Researchers from the United States have now found that the immune recall induced by a two-dose primary series of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) vaccination after an initial natural infection highly improves the longevity of the antibodies as well as induces an enhanced breadth of protection across the SARS-CoV-2 variants. The study is published in the journal Science Immunology.
The team also found that a homologous 3rd mRNA vaccine dose in naïve individuals [with no history of coronavirus disease 2019 (COVID-19) infection before vaccination] induces superior and more consistent cross-variant neutralization potency with durability similar to that induced by hybrid immunity from infection plus vaccination.
In addition, the team found that a recall of endemic coronavirus immune memory induced by SARS-CoV-2 infection improved the longevity of immune protection.
 Study: Immune recall improves antibody durability and breadth to SARS-CoV-2 variants. Image Credit: Billion Photos / Shutterstock
Study: Immune recall improves antibody durability and breadth to SARS-CoV-2 variants. Image Credit: Billion Photos / Shutterstock
Background
Antibody-producing-B cells can evolve to better recognize the repeatedly attacking pathogen through B cell clonal selection and somatic hypermutation (SHM), subsequently producing a gush of antibodies immediately upon the next pathogen encounter. B cells can differentiate into either long-lived plasma cells (LLPCs) or memory B cells, of which the memory B cells differentiate into antibody-secreting plasma cells upon subsequent pathogen encounter, while the already existing specific antibodies produced from LLPCs provide instant defense.
While the vaccines against COVID-19, to a greater extent, have successfully controlled the overwhelming situation created by the pandemic, the emergence of new viral variants with immune escape potential and gradually waning immune responses are now becoming matters of more significant concern on this front.
Therefore, it is imperative to understand how far the antibody memory spectrum stretches across the viral variants, how repeated invasions influence the durability of specific antibody responses, and how available vaccines can elicit them.
The team designed the present study to evaluate the virus-specific antibody longevity, cross-variant breadth, and evenness of cross-variant neutralization (neutralization breadth per unit function) after either SARS-CoV-2 infection, mRNA vaccination, or mRNA vaccination after SARS-CoV-2 infection.
What did the researchers do?
The study was conducted on three population cohorts:
- COVID-19 convalescents without vaccination followed up for >200 days
- COVID-19 convalescents followed up without vaccination for >200 days and later for an additional >200 days after mRNA vaccination (COVID-19 vaccinees)
- COVID-19-naive individuals/ naive vaccinees with mRNA vaccination followed up for >200 days.
Pseudotype virus neutralization assays were performed on plasma to examine the antibody breadth across the SARS-CoV-2 variants.
The team calculated a breadth index to assess the evenness of cross-variant neutralization over time. Antibody durability indexes over time were also generated.
In order to assess antibody cross-reactivity to other endemic coronaviruses, the team sorted spike S+ memory B cells and cloned respective antibody genes to purify monoclonal antibodies (mAb) from sustainers and decayers, respectively, around 40- and 110-days post symptom onset.
What did the researchers find?
Compared to COVID-19 convalescents around 40 days post symptom onset, both COVID-19 vaccinees and naive vaccinees had higher anti-S and anti-RBD IgG peak levels. However, later, around >220 days, COVID-19 vaccinees had significantly higher anti-S and anti-RBD IgG than naive vaccinees and COVID-19 convalescents.
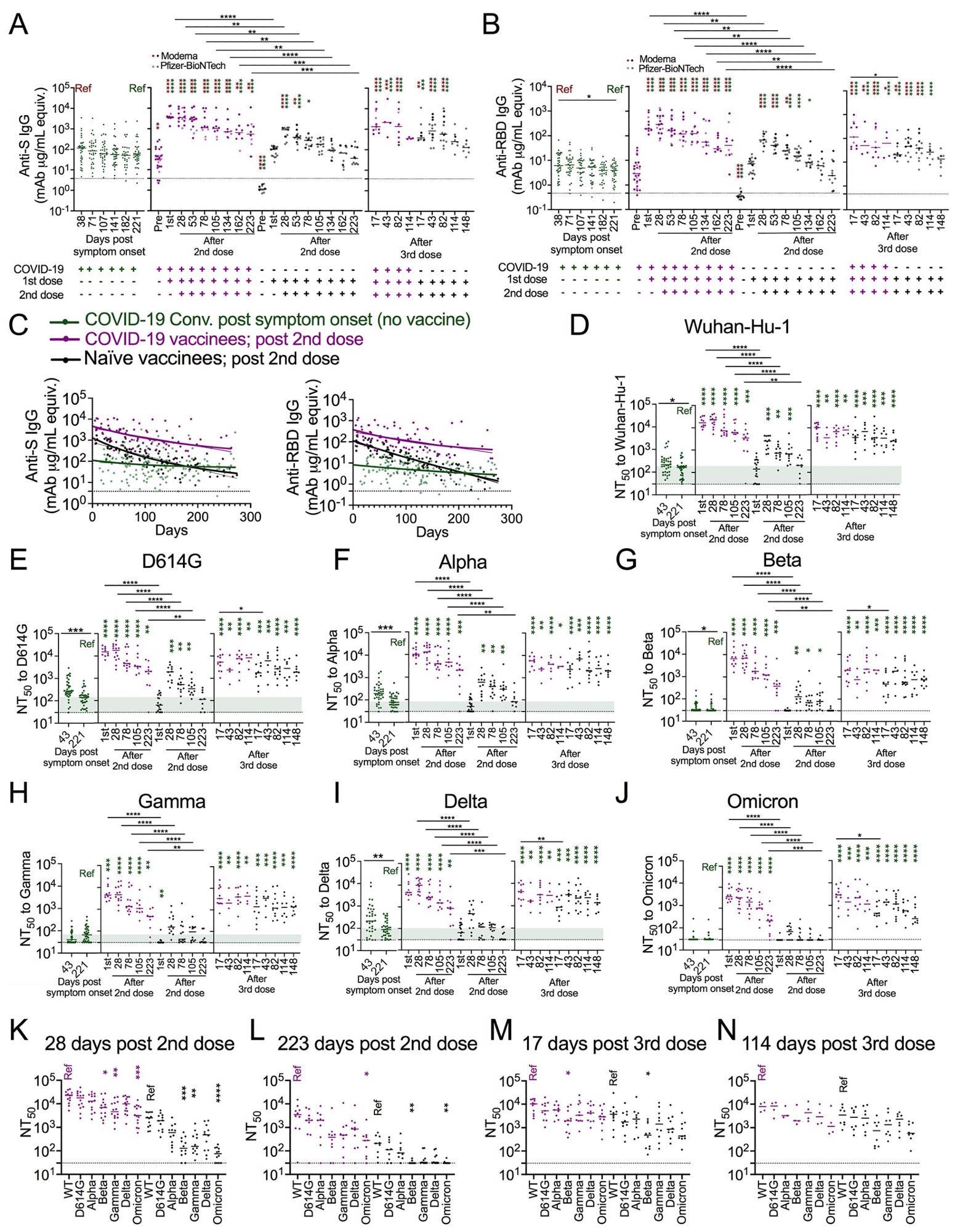 Immune recall by vaccination after prior infection confers greater antibody magnitude and stability. (A, B) Dot plots showing anti-spike (S) (A) and anti-RBD (B) IgG antibody levels in seroconverted COVID-19 convalescents (green, n = 28-34, left), COVID-19 vaccinees (purple, n = 4-28), and naïve vaccinees (black, n = 8-18) over time as indicated. For vaccinees, ANOVA with Tukey’s multiple comparison test was performed to test significant difference of log-transformed antibody data. (C) Dot and line graphs showing anti-S IgG (left) and anti-RBD IgG (right) trajectories in COVID-19 convalescents after natural infection (green, n = 34), and COVID-19 vaccinees (purple, n =28) as well as naïve vaccinees (black, n = 18) after the 2nd vaccine dose. Overlapping fitted curves for both one phase decay (thick line) and linear regression (thin line) models are shown. Green line indicates one-phase decay model and linear regression model fitted curves for log transformed antibody data from the COVID-19 convalescents. The fitted curves from the three groups were significantly different with each other (Extra sum-of-squares F Test, p<0.0001). The Dashed lines represent twice the average of pre-COVID (negative) controls. Extra sum-of squares F test found no significant differences between one phase decay and simple linear regression fitted curves in all cases except anti-S IgG log transformed data from naïve vaccinees. Extra sum of squares F test was used to test for significance of differences between the one phase decay model fitted curves. Analysis of Covariance (ANCOVA) was performed to test the significance of differences between slopes generated from linear regression. (D-J) Dot plots showing 50% pseudovirus neutralization titers (NT50) to indicated SARS-CoV-2 variants in plasma collected from COVID-19 convalescents after natural infection (green, n = 34), and COVID-19 vaccinees (purple, n =3-14) as well as naïve vaccinees (black, n = 6-18) after 2nd and 3rd vaccination as indicated. Kruskal Wallis test and Mann-Whitney U test. (K-N) Dot plots showing NT50 across all the tested variants in plasma collected from COVID-19 vaccinees (purple, n =3-14) and naïve vaccinees (black, n = 8-12) at the time as indicated. Kruskal-Wallis test. Dashed lines in A-C represent twice the average of negative controls, and dashed lines in D-M represent the limit of neutralization detection (i.e., 30). Green thick ribbon indicates the median neutralization level in COVID-19 convalescents at median day of 221 days after symptom onset. Colored asterisks indicate comparisons to the corresponding colored “Ref” in each panel. *p<0.05, **p<0.01 ***p<0.001; ****p<0.0001
Immune recall by vaccination after prior infection confers greater antibody magnitude and stability. (A, B) Dot plots showing anti-spike (S) (A) and anti-RBD (B) IgG antibody levels in seroconverted COVID-19 convalescents (green, n = 28-34, left), COVID-19 vaccinees (purple, n = 4-28), and naïve vaccinees (black, n = 8-18) over time as indicated. For vaccinees, ANOVA with Tukey’s multiple comparison test was performed to test significant difference of log-transformed antibody data. (C) Dot and line graphs showing anti-S IgG (left) and anti-RBD IgG (right) trajectories in COVID-19 convalescents after natural infection (green, n = 34), and COVID-19 vaccinees (purple, n =28) as well as naïve vaccinees (black, n = 18) after the 2nd vaccine dose. Overlapping fitted curves for both one phase decay (thick line) and linear regression (thin line) models are shown. Green line indicates one-phase decay model and linear regression model fitted curves for log transformed antibody data from the COVID-19 convalescents. The fitted curves from the three groups were significantly different with each other (Extra sum-of-squares F Test, p<0.0001). The Dashed lines represent twice the average of pre-COVID (negative) controls. Extra sum-of squares F test found no significant differences between one phase decay and simple linear regression fitted curves in all cases except anti-S IgG log transformed data from naïve vaccinees. Extra sum of squares F test was used to test for significance of differences between the one phase decay model fitted curves. Analysis of Covariance (ANCOVA) was performed to test the significance of differences between slopes generated from linear regression. (D-J) Dot plots showing 50% pseudovirus neutralization titers (NT50) to indicated SARS-CoV-2 variants in plasma collected from COVID-19 convalescents after natural infection (green, n = 34), and COVID-19 vaccinees (purple, n =3-14) as well as naïve vaccinees (black, n = 6-18) after 2nd and 3rd vaccination as indicated. Kruskal Wallis test and Mann-Whitney U test. (K-N) Dot plots showing NT50 across all the tested variants in plasma collected from COVID-19 vaccinees (purple, n =3-14) and naïve vaccinees (black, n = 8-12) at the time as indicated. Kruskal-Wallis test. Dashed lines in A-C represent twice the average of negative controls, and dashed lines in D-M represent the limit of neutralization detection (i.e., 30). Green thick ribbon indicates the median neutralization level in COVID-19 convalescents at median day of 221 days after symptom onset. Colored asterisks indicate comparisons to the corresponding colored “Ref” in each panel. *p<0.05, **p<0.01 ***p<0.001; ****p<0.0001
The naïve vaccinees demonstrated a more striking antibody decay over time. Although anti-S and anti-RBD IgG antibodies were significantly higher in naive vaccinees than in the COVID-19 convalescents, the levels were not maintained for a long time and came down to long-term plateau levels similar to COVID-19 convalescents. The COVID-19 vaccinees, on the hand, demonstrated the highest level of peak antibodies among the three groups, as well as a more stable linear regression slope and plateau phase over time than naive vaccinees.
A homologous 3rd dose of mRNA vaccination (both Pfizer and BioNTech) in COVID-19 naive individuals were seen to boost the antibody levels and the cross-variant neutralization breadth, including Omicron that remained robust throughout median 148 days post 3rd dose.
Consistent with higher anti-S and anti- RBD IgG levels, the plasma of COVID-19 vaccinees also had a wider cross-variant neutralization function than plasma from naive vaccinees and unvaccinated COVID-19 convalescents at all time points.
COVID-19 convalescents who demonstrated greater breadth indexes for Gamma and Delta than naive vaccinees also demonstrated greater breadth indexes for Beta and Omicron after vaccination.
“It is possible that replicating intact viruses in natural infection may elicit more even cross-variant anti-spike antibody recognition coverage than the coverage elicited by the stabilized version of spike in current vaccines,” suggests the team.
With the third dose of mRNA vaccine, however, all of the SARS-CoV-2 variants were significantly covered, reaching levels that resembled hybrid immunity in COVID-19 vaccinees.
As compared to the decayers with anti-S IgG durability index < 1, the long-term sustainers (anti-S IgG durability index ≥1) were observed to have milder disease, shorter symptom duration, and were mostly younger. For up to 270 days after symptom onset, the sustainers had relatively more anti-S and anti-RBD IgG than decayers.
An increased S+ memory circulating CD4+ T follicular helper cell frequencies were observed in anti-S IgG sustainers, pointing to a link between S-specific CD4+ T cell responses and anti-S IgG durability.
Monoclonal binding analysis demonstrated that sustainers also possessed significantly higher frequencies of memory B cells reactive to endemic human coronaviruses HKU1 and OC43 spikes, which tended to have higher SHM with high avidity to S. Additionally, sustainers were seen to have a significantly higher frequency of memory B cells binding to S2, which is highly conserved across coronaviruses.
At approximately 110 days after symptom onset, S2 and endemic coronavirus cross-reactive clones tended to decrease, while S1 and RBD-binding clones increased.
“Data suggest that early cross-reactive memory B cells recalled from prior endemic coronavirus infection are linked to stronger antibody durability over time,” highlights the team.
Cross-variant neutralization was observed to be superior in sustainers than decayers and was suggested to be linked to anti-S and anti-RBD antibody levels through correlation analysis. As time progressed, however, vaccination after infection normalized antibodies and cross-variant neutralizations in sustainers and decayers.