The unprecedented coronavirus disease 2019 (COVID-19) pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) originated in late December 2019 in Wuhan, China. SARS-CoV-2 rapidly spread throughout the world, leading to the introduction of lockdown measures in many countries.
In pandemic scenarios like SARS-CoV-2, rapid information aggregation is vital. In the era of eHealth and data-driven medicine, publicly available symptom tracking tools offer scalable and efficient means to collect and analyze large amounts of information. Consequently, front-line providers can be updated with knowledge gained.
Various improvements have been made in these applications in different countries during the SARS-CoV-2 pandemic.
The most common symptoms associated with the SARS-CoV-2-infection are fever, cough, and fatigue. Anosmia (loss of smell) was one of the first apparent symptoms that varies significantly in COVID-19 patients.
Various studies have drawn attention towards extrapulmonary symptoms and symptom constellations. For example, one study found that a less common symptom, such as anosmia, combined with the more established symptoms like fever and cough, were the strongest predictors of COVID-19.
Development of the COVID–Online application
In a research paper published in the journal PLOS ONE, the authors developed COVID–Online, a symptom checking application in Germany. The main objective of this COVID-Online app was patient care navigation by guiding the participant through the next steps in case of increased risk, which serves as a guidance system for the regional health authority to enable an efficient allocation of resources in case of an outbreak.
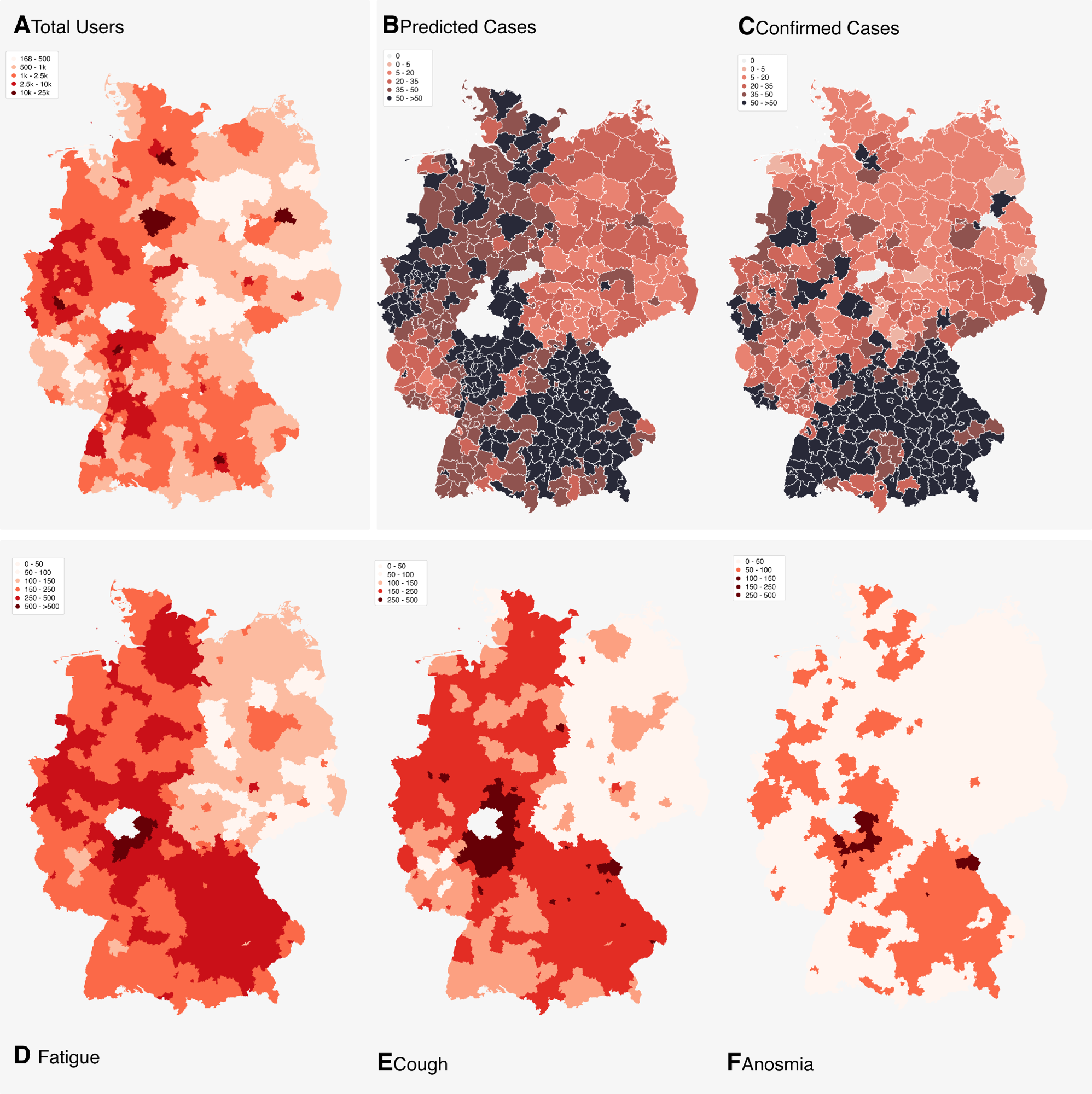 Maps illustrating the total number of users of COVID-Online (A), the number of predicted infections (B), the number of confirmed infections (C), the frequency of fatigue (D), the frequency of anosmia (E) and the frequency of fever (F). Data of maps A–F is based on the time period 03.04–10.04.2020. A, B, D, E, F: The district “Marburg-Biedenkopf” has been excluded from these charts as it contained too many records from internal tests carried out by associated personnel of COVID-Online and was also influenced by regional media reports. Unfortunately, due to the great time pressure in times of crisis, no test or staging instance could be installed for such purposes.
Maps illustrating the total number of users of COVID-Online (A), the number of predicted infections (B), the number of confirmed infections (C), the frequency of fatigue (D), the frequency of anosmia (E) and the frequency of fever (F). Data of maps A–F is based on the time period 03.04–10.04.2020. A, B, D, E, F: The district “Marburg-Biedenkopf” has been excluded from these charts as it contained too many records from internal tests carried out by associated personnel of COVID-Online and was also influenced by regional media reports. Unfortunately, due to the great time pressure in times of crisis, no test or staging instance could be installed for such purposes.
The COVID-Online application had a questionnaire with 38 items, including ten adaptive items. The questions consisted of a single choice, multiple-choice, and free text input fields. In addition, the questionnaire included three segments: basic epidemiological data, current symptoms, and individual risk factors, along with their postal code.
Triple combination symptoms are more specific for COVID-19
A linear regression model from the literature using the same symptom combination as features was applied to the data.
The data from the study revealed that the triple combination symptoms of anosmia, fever, and cough were most important in differentiating between groups of participants with and without confirmed contacts. However, single symptoms such as anosmia, cough, or fatigue alone were not sufficient.
Also, the frequencies of fall of symptoms were much faster in the group without confirmed contact than the frequencies of symptom combinations within the confirmed contact group. This indicated that single symptoms might be less suitable for the geographical identification of COVID-19 patients than the combination of multiple symptoms.
Furthermore, it was also suggested that in the group of confirmed contacts, more positive cases must exist. The frequency of predicted infections closely matched the regional distribution of confirmed cases across Germany, indicating that the northern federal states might be much more severely affected.
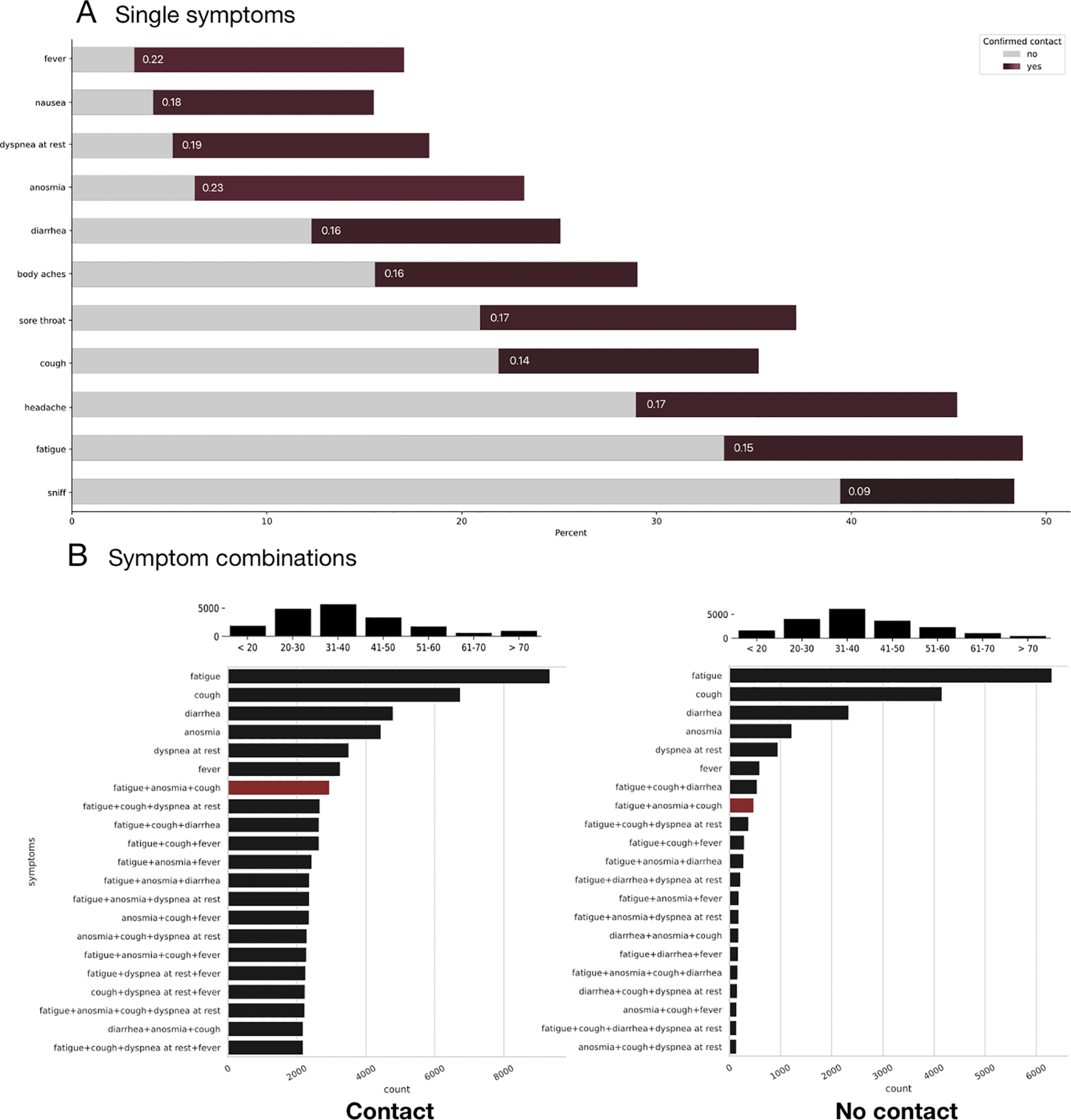 Comparison of symptom distribution between patients with and without confirmed contact. A) Single symptoms: Color-coded Cramer’s V correlation of symptoms with the confirmed contact variable (dark red tones) & symptom frequency count in percent of positive statements broken down by groups with and without confirmed contact. Anosmia seems to be the strongest predictor followed by fever and dyspnea at rest. On the contrary, the least single important symptoms are sniff, fatigue, and cough by themselves. B) Complex symptoms: Symptom frequency count (total) with combinations. The symptom combination of fatigue, anosmia, and cough has been highlighted in red to illustrate the shift of importance between the two groups. The age distribution of the two groups is depicted in each case above graph B. A random sample of 19128 was taken from the population without confirmed contact for comparison. The percentage of positive cases in the total number of participants without contact was 6·24% whereas the percentage of positive cases in the group with confirmed contact was 23·21%.
Comparison of symptom distribution between patients with and without confirmed contact. A) Single symptoms: Color-coded Cramer’s V correlation of symptoms with the confirmed contact variable (dark red tones) & symptom frequency count in percent of positive statements broken down by groups with and without confirmed contact. Anosmia seems to be the strongest predictor followed by fever and dyspnea at rest. On the contrary, the least single important symptoms are sniff, fatigue, and cough by themselves. B) Complex symptoms: Symptom frequency count (total) with combinations. The symptom combination of fatigue, anosmia, and cough has been highlighted in red to illustrate the shift of importance between the two groups. The age distribution of the two groups is depicted in each case above graph B. A random sample of 19128 was taken from the population without confirmed contact for comparison. The percentage of positive cases in the total number of participants without contact was 6·24% whereas the percentage of positive cases in the group with confirmed contact was 23·21%.
Limitations of the study
There are a few limitations associated with this work. The major limitation was the absence of data on confirmatory laboratory test results of the study participants, which restricted the adoption of a model proposed in the literature.
Less tech-savvy individuals were not included in the study, which created a contributor bias and also ruled out the fact that certain symptom constellations might be more common in such individuals.
Since the data was self-reported, the validity of the data could not be ascertained. Also, only snapshots of symptoms could be collected due to data security and privacy, which cannot be tracked over time.
Moreover, as the users can access the web app, there might be a chance of submitting more than one questionnaire by the same individual.
Conclusions
The study data confirmed that the triple combination symptoms of anosmia, fatigue and cough are indeed more specific for COVID-19 than single symptoms alone and are sufficient to recommend self-quarantine.
The results also demonstrated that the crowdsourced data could support traditional public health surveillance methods by providing information on SARS-CoV-2-infected individuals in a wide area. In the future, the validity of data from online crowdsourcing platforms can be increased by also considering the diagnostic test status.
“The near real-time analysis of symptom tracking applications might help to more accurately understand the spread and infectivity of infectious diseases such as SARS-CoV-2.”