Researchers in the U.S. have used a new web-based platform to help identify synergistic and antagonistic drug combinations and their underlying mechanisms of action in the context of coronavirus disease 2019 (COVID-19).
The tool, called COVID-KOP, prioritized 73 combinations of 32 drugs as potential treatments for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), allowing the team then to identify 16 synergistic combinations and eight antagonistic combinations.
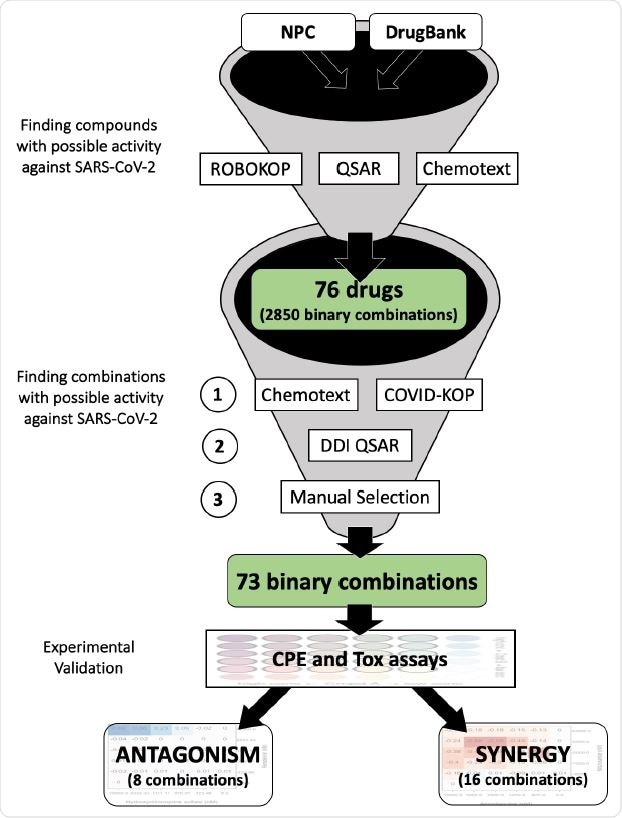
Study design for selecting possible synergistic drug combinations. In this study we report only 73 binary combinations. 95 ternary combinations identified in a similar fashion will be reported in a future study.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
“COVID-KOP, a tool recently developed by our group, integrates existing biochemical knowledge with that contained in literature on COVID-19 in the form of knowledge graphs,” say the researchers from the University of North Carolina and the National Center for Advancing Translational Sciences.
“Overall, our results emphasize the importance of both drug repurposing and preclinical testing of drug combinations for potential therapeutic use against SARS-CoV-2 infections,” writes the team.
A pre-print version of the paper can be accessed in the server bioRxiv*, while the article undergoes peer review.
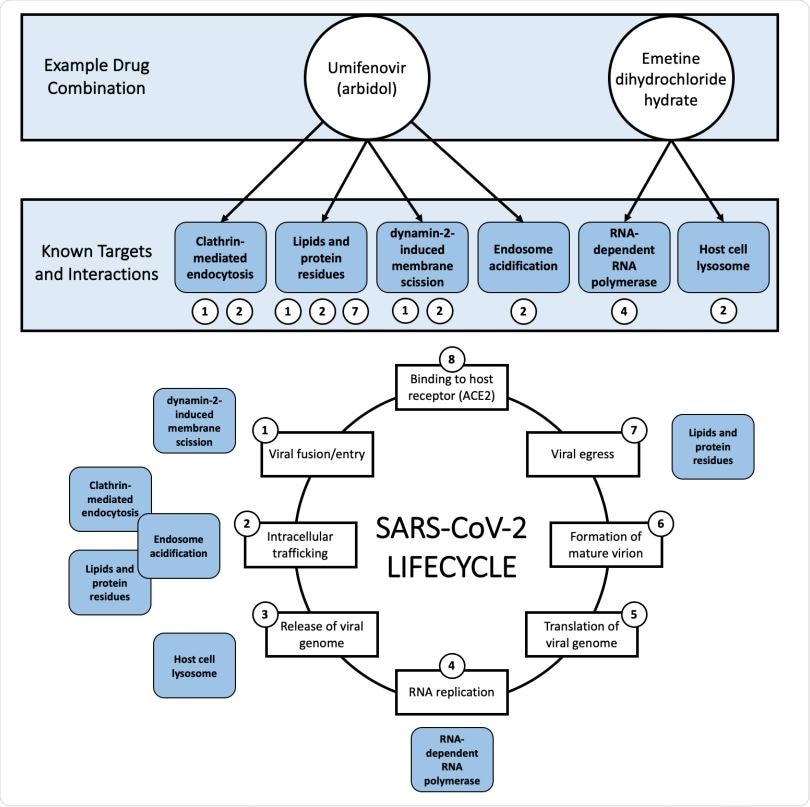
Example (umifenovir + emetine dihydrochloride hydrate) of the rationale behind mixture selection, i.e., interference with different steps of the COVID-19 lifecycle. All known targets and interactions were taken from the literature and are not necessarily specific to SARS-CoV-2. Umifenovir’s proposed mechanisms of action involve clathrin-mediated endocytosis, lipids and protein residues, dynamin-2-induced membrane scission, and endosome acidification. Emetine’s proposed mechanisms of action involve RNA replication/RNA-dependent RNA polymerase and the host cell lysosome. The viral lifecycle of SARS-CoV-2 was inspired by Fig. 1. of da Costa et. al 2020.
Drug combinations for treating viral infection
Since the COVID-19 outbreak began in Wuhan, China, late last year, the pandemic has affected millions of people worldwide, infecting more than 10.9 million and causing more than 519,000 deaths.
“COVID-19 is undoubtedly the most impactful viral disease of the current century,” said Eugene Muratov and colleagues.
Drug combinations have previously proved particularly effective at treating viral infections, including HIV, the Ebola virus, and poliovirus. These combinations significantly lower the risk of resistance developing to any one drug and can provide synergy – where the antiviral action of the combination is more potent than that of one of the drugs alone.
Although many current or upcoming trials are testing drug combinations for the treatment of COVID-19, few have completed extensive preclinical studies before being administered to patients.
More investigation into these regimens is needed so that synergistic combinations can be more efficiently prioritized and any antagonistic effects identified prior to preclinical testing.
Prioritizing drug combinations using COVID-KOP
Muratov and the team recently used data and text mining techniques to identify 281 combinations of 38 drugs for potential repurposing against SARS-CoV-2.
Using their new COVID-KOP web-platform, the researchers refined the original list and prioritized 73 combinations of 32 before testing them using cytopathic and cytotoxicity assays.
Of the 16 combinations showing synergy against SARS-CoV-2, the most notable were combinations of the FDA-approved drug nitazoxanide with three other agents: remdesivir, amodiaquine, and umifenovir.
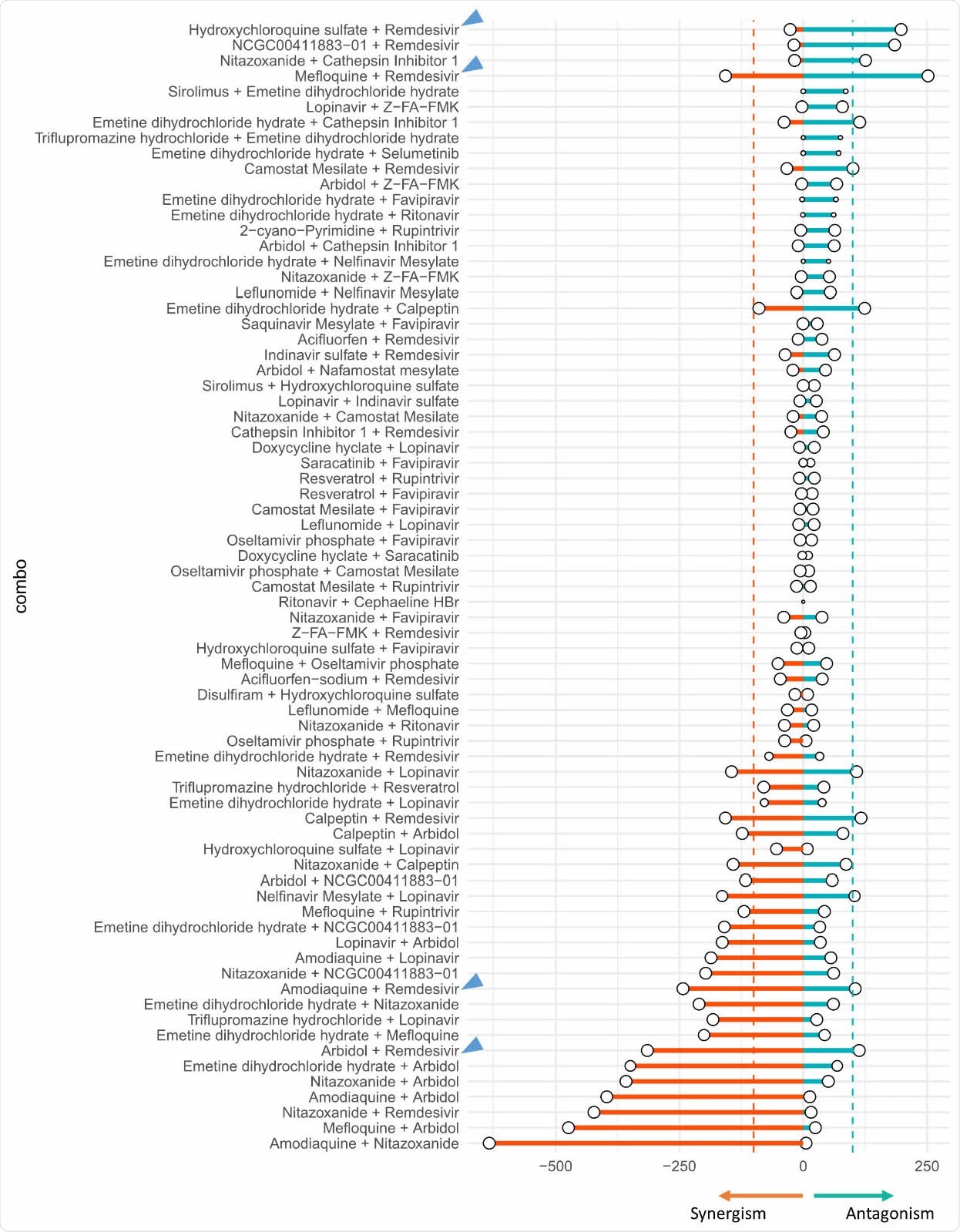
Summary of synergism or antagonism across 73 tested combinations. Due to biphasic dose response, synergism was separated from antagonism. Synergism is calculated as the sum of HSA.neg values from non-toxic dose combinations (Tox > 50%), and vice versa. The size of circle reflected the confidence of the observed synergism/antagonism (bigger circle = less doses were excluded due to toxicity).The inconclusive blocks (Nnontoxic < 25 or rough activity landscape) were shaded. Two dashed lines indicated the cutoff of HSA synergism (-100) or antagonism (100). Blue arrows highlighted the combinations between remdesivir and tertiary amine compounds from conclusive blocks.
Nitazoxanide with remdesivir looks promising from a clinical perspective
Among these combinations, a strong synergistic interaction was observed for nitazoxanide with remdesivir.
Nitazoxanide is a broad-spectrum anti-infective agent that has recently been explored for activity against SARS-CoV-2 due to its previously demonstrated antiviral activity.
In studies of the Ebola virus, nitazoxanide has been shown to activate protein kinase-R, which induces type-I interferon production and signaling.
“Current knowledge suggests an interferon-inducing mechanism of nitazoxanide in vitro,” say Muratov and team. “SARS-CoV-2 is known for its unique immunopathology of reduced production of, but extra sensitivity to, type-I/III interferon.”
The researchers say that as of June 18th, 2020, 18 trials of nitazoxanide for COVID-19 were registered with clinicaltrials.gov.
Moreover, nitazoxanide with remdesivir seems to be the most hopeful combination from a clinical perspective, since nitazoxanide is already available as an FDA-approved drug and remdesivir have been approved with FDA Emergency Use Authorization (EUA) for the treatment of COVID-19.
“The most striking antagonism”
The researchers say that among the eight antagonistic interactions they identified, the most notable and unexpected combination was remdesivir with hydroxychloroquine, which is currently undergoing clinical trials.
“The most striking antagonism was observed in the combination of the only two drugs ever approved with FDA Emergency Use Authorization (EUA): hydroxychloroquine and remdesivir (though we note the EUA for hydroxychloroquine has been withdrawn by the FDA),” writes the team.
Further analysis using COVID-KOP suggested that the mechanism of action for this antagonism may involve pathways related to in viral entry/egress from cells.
A literature search also showed that hydroxychloroquine inhibits an enzyme called glycogen synthase kinase-3β (GSK-3β), which regulates viral replication in the cases of dengue virus and SARS-CoV-1.
“Thus, our computational analysis suggests that hydroxychloroquine’s inhibition of GSK-3β may play a role in its antagonism of remdesivir,” writes the team.
The importance of preclinical research
Muratov and colleagues say their study shows the importance of conducting preclinical research into antiviral drug combinations before applying them to patients, as well as the value of data and text mining in exploring the mechanisms underlying synergism and antagonism within the context of COVID-19.
“Altogether, our findings demonstrate the utility of in silico tools for rational selection of drug combinations, the importance of preclinical testing of drug combinations prior to their administration in patients, and the overall promise of using drug repurposing and combination therapies against SARS-CoV-2,” concludes the team.
All the protocols and results reported in this study are freely accessible at https://opendata.ncats.nih.gov/covid19/matrix

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources