars indicate SD of each serum samples, *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, two-tailed Student’s t-test (a–c). One-way ANOVA analysis (g). Source data are provided as a Source Data file.
Previous research by the same group of scientists on the Dengue and Zika viruses reported that, in the case of the Dengue virus, the M protein caused tissue damage and vascular leakage in mice by activating the NLRP3 inflammasome. In the case of the Zika virus, the NS5 protein promoted the entry of the virus into the brain.
The N protein of SARS-CoV-2 had been reported to play an important role in its replication process. The current study demonstrated that the SARS-CoV-2 N protein exists as the viral E protein that enters the cytoplasm and activates NLRP3 before viral assembly.
Additionally, this study showed that the levels of IL-1β and IL-6 in the lungs and sera were increased by the lung-specific expression of N protein. However, the expressions of IL-1β and IL-6 were significantly reduced following the administration of MCC950 and Ac-YVAD-cmk to the infected mice.
Utility in treating COVID-19
The authors of this study are extremely optimistic that inhibition of NLRP3 inflammasome can effectively reduce the “cytokine storm” and lung injury caused by COVID-19. In other words, they suggested that the NLRP3 inflammasome could be an alternative target for COVID-19 therapy.
Interestingly, this research revealed that SARS-CoV-2 N-induced lung injury and cytokine production are repressed by MCC950, which is a specific NLRP3 inhibitor, and Ac-YVAD-cmk, which is an inhibitor of the caspase-1. Therefore, these two candidates can be potentially used as therapeutic agents for the prevention and treatment of COVID-19.