The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which is the virus responsible for the coronavirus disease 2019 (COVID-19), is a highly infectious positive-stranded ribonucleic acid (RNA) virus belonging to the family Coronaviridae. Among RNA viruses, coronaviruses contain the largest genome and are associated with complex replication processes.
The complexity of coronavirus replication is because this process involves frame shift slipping, during which the replicase jumps and multiple RNA duplexes are generated. Coronaviruses also develop numerous membrane compartments to protect their viral components, which increases replication efficiency and helps these viruses to evade innate immune recognition.
 Study: Correlative multi-scale cryo-imaging unveils SARS-CoV-2 assembly and egress. Image Credit: CROCOTHERY / Shutterstock.com
Study: Correlative multi-scale cryo-imaging unveils SARS-CoV-2 assembly and egress. Image Credit: CROCOTHERY / Shutterstock.com
The structural proteins of SARS-CoV-2
SARS-CoV-2 consists of four different structural proteins, which include the spike (S), membrane (M), envelope (E), and nucleocapsid (N) proteins. The primary function of the S protein, which consists of S1 and S2 domains, is to mediate the entry of the virus into the host cell.
SARS-CoV-2 infection involves two steps of pre-fusion and post-fusion. Whereas pre-fusion involves both S1 and S2 subunits, the latter involves the S2 subunit alone.
The main role of the SARS-CoV-2 N protein is encapsulating and protecting the viral genome. The E protein is the smallest structural protein and acts as an ion channel. Lastly, the transmembrane M protein, which is the most abundantly present structural protein in SARS-CoV-2, lines the inner surface of the virus lipid membrane.
SARS-CoV-2 replication cycle
The first step of SARS-CoV-2 infection involves the interaction between the virus’ S protein and the angiotensin-converting enzyme 2 (ACE2) receptor present on the surface of the host cell. Subsequently, S2 either cleaves at the cell surface in the presence of cell protease TMPRSS2 or triggers endocytosis of the viral particles in absence of TMPRSS2.
When the viral components are introduced into the cytoplasm of the host cell, the precursor polyproteins Pp1a and Pp1ab are synthesized. The SARS-CoV-2 non-structural proteins like nsp3, nsp4, and nsp6 promptly induce the formation of membranous compartments called double-membrane vesicles (DMVs). The viral genome replication and transcription processes occur within these membranous compartments.
Scientists are curious about the mechanism by which messenger RNAs (mRNAs) reach the cytoplasm to be translated by the cellular ribosomes from sealed DMVs. A recent study on murine hepatitis coronavirus (MHV) and SARS-CoV-2 reported the presence of a molecular pore that served as an export portal for transferring mRNA and positive-strand viral genome copies.
Previous studies have also reported that the assembly of the viral particles takes place at the modified cellular membranes derived from the endoplasmic reticulum (ER), Golgi, and ER–Golgi intermediate compartment (ERGIC). These studies further added that the virus gets released via exocytosis.
Although extensive research has been conducted on the structural components of SARS-CoV-2, there is a limited amount of structural and ultrastructural evidence on how SARS-CoV-2 infection progresses in native cells. Additionally, there is a gap in the research related to the understanding of viral assembly and egress.
A multi-scale, cryo-correlative platform to visualize SARS-CoV-2 infection
A new study published in Nature Communications has correlated the cytopathic episodes triggered by SARS-CoV-2 with virus replication processes in frozen-hydrated cells. The authors of this study have used a unique correlative, multi-modal, and multi-scale cryo-imaging technique to examine the replication progress of SARS-CoV-2 using Vero cells.
The technique utilized in the current study is unique because it provides a holistic image of SARS-CoV-2 infection that includes the entire cell fused to individual virus spike molecules. This approach also revealed pathways associated with the virus assembly, as well as the egress and cytopathic effects of SARS-CoV-2 infection.
In this study, researchers combined different techniques including serial cryo serial focused ion beam scanning electron microscopy with scanning electron microscopy (cryoFIB/SEM) and soft X-ray tomography to study the whole-cell morphology. The data obtained from these techniques were then correlated with high-resolution cryo-electron tomography (cryoET). One of the advantages of cryoET is the use of frozen-hydrated conditions that offers rapid and simple sample preparation.
The images obtained on the replication of SARS-CoV-2 revealed a spatially well-organized and highly efficient process. Scientists observed that each step of the replication process including genome replication, protein synthesis, and virus assembly, to name a few, takes place in specific cytoplasmic compartments.
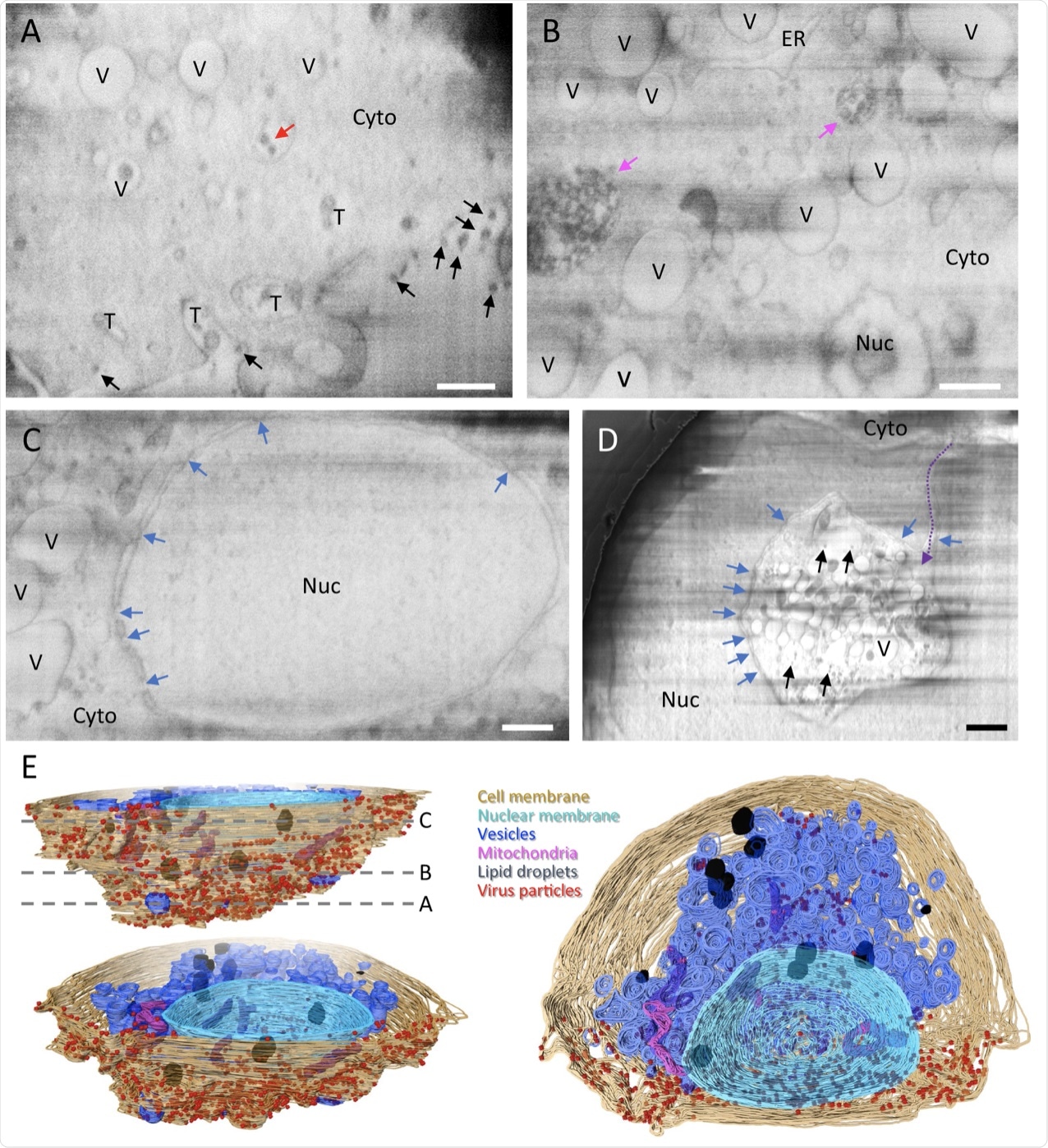 A–D Representative cryoFIB/SEM slices of a SARS-CoV-2-infected cell at the cell periphery (A), cytoplasm (B), cell nucleus (C), and invagination of cytoplasm into the nuclear space (note, from a different cell) (D). Scale bars, 500 nm in A–C, 1 µm in D. Black and red arrows, extracellular and intracellular virus particles, respectively; blue arrows, nuclear pores; pink arrows, complex membrane compartment; dashed purple arrow, invagination path. V vesicles, T tunnels, Nuc nucleus, Cyto cytoplasm, ER endoplasmic reticulum. E Surface rendering of the segmented volume of SARS-CoV-2-infected cell shown in A–C. Segmented organelles and virus particles are labeled with the colors indicated. The dashed lines (E, top left panel) indicate the positions of slices shown in A–C, respectively.
A–D Representative cryoFIB/SEM slices of a SARS-CoV-2-infected cell at the cell periphery (A), cytoplasm (B), cell nucleus (C), and invagination of cytoplasm into the nuclear space (note, from a different cell) (D). Scale bars, 500 nm in A–C, 1 µm in D. Black and red arrows, extracellular and intracellular virus particles, respectively; blue arrows, nuclear pores; pink arrows, complex membrane compartment; dashed purple arrow, invagination path. V vesicles, T tunnels, Nuc nucleus, Cyto cytoplasm, ER endoplasmic reticulum. E Surface rendering of the segmented volume of SARS-CoV-2-infected cell shown in A–C. Segmented organelles and virus particles are labeled with the colors indicated. The dashed lines (E, top left panel) indicate the positions of slices shown in A–C, respectively.
A multi-scale view of SARS-CoV-2 genome replication
The researchers of this study have illustrated five different steps associated with the replication of the SARS-CoV-2 genome, which includes:
- Step 1: RNA replication occurs in DMVs, thus separating vRNA and subgenomic mRNA from the host cell innate immune response.
- Step 2: The newly synthesized vRNAs are transported from DMVs to virus assembly sites through the transmembrane portals. The mRNAs are also transported by the same portals to the cytoplasm, where it produces proteins.
- Step 3: The viral S proteins are transported to assembly sites through small transport vesicles.
- Step 4: These vesicles fuse with single membrane vesicles (SMV), which is where viral spikes gather at the assembly site and the budding process begins. These events occur in the presence of vRNA. N proteins are also formed in this step.
- Step 5: After the assembly and budding processes, a virus-containing vesicle is formed that can contain multiple virus particles. These particles can then exit through tunnels or lysosomal exocytosis.
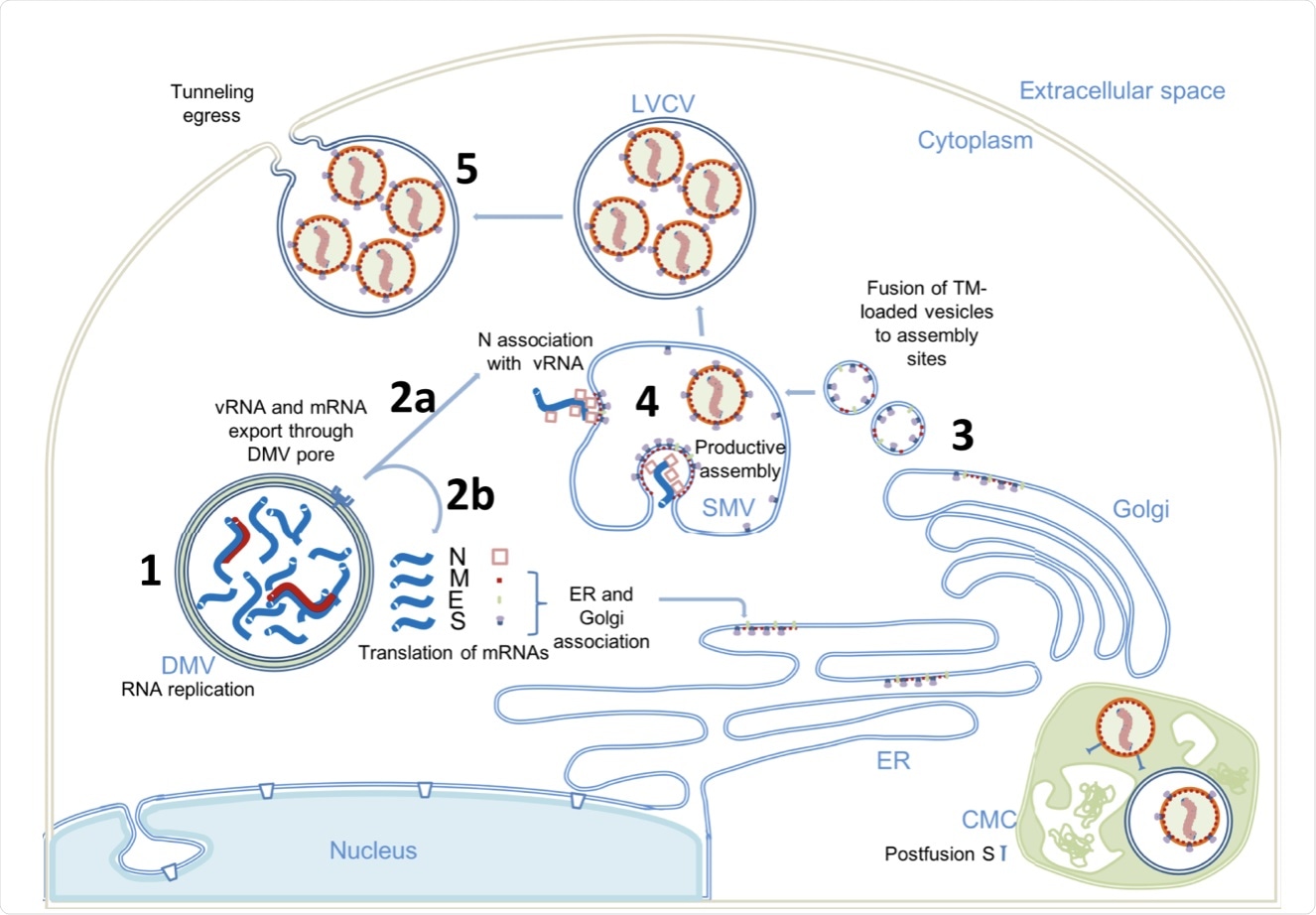 (1) Viral genome replication occurs inside the DMVs, generating the negative-strand viral and subgenomic RNAs (red), positive vRNA genomic copy, and subgenomic mRNAs (blue). (2) Positive RNAs are exported to cytoplasm through the DMV pores. Subgenomic mRNAs are translated (2b). Structural proteins M, E, and S associate with ER and Golgi membranes. Genomic vRNA becomes complexed with newly synthetized N (2a). (3) S, E, and M are transported in dense vesicles, which are fused with SMVs. (4) Productive viral assembly happens in the SMV clustering the viral spikes and encapsidating the genome in RNPs. Viruses bud to the internal space of the SMV. (5) Egress occurs through tunnels. The complex membrane compartment (CMC) is depicted in green, which encloses both postfusion spike-containing virus particles and prefusion spike-containing virus particles. DMV double membrane vesicle, SMV single membrane vesicle, CMC complex membrane compartment, LVCV large virus-containing vesicle.
(1) Viral genome replication occurs inside the DMVs, generating the negative-strand viral and subgenomic RNAs (red), positive vRNA genomic copy, and subgenomic mRNAs (blue). (2) Positive RNAs are exported to cytoplasm through the DMV pores. Subgenomic mRNAs are translated (2b). Structural proteins M, E, and S associate with ER and Golgi membranes. Genomic vRNA becomes complexed with newly synthetized N (2a). (3) S, E, and M are transported in dense vesicles, which are fused with SMVs. (4) Productive viral assembly happens in the SMV clustering the viral spikes and encapsidating the genome in RNPs. Viruses bud to the internal space of the SMV. (5) Egress occurs through tunnels. The complex membrane compartment (CMC) is depicted in green, which encloses both postfusion spike-containing virus particles and prefusion spike-containing virus particles. DMV double membrane vesicle, SMV single membrane vesicle, CMC complex membrane compartment, LVCV large virus-containing vesicle.