Based on structural modeling of ribonucleic acid (RNA) and proteins, the researchers from Duke University argue that newly identified regions of positive selection in the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) likely affect species-specific RNA instead of protein function – improving our understanding of the molecular mechanisms responsible for highly unique biological properties of this virus. The study is currently available on the bioRxiv* preprint server.
.jpg)
Transmission electron micrograph of SARS-CoV-2 virus particles, isolated from a patient. Image captured and color-enhanced at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. Credit: NIAID
The pandemic spread of a novel coronavirus (SARS-CoV-2) linked to coronavirus disease (COVID-19) has incited efforts to understand the genetic basis for its unique traits and species exchange from non-primate hosts to humans. One of the crucial characteristics is that viral shedding starts before symptom onset; conversely, in the original SARS epidemic in 2002-2004, viral shedding began 2-10 days after the onset of symptoms.
This striking difference, ultimately responsible for the global spread of SARS-CoV-2, indicates that one or more molecular mechanisms during cell invasion, viral replication, or immune evasion may have changed.
In any case, mutations aiding viral transmission would likely be favored by natural selection, which makes tests for positive selection a rather useful tool for pinpointing candidate genetic changes responsible for such unique properties of SARS-CoV-2.
Several recent publications around the SARS-CoV-2 genome have found signals of positive selection and conservation within the gene encoding spike glycoprotein, based on the ratio of synonymous to nonsynonymous substitution. Nonetheless, such tests cannot detect changes in the function of RNA molecules.
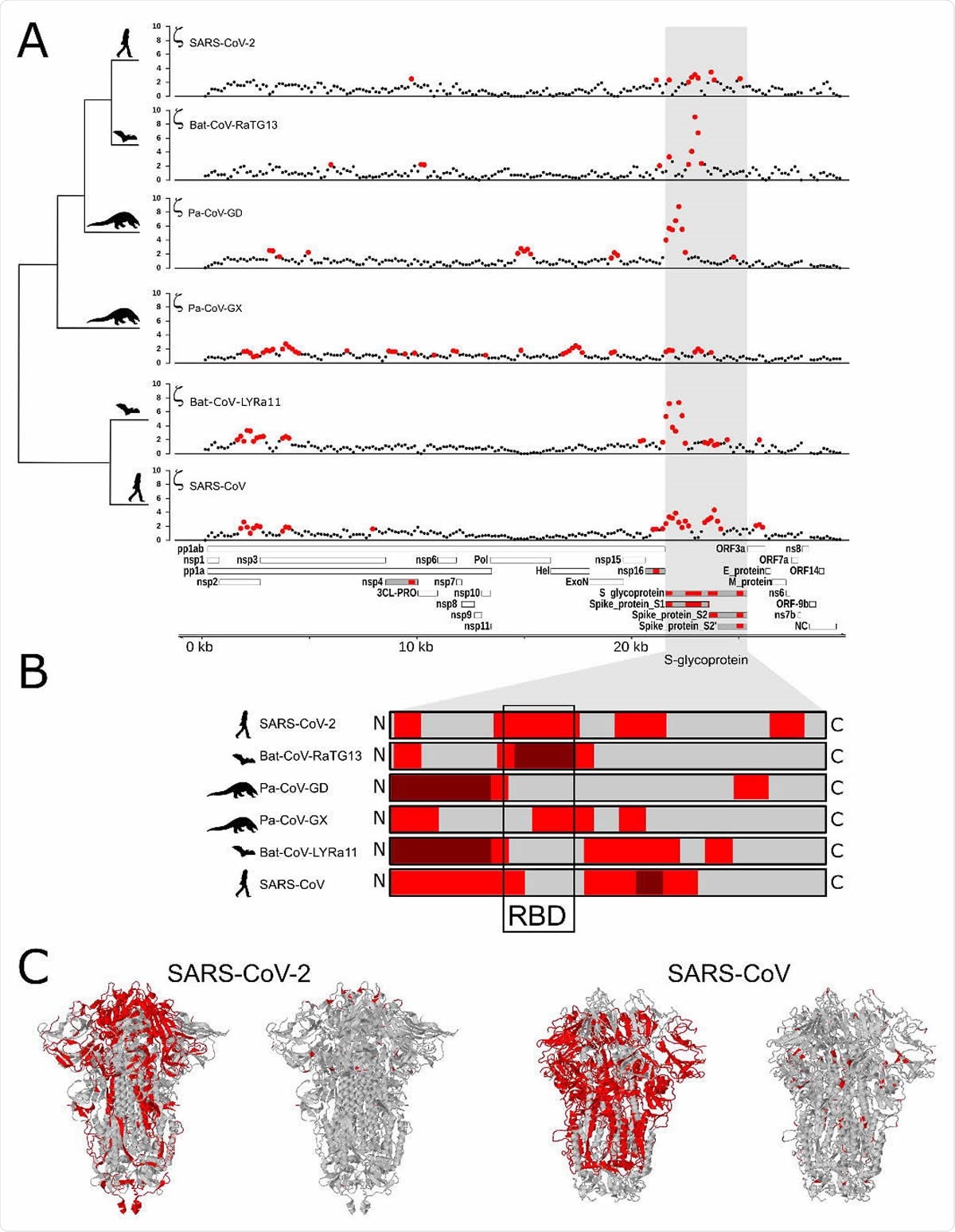
In this exciting new paper, Dr. Alejandro Berrio, Dr. Valerie Gartner, and Dr. Gregory A Wray from Duke University in Durham, North Carolina (United States) hunted for regions of possible and probable positive selection within the genomes of six coronavirus species – including SARS-CoV and SARS-CoV-2.
Chasing mutations and conserved regions
In this study, the researchers have applied a test for branch-specific oversubstitution of mutations within confined windows of the genome and without reference to the genetic code. In other words, the method they have used tests for a surplus of branch-specific nucleotide substitutions within a designated window relative to a neutral expectation for divergence in that window.
They have tested for branch-specific selection on nucleotide sequences in coronavirus genomes, focusing their attention on six species from the Sarbecovirus subgenus from the Coronaviridae family, as well as on Bat coronavirus BM48-31/BGR/2008 as an outgroup.
To gain comprehensive insights into the evolutionary mechanisms that have shaped genetic variation within the SARS-CoV-2 genome relatively recently, they have compiled a list of known mutations, primarily based on accessions sequenced since the start of the ongoing COVID-19 pandemic.
Furthermore, highly conversed regions of the genome across the sarbecovirus genomes were examined with the use of PhastCons, which is a special type of statistical model/program to identify and score conserved elements.
Adaptive modification of the SARS-CoV-2 genome
"The results reveal several signals of positive selection that are unique to a single species and others that are recapitulated in multiple species," say study authors. "The latter finding suggests that some segments of the viral genome have repeatedly experienced adaptive modification," they add.
Generally speaking, the distribution of positive selection is more in line with closely related species when compared to divergent ones – implying that certain molecular functions have been modified over an interval that expands beyond the origin of a single species, although not across the entire Sarbecovirus expansion.
When mutations were concerned, the vast majority of variants were singletons, representing either mutations without segregation or potential sequencing errors. More specifically, the researchers identified Nsp4 and Nsp16 as prime regions of the SARS-Cov-2 genome with mutations that may contribute to its unique biological, pathological, and epidemiological features.
These mutations are highly specific for SARS-CoV-2 and may, in turn, affect molecular processes that are mediated by the positive or negative RNA molecules, which includes RNA stability, transcription, and translation processes, as well as the evasion of the host innate immune system.
Considering multiple perspectives
"By shining a light on regions of the SARS-CoV-2 genome that appear to be under positive selection yet are unlikely to alter protein function, our results illustrate the value of evaluating the potential for adaptive changes in secondary structures within the genomes of RNA viruses", conclude study authors in this bioRxiv paper.
Consequently, these results emphasize the importance of taking into account mutations in viral genomes not only from the perspective of their effect on protein structure but also how they may influence other molecular processes pivotal to the viral life cycle.
And while it is enticing to speculate about the possible adaptive role of RNA changes within Nsp4 or Nsp16 accelerated regions, the authors suggest that this should always be pursued in the context of robust experimental results.
For example, the primary sequence of the genome can be modified to encode the same protein sequence while disrupting secondary structure within the aforementioned regions, subsequently testing for the consequences in viral replication and molecular functions.
In any case, this specific study may inspire other researchers to conduct studies with the aim of better understanding the evolving functions of RNA secondary structure found within the SARS-CoV-2 genome. As a result, this may open the door for novel therapeutic and preventative measures against this pandemic virus.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Berrio, A., Gartner, V. & Wray, G.A. Positive selection within the genomes of SARS-CoV-2 and other Coronaviruses independent of impact on protein function. bioRxiv. https://doi.org/10.1101/2020.09.16.300038.
- Peer reviewed and published scientific report.
Berrio, Alejandro, Valerie Gartner, and Gregory A. Wray. 2020. “Positive Selection within the Genomes of SARS-CoV-2 and Other Coronaviruses Independent of Impact on Protein Function.” PeerJ 8 (October): e10234. https://doi.org/10.7717/peerj.10234. https://peerj.com/articles/10234/.