The ongoing coronavirus disease 2019 (COVID-19), caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has resisted successful containment by most non-pharmaceutical measures. Vaccination and natural immunity appear to be the only definitive methods to attain population immunity, and thus lead to a return to global normalcy. However, effective therapies continue to be sought after to reduce the high mortality rate from severe COVID-19 disease.
The bogey of mutational escape
However, even as the first vaccines are being rolled out, new and more infective variants are emerging in different regions of the world. This could result in mutational escape from antibodies elicited by natural infection as well as vaccination. This underlines the need for effective pharmaceutical agents to treat the infection, and bring down mortality rates.
Some important mutations include K417E/N/T, E484K and N501Y, which increase transmissibility, and in some cases, the virulence of the virus. Other mutations confer escape from antibody neutralization, including N439K, Y453F, and G446V.
To retain the potency of neutralizing monoclonal antibodies, they were proposed to be used in cocktails, targeting distinct neutralizing epitopes and thus preventing selection of escape mutations. However, this pushes up the costs of manufacture and limits the supply.
DARPin platforms
The current study focused on the use of the DARPin platform to enable the rapid and inexpensive production of highly neutralizing multi-specific therapeutic agents against the virus. DARPins are small artificial scaffold proteins based on ankyrin repeats.
Ankyrins are scaffolding proteins that attach those proteins that form part of the cell membrane to the membrane cytoskeleton. Natural proteins built on ankyrin repeats bind many different targets. This allows organization of the cytoskeleton, as well as enabling the regulation of enzyme activity.
This feature was exploited to construct a motif that contains variable regions, and that can join with adjacent motifs to form one single rigid structure. DARPin molecules thus comprise several DARPin domains joined to form a single chain.
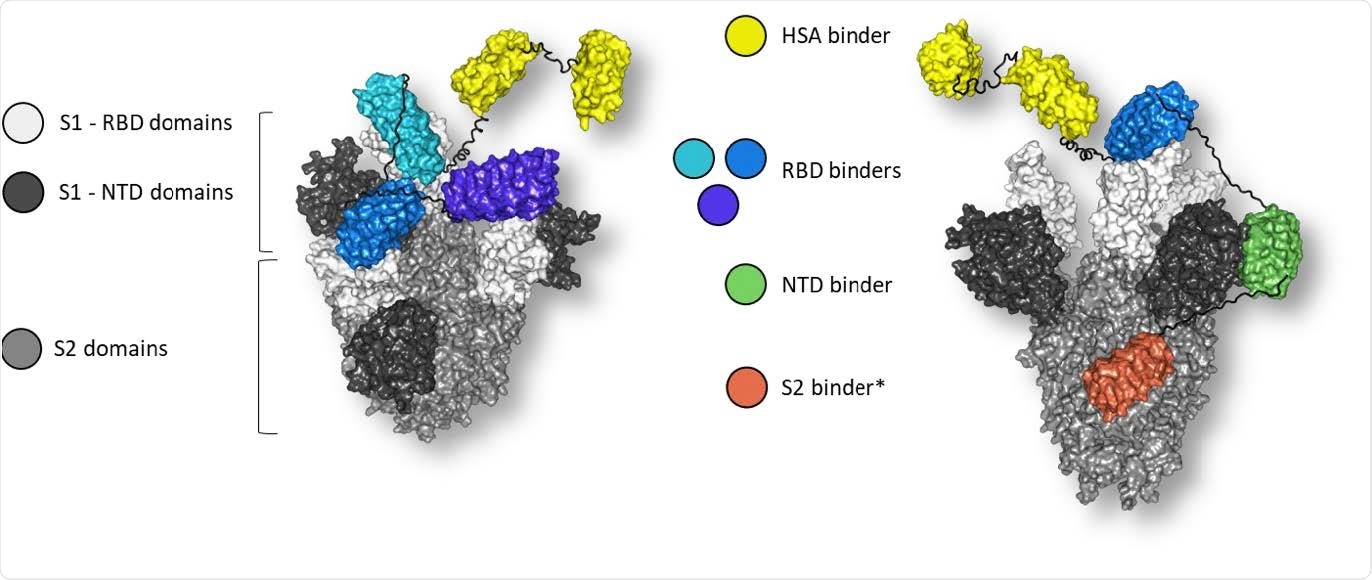
Molecular model of multi-specific DARPin® antivirals MP0423 (right), consisting of five DARPin® domains (yellow: HSA-binding domains, blue: RBD-binding domain, green: NTD-binding domain, orange: S2-binding domain) bound to the spike ectodomain. Linkers are shown in black. Molecular model of multi-paratopic DARPin® candidate ensovibep (left) consisting of five DARPin® domains (yellow: HSA-binding domains, shades of blue: RBD-binding domains) bound to the RBDs (white) of the spike ectodomain (grey). Linkers are shown in black. Position of RBD and NTD binders guided by Cryo-EM data. *Positioning on spike protein remains uncertain for the S2 binder and is guided by manual docking and not based on Cryo-EM data15.
DARPins form promising therapeutic approach
DARPin molecules are very thermostable, soluble and produce high yields, making them a very viable manufacturing option relative to antibody combinations. For the current application, the DARPin molecules were modified for a longer half-life by adding two N-terminal human serum albumin (HSA) binding domains.
The two DARPin therapeutics discussed here include ensovibep (MP0420) and MP0423. Ensovibep has three domains targeting the spike receptor-binding domain (RBD), while MP0423’s domains bind to the RBD, the N terminal domain (NTD) and the S2 domain.
With the latter molecule, the neutralization is not dependent on RBD binding alone. As a result, even if one of the three domains fails to recognize its epitope, as by an escape mutation, it does not lose its neutralization potency.
Neutralization potency
The study showed that both the DARPin molecules are very powerful neutralizing agents against the newer SARS-CoV-2 variants B.1.1.7 (UK variant) and B.1.351 (South African variant) in circulation. Ensovibep showed comparable binding efficiency to both variants compared to the wildtype virus, with the 50% inhibitory concentrations (IC50) being in the very low ng/mL range.
MP0423 showed a 20-fold lower potency against the UK variant but not the South African variant. However, the RBD-binding domain of this DARPin molecule had similar neutralization ability for the UK variant as the wildtype. Its neutralization potency is unaffected by loss of NTD binding. Therefore, the loss of potency for the UK variant must be because of S2 domain exposed mutations, either alone or acting together with mutations in the NTD.
Further studies are ongoing to elucidate this aspect. Nonetheless, even at reduced potency, MP0423 is still within the range of potency of many currently available monoclonal antibodies.
Neutralization of spike mutants
The DARPin platforms also neutralize the most common mutations in the spike protein. Notably, ensovibep, as well as the three individual RBD binders, failed to protect against the F486V mutation. F486 is required for spike binding to the human cell receptor, angiotensin-converting enzyme 2 (ACE2).
As a result, it is highly conserved by selection, and provides an anchoring element for ensovibep binding. The F486V mutation thus reduces the binding of all three ensovibep binding domains to the spike protein. It is currently found at low frequency in all circulating strains of the virus.
The triple epitope binding of the MP0423 molecule shows its value here, since the loss of RBD binding is overcome by the binding of the other two domains.
Prevention of escape mutations
When a virus is exposed to a monoclonal neutralizing antibody, escape mutants rapidly emerge by selection. However, in the current study, passaging of the virus through cultured cells demonstrated that both these agents prevented mutational escape. The extent of protection provided by either of them or both of them in combination was similar to that provided by the most potent antibody cocktails (REGN10933 & REGN10987).
When used in combination, the DARPin antivirals were effective at less than 0.1 μg/mL, while in isolation, the required concentration was 2 and 10 μg/mL for ensovibep and MP0423, respectively. In contrast, the Regeneron monoclonal antibody cocktail REGN10933 & REGN10987, which is currently in use under its emergency use authorization (EUA) status, was effective at 0.4 μg/mL.
The researchers emphasize that the manner in which the three binding domains in each DARPin molecule act together to neutralize the viral spike protein is central to its potent neutralizing activity.
What are the implications?
The inherent manufacturing advantages of the DARPin platforms, coupled with the resistance conferred by the tripartite binding of these agents, suggest that they are extremely promising leads for the development of very effective neutralizing therapeutic agents.
These results suggest ensovibep and MP0423 as superior alternatives to monoclonal antibody cocktails for global supply and demonstrate the strength of the DARPin® platform for achieving potent and lasting virus inhibition for SARS-CoV-2 and possibly other viruses,” say the researchers.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Rothenberger, S. et al. (2021). Multi-specific DARPin® therapeutics demonstrate very high potency against mutated SARS-CoV-2 variants in vitro. bioRxiv preprint server. doi: https://doi.org/10.1101/2021.02.03.429164, https://www.biorxiv.org/content/10.1101/2021.02.03.429164v1
- Peer reviewed and published scientific report.
Rothenberger, Sylvia, Daniel L. Hurdiss, Marcel Walser, Francesca Malvezzi, Jennifer Mayor, Sarah Ryter, Hector Moreno, et al. 2022. “The Trispecific DARPin Ensovibep Inhibits Diverse SARS-CoV-2 Variants.” Nature Biotechnology, July. https://doi.org/10.1038/s41587-022-01382-3. https://www.nature.com/articles/s41587-022-01382-3.