A research group from the US (led by experts from Yale University School of Medicine) recently presented a case of an immunocompromised female patient with acquired deficiency of B lymphocytes who subsequently developed a protracted course of the infection with the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The research paper is currently available on the medRxiv* preprint server while it undergoes peer review.
As the vaccine rollout strategy against the coronavirus disease 2019 (COVID-19) is keeping pace in many countries around the world, we are yet to see a highly effective drug. Even though there is no consensus on the use of remdesivir, it is still a common choice for patients hospitalized with the disease.
Moreover, remdesivir has also been a go-to option for treating protracted cases of SARS-CoV-2 infection in immunocompromised individuals, where various case reports in the medical literature have found that the antiviral drug showed transient clinical improvement.
One advantage thus far has been no resistance in patients treated with remdesivir, although they have been observed in different in vitro studies. The latter phenomenon was also used to suggest the evolutionary predictability of SARS-CoV-2.
It is then of utmost importance to identify resistance-prone viral strains, pinpoint exact mutations and monitor the emergence of such variants, which is why case reports akin to this new one from US researchers will be very valuable going forward.
How did the case unfold?
In this paper, the authors present a woman in her seventies who received a specific treatment protocol for non-Hodgkin's lymphoma (NHL), complicated by a low number of lymphocytes and a low level of antibodies. This was complicated by a SARS-CoV-2 infection, presenting as acute fever, cough, runny nose and a loss of smell.
Due to a persistent fever and computerized tomography findings, the patient was put on a 10-day course of remdesivir, which resulted in normalization of body temperature and inflammatory markers, as was as a radiological improvement. Nonetheless, monoclonal antibody therapy was instituted due to a recrudescence of viral shedding during and after treatment with remdesivir.
The researchers aimed to identify SARS-CoV-2 mutations that have been generated during the course of illness. Thus they used the patient's tissues and secretions from different sources and performed whole-genome sequencing on a next-generation sequencing platform.
The emergence of a novel mutation
In short, such detailed analysis of viral genomes identified the emergence of mutation harboring remdesivir resistance, known as E802D, which arose in the immunocompromised patient with persistent SARS-CoV-2 infection. More specifically, using patient specimens, the occurrence of the mutation has been temporally linked to the treatment with this drug.
It is then plausible that the treatment of the patient with remdesivir selected for variants that harbored E802D mutation. Furthermore, as the patient's cytopenia resolved after implementing monoclonal antibodies, a causal link between SARS-CoV-2 and the low number of cells in this setting has been established.
Finally, the case vividly illustrates the benefits of using monoclonal antibodies in immunocompromised patients that present with enduring SARS-CoV-2 infection. In this instance, they have cleared viral shedding, resolved the issue of smell, and improved levels of cells in the blood.
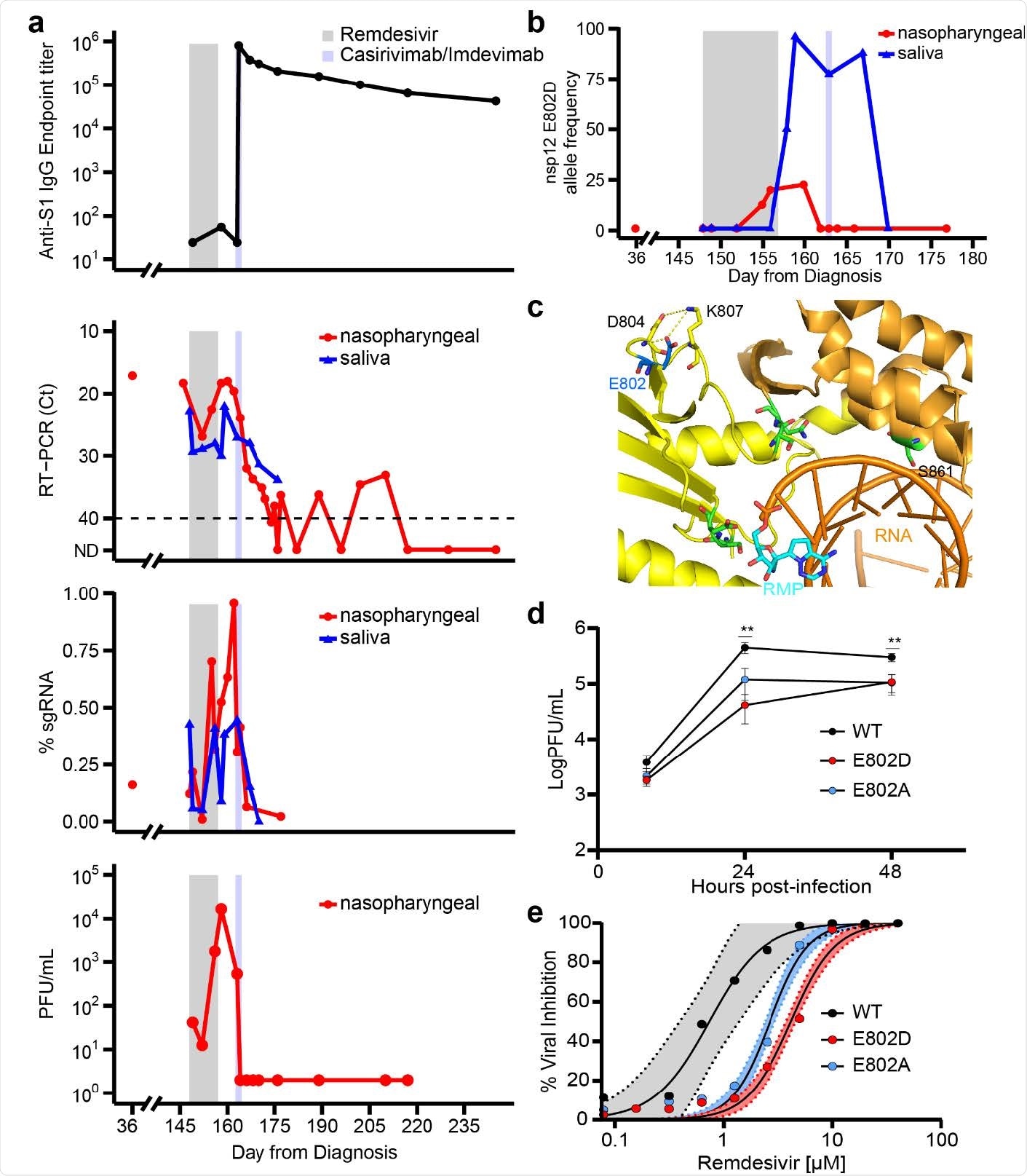
de novo emergence of remdesivir resistance mutation during and following treatment with the antiviral agent
Implications for genomic surveillance
Notwithstanding the fact that these insights are limited to a single case (which introduces generalizability issues), it still suggests that remdesivir can exert substantial selective pressures in vivo, which drives the evolution of the virus.
"E802D is associated with a fitness cost in vitro which may limit the broader impact of this mutation on the development of secondary resistance during treatment and the risk for primary resistance through the transmission of resistant variants", say the authors of this medRxiv paper.
"Yet, our findings underscore the importance of immunocompromised hosts with uncontrolled viral replication as a source of genetic diversification and selection of mutations that may potentially impart adverse consequences for antiviral therapy", they add.
Ergo, it may be important to include improved genomic surveillance of immunocompromised patients. Furthermore, the introduction of anti-SARS-CoV-2 monoclonal antibodies may be considered as a treatment option that aims to achieve a swift and sustained virologic response, as well as improve the overall clinical outcome.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Gandhi, S. et al. (2021). De novo emergence of a remdesivir resistance mutation during treatment of persistent SARS-CoV-2 infection in an immunocompromised patient: A case report. medRxiv. https://doi.org/10.1101/2021.11.08.21266069, https://www.medrxiv.org/content/10.1101/2021.11.08.21266069v1
- Peer reviewed and published scientific report.
Gandhi, Shiv, Jonathan Klein, Alexander J. Robertson, Mario A. Peña-Hernández, Michelle J. Lin, Pavitra Roychoudhury, Peiwen Lu, et al. 2022. “De Novo Emergence of a Remdesivir Resistance Mutation during Treatment of Persistent SARS-CoV-2 Infection in an Immunocompromised Patient: A Case Report.” Nature Communications 13 (1): 1547. https://doi.org/10.1038/s41467-022-29104-y. https://www.nature.com/articles/s41467-022-29104-y.