Viruses are essentially parasites that rely on host cell machinery to support their entry, replication, and the synthesis of viral progeny inside the cell. Inhibition of the host factors necessary for viral invasion is thus a potent strategy in the development of antiviral agents. Given the scarcity of effective antiviral drugs to fight the current COVID-19 pandemic, this is an urgent global necessity.
Many viruses, including severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) – the causative agent of coronavirus disease 2019 (COVID-19) – hijack the host protein synthesis mechanisms in order to synthesize and assemble their own viral proteins. Hence, small molecule inhibitors of viral protein entry into the endoplasmic reticulum (ER) have recently gained attention as potential broad-spectrum antiviral agents. Such inhibitors reduce viral pathogenicity through a dual approach – 1. by directly inhibiting the synthesis of key viral proteins; and 2. by reducing the levels of host proteins that inhibit viral infection.
The role of viral and host proteins in SARS CoV-2 infection and pathogenesis
SARS-CoV-2 infection involves the synthesis of 3 major structural proteins – the spike (S), envelope (E), and membrane (M) proteins – in the ER of infected host cells before the assembly of the new viral particles. Thus, the inhibition of membrane protein synthesis is a promising strategy for reducing the infectivity of SARS-CoV-2 and other similar viral pathogens.
In the case of SARS-CoV-2, the human angiotensin-converting enzyme 2 (ACE2) is one of the crucial host cell receptors for viral entry and invasion. Hence, ACE2 is also a potential candidate for down-regulation using small molecule inhibitors of ER entry.
Ipom-F blocks the translocation of the three vital protein targets for SARS-CoV-2 infection
Recently, a team of researchers from the University of Manchester, UK; Ball State University, Indiana, USA; and University of Maryland, USA, showed the small molecule inhibitor, ipomoeassin F (Ipom-F), blocks the translocation/insertion of the three protein targets for SARS-CoV-2 infection. Their study is published on the preprint server, bioRxiv*.
The researchers used an in vitro system to demonstrate that Ipom-F inhibits the Sec61-mediated ER membrane translocation/insertion of the SARS-CoV-2 spike and ORF8 proteins and the host cell plasma membrane receptor, ACE2.
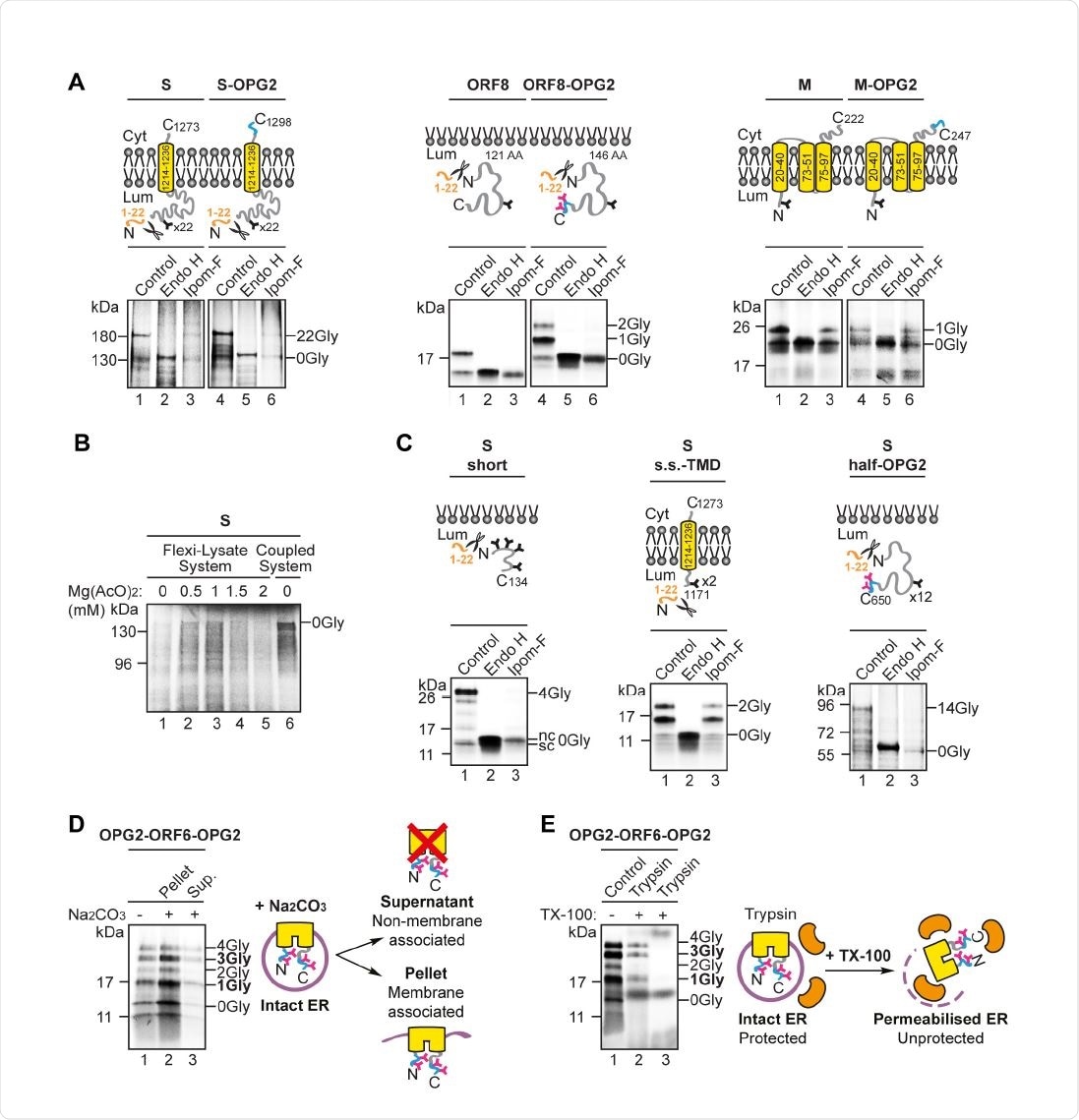
Ipom-F selectively inhibits the ER membrane translocation of SARS-CoV-2 proteins. (A) Schematic of the in vitro ER import assay in which canine pancreatic microsomes are used as a source of ER membrane. Following translation, fully translocatedor membrane inserted radiolabelled precursor proteins are recovered with the ER membrane fraction and analysed by SDS-PAGE and phosphorimaging. The identity of N-glycosylated species was confirmed by treatment with endoglycosidase H (Endo H). (B) Protein precursors of the human angiotensin-converting enzyme 2 (ACE2) and OPG2-tagged versions of the SARS-CoV-2 ORF8 (ORF8-OPG2), spike (S-OPG2), envelope (OPG2-E), membrane (M158 OPG2) and ORF6 (a doubly-OPG2 tagged version, OPG2-ORF6-OPG2, and two singly159 OPG2 tagged forms, OPG2-ORF6 and ORF6-OPG2, with their respective predominant N160 glycan species highlighted in bold) were synthesised in rabbit reticulocyte lysate perpetuity. It is made available under aCC-BY-ND 4.0 International license. preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in supplemented with ER-derived microsomes in the absence or presence of Ipom-F (lanes 1 and 3). Phosphorimages of membrane-associated products resolved by SDS-PAGE, together with representative substrate outlines, are shown. N-glycosylation was used as a read-out for the efficiency of membrane translocation/insertion and N-glycosylated (X-Gly) versus non-N165 glycosylated (0Gly) species were identified by treatment with Endo H (see lane 2). (C) The relative efficiency of membrane translocation/insertion in the presence of Ipom-F was calculated using the ratio of N-glycosylated protein to non-glycosylated protein, relative to the DMSO treated control (set to 100% efficiency). Quantifications are given as means ±SEM for independent translation reactions performed in triplicate (n=3) and statistical significance (one170 way ANOVA, DF and F values shown in the figure) was determined using Dunnett’s multiple comparisons test. Statistical significance is given as n.s., non-significant >0.1; ****, P < 0.0001.
Ipom-F sensitivity of the SARS-CoV-2 proteins is due to their dependence on the Sec61 translocation complex
The study findings highlight the utility of ER protein translocation inhibitors like Ipom-F as broad-spectrum, host-targeting, antiviral drugs. The researchers showed that the key molecular basis for the Ipom-F sensitivity of the SARS-CoV-2 proteins is their dependence on the Sec61 translocation complex. Overall, this in vitro study suggests that Ipom-F is a potent candidate for the development of antiviral agents that can inhibit SARS-CoV-2 protein synthesis.
“We conclude, that Sec61-selective protein translocation inhibitors like Ipom-F hold promise as broad-spectrum antivirals that may exert a therapeutic effect by selectively inhibiting the ER translocation of viral and/or host proteins which are crucial to viral infection and propagation.”
Ipom-F is a potent inhibitor of Sec61-mediated protein translocation and is well tolerated in mice
With respect to SARS-CoV-2, the integration of the S protein and its host cell receptor, ACE2, into the ER membrane is considerably reduced by Ipom-F. Similarly, the viral ORF8 protein translocation across the ER membrane is also significantly diminished. The viral S protein binding to host cell ACE2 is a key step in SARS-CoV-2 infection, while ORF8 protein may protect infected cells against cytotoxic T lymphocytes of the host. This makes all of these proteins possible therapeutic targets.
Since Ipom-F has been identified as a potent inhibitor of Sec61-mediated protein translocation in cell culture models and seems to be well tolerated in mice, the researchers propose that more studies should investigate the effects of Ipom-F on SARS-CoV-2 entry and propagation in the future.
“Taken together, our data establish that, analogous to human membrane and secretory proteins, the principal molecular basis for the Ipom-F-sensitivity of the SARS-CoV-2 ORF8 and S proteins is their dependence on Sec61-mediated protein translocation into and across the ER membrane,”

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
O'Keefe, Sarah and Peristera Roboti, Kwabena B Duah, Guanghui Zong, Hayden Schneider, Wei Q Shi, Stephen High. Ipomoeassin-F inhibits the in vitro biogenesis of the SARS-CoV-2 spike protein and its host cell membrane receptor. (2020) bioRxiv preprint server. doi: https://doi.org/10.1101/2020.11.24.390039, https://www.biorxiv.org/content/10.1101/2020.11.24.390039v1
- Peer reviewed and published scientific report.
O’Keefe, Sarah, Peristera Roboti, Kwabena B. Duah, Guanghui Zong, Hayden Schneider, Wei Q. Shi, and Stephen High. 2021. “Ipomoeassin-F Inhibits the in Vitro Biogenesis of the SARS-CoV-2 Spike Protein and Its Host Cell Membrane Receptor.” Journal of Cell Science 134 (4). https://doi.org/10.1242/jcs.257758. https://journals.biologists.com/jcs/article/134/4/jcs257758/224127/Ipomoeassin-F-inhibits-the-in-vitro-biogenesis-of.