The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which is the virus responsible for the coronavirus disease 2019 (COVID-19), is a positive-stranded and enveloped ribonucleic acid (RNA) virus that belongs to the family Coronaviridae. To date, SARS-CoV-2 has infected over 200 million individuals and claimed more than 4 million lives worldwide.
Several vaccines and therapeutics are currently available for the management of COVID-19; however, more research is required for the development of new effective strategies to combat the numerous SARS-CoV-2 variants of concern (VoCs).
 Study: A method for the generation of pseudotyped virus particles bearing SARS coronavirus spike protein in high yields. Image Credit: Blue Planet Studio / Shutterstock.com
Study: A method for the generation of pseudotyped virus particles bearing SARS coronavirus spike protein in high yields. Image Credit: Blue Planet Studio / Shutterstock.com

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Importance of characterizing SARS-CoV-2
Similar to that of SARS-CoV, the envelope of SARS-CoV-2 contains three structural proteins that include the spike (S), membrane (M), and small envelope (E) proteins.
Researchers have classified the SARS-CoV-2 S protein as a class I viral fusion protein. Other members of this class include retrovirus envelope proteins and influenza virus hemagglutinin proteins.
The SARS-CoV-2 S protein is a homotrimer whose key functional role is in the interaction of the virus with host cell receptors. The S protein promotes the entry of the virus into the host cell via endocytosis or membrane fusion. The S1 subunit region of the S protein binds to the angiotensin-converting enzyme 2 (ACE2) receptor on the host cell.
Thereafter, in the presence of transmembrane protease serine 2 (TMPRSS2) and furin, the S protein undergoes proteolytic cleavage into S1 and S2 fusion subunits at the S1/S2 site. Other cleavage sites located in the S2 region are cleaved by host cell proteases such as cathepsins, furin-like proprotein convertases, and trypsin-like serine protease.
Scientists have claimed that it is extremely important to understand the role of the SARS-CoV-2 S protein at the molecular level, as it would provide a better understanding of the mode by which virus enters the host cell. However, conducting this research is challenging because it requires performing experiments using live viruses. As a result, researchers conducting these types of studies must follow biosafety level 3 (BSL-3) requirements.
What is a pseudovirus system?
Pseudotyped virus systems play an important role in studying the mode of entry of the virus into the host cells, as well as the characterization of viral envelope proteins. Researchers have developed pseudovirus particles that consist of a surrogate viral core and virus envelope proteins that are analogous to the original virus.
Scientists have successfully modified the genome of the original virus such that they removed the virulence-specific genes to prevent the production of infectious progeny viruses. By using these avirulent pseudotype viruses, researchers can safely conduct their research at a BSL-2 facility instead of a BSL-3 or BSL-4 facility.
Typically, Vesicular Stomatitis Virus (VSV), which is a negative-strand RNA virus, is used to generate pseudotyped virus systems and recombinant viruses to study structural proteins. When studying the SARS-CoV-2 S protein using VSV, researchers used a conventional transfection protocol to reveal that most of the pseudotyped particles produced by this method lacked S protein.
A new study published on the preprint server bioRxiv* discusses the development of an improved protocol based on cell lines that can consistently express S proteins. The authors of this study claim that the new protocol would immensely benefit researchers who are interested in understanding the mode of entry of SARS-CoV-2 into the host cell. This research would also help in the development of new antiviral agents.
Study findings
Initially, the researchers hoped to assess the loading efficiency of S proteins on VSV pseudotyped virus particles. This was performed using HEK293T cells that generated pseudotyped virions.
Immunofluorescence analysis of these virions was conducted after staining them with lipophilic carbocyanine dye DiI. Confocal fluorescence imaging showed that only a small portion of these particles contained the S protein. The current study revealed that for the production of pseudotyped virus particles bearing uniform S protein, a low and persistent level of S protein expression is essential.
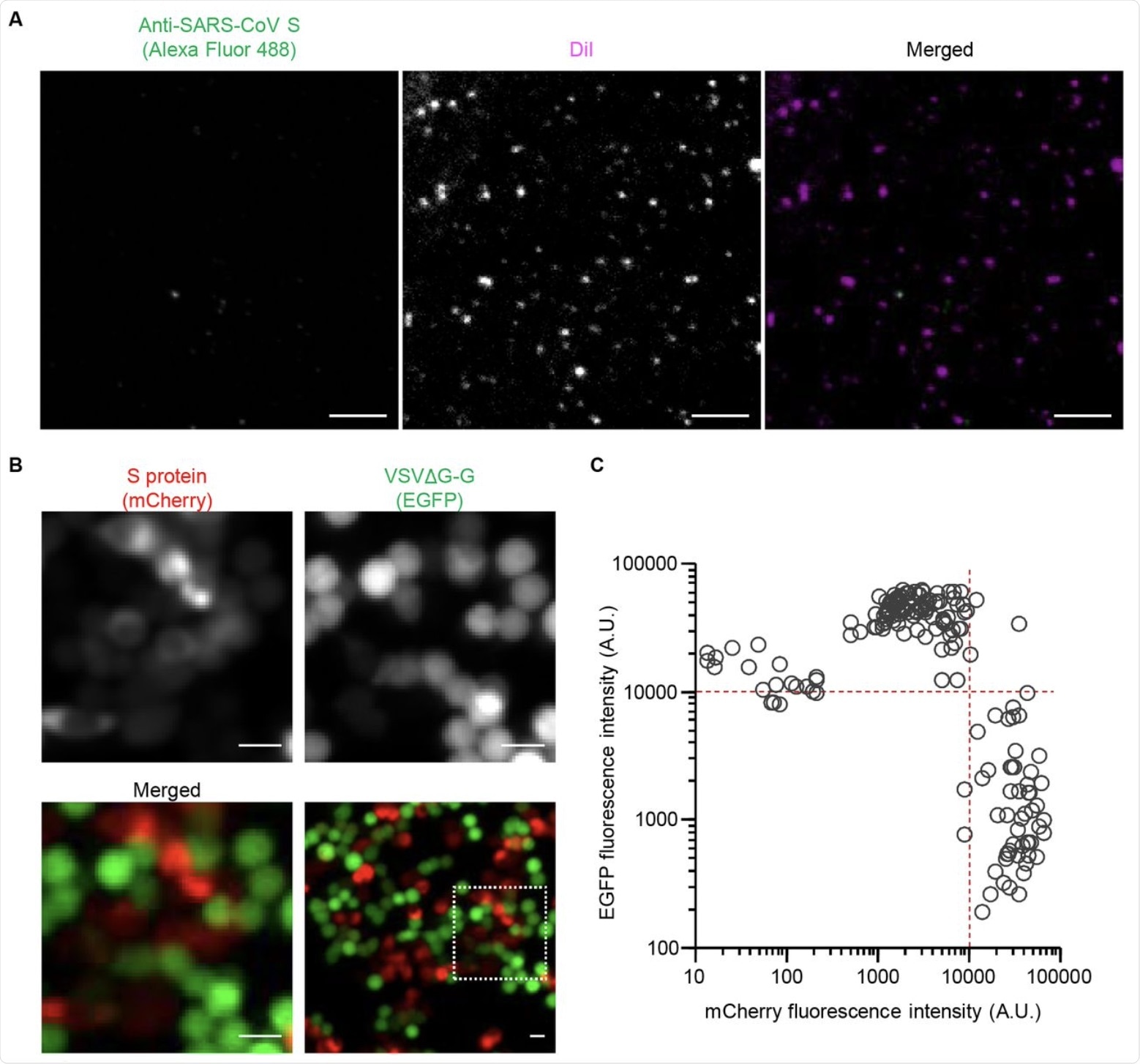 (A) HEK293T cells were transfected with an expression vector for SARS-CoV S protein as well as infected with VSVΔG-G. The culture supernatant was collected and stained with DiI as well as subjected to immunofluorescence staining with antibodies to SARS-CoV S protein and Alexa Fluor 488–conjugated secondary antibodies. The virus particles were then adsorbed onto a polyethylenimine-coated glass-based plate and observed with a confocal fluorescence microscope. Representative images are shown. Bars, 10 μm.
(A) HEK293T cells were transfected with an expression vector for SARS-CoV S protein as well as infected with VSVΔG-G. The culture supernatant was collected and stained with DiI as well as subjected to immunofluorescence staining with antibodies to SARS-CoV S protein and Alexa Fluor 488–conjugated secondary antibodies. The virus particles were then adsorbed onto a polyethylenimine-coated glass-based plate and observed with a confocal fluorescence microscope. Representative images are shown. Bars, 10 μm.
(B) HEK293T cells were transfected with an expression vector for mCherry-tagged SARS-CoV S protein and infected with VSVΔG-G for 16 h. They were then observed with a fluorescence microscope for detection of mCherry (red) and GFP (green) fluorescence. Representative images are shown. The boxed region in the lower right panel is shown at higher magnification in the other panels. Bars, 10 μm.
(C) Fluorescence intensities of GFP and mCherry in individual cells as in (B).
Subsequently, the researchers isolated transfected VeroE6 cells and found that all of the virus particle products were positive for the S protein. This showed that VeroE6 cells lines could more consistently and stably produce virus particles bearing the S protein as comparied to transiently transfected HEK293T cells. Additionally, this quantitative study revealed a significantly higher amount of viral particles with the S protein in the stable VeroE6 cells lines as compared to transiently transfected HEK293T cells.
Taken together, the novel protocol discussed here has proved to be highly efficient in producing pseudoviruses that can be used to study the mode of cell entry by SARS-CoV or SARS-CoV-2.
Conclusion
The current study has successfully developed a modified protocol to generate pseudoviruses based on stable cell lines. The virus particles produced by this method harbored a large amount of the S protein, which is known to play a vital role in SARS-CoV-2 infection. This research should aid in the development of vaccines and therapeutics for COVID-19.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Fujioka, Y., Kashiwagi, S., Yoshida, A., et al. (2021). A method for the generation of pseudotyped virus particles bearing SARS coronavirus spike protein in high yields. bioRxiv. doi:10.1101/2021.07.30.454063. https://www.biorxiv.org/content/10.1101/2021.07.30.454063v1
- Peer reviewed and published scientific report.
Fujioka, Yoichiro, Sayaka Kashiwagi, Aiko Yoshida, Aya O. Satoh, Mari Fujioka, Maho Amano, Yohei Yamauchi, and Yusuke Ohba. 2022. “A Method for the Generation of Pseudovirus Particles Bearing SARS Coronavirus Spike Protein in High Yields.” Cell Structure and Function. https://doi.org/10.1247/csf.21047. https://www.jstage.jst.go.jp/article/csf/47/1/47_21047/_article.