Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a novel pathogen that emerged in late 2019 and caused the ongoing coronavirus disease 2019 (COVID-19) pandemic. Coronaviruses (CoV) are single-stranded with positive-sense RNA genomes. Severe COVID-19 causes increased levels of pro-inflammatory cytokines and can lead to a potentially fatal cytokine storm. The immune cell infiltration in the lungs can result in massive lung damage and multi-organ failure.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
The emergence of new and more infectious SARS-CoV-2 variants with mutations in the spike (S) protein and alternative viral entry mechanisms highlights the importance of a comprehensive understanding of viral entry into host cells, viral replication, and host cell response to the virus. A better knowledge of these aspects might help develop new therapeutics and vaccines to alleviate the present pandemic and prevent future viral outbreaks.
The SARS-CoV-2 spike protein binds to the angiotensin-converting enzyme 2 (ACE2) receptor on the host cell and mediates the virus's entry. RNA in situ mapping shows the presence of ACE2 receptors throughout the respiratory tract, especially in the nasal epithelium and in decreasing amounts throughout the lower respiratory tract. However, compared to ACE2 in the gastrointestinal tract, kidney, and myocardium, ACE2 expression is relatively low in the respiratory tract. Studies show that low ACE2 expression may be compensated by other factors to enhance viral entry.
Apart from ACE2 expression, cellular tropism of SARS-CoV-2 may be influenced by intrinsic and innate immune defenses. In vitro, SARS-CoV-2 infection models are few, use cell lines of non-human origin, and are engineered to overexpress ACE2.
Identifying new lung and upper airway cell models that are naturally supportive of SARS-CoV-2 infection
Researchers in the US recently infected a series of human lung and head/neck cancer cell lines that expressed varying levels of ACE2 and the TMPRSS2 protease. Their objective was to identify new lung and upper airway cell culture models that can be naturally permissive to SARS-CoV-2 infection. They found that human H522 lung adenocarcinoma cells, which do not express ACE2, support SARS-CoV-2 replication. The ACE2-independent infection of H522 cell line required the SARS-CoV-2 spike protein. The team’s findings have been released on the bioRxiv* preprint server.
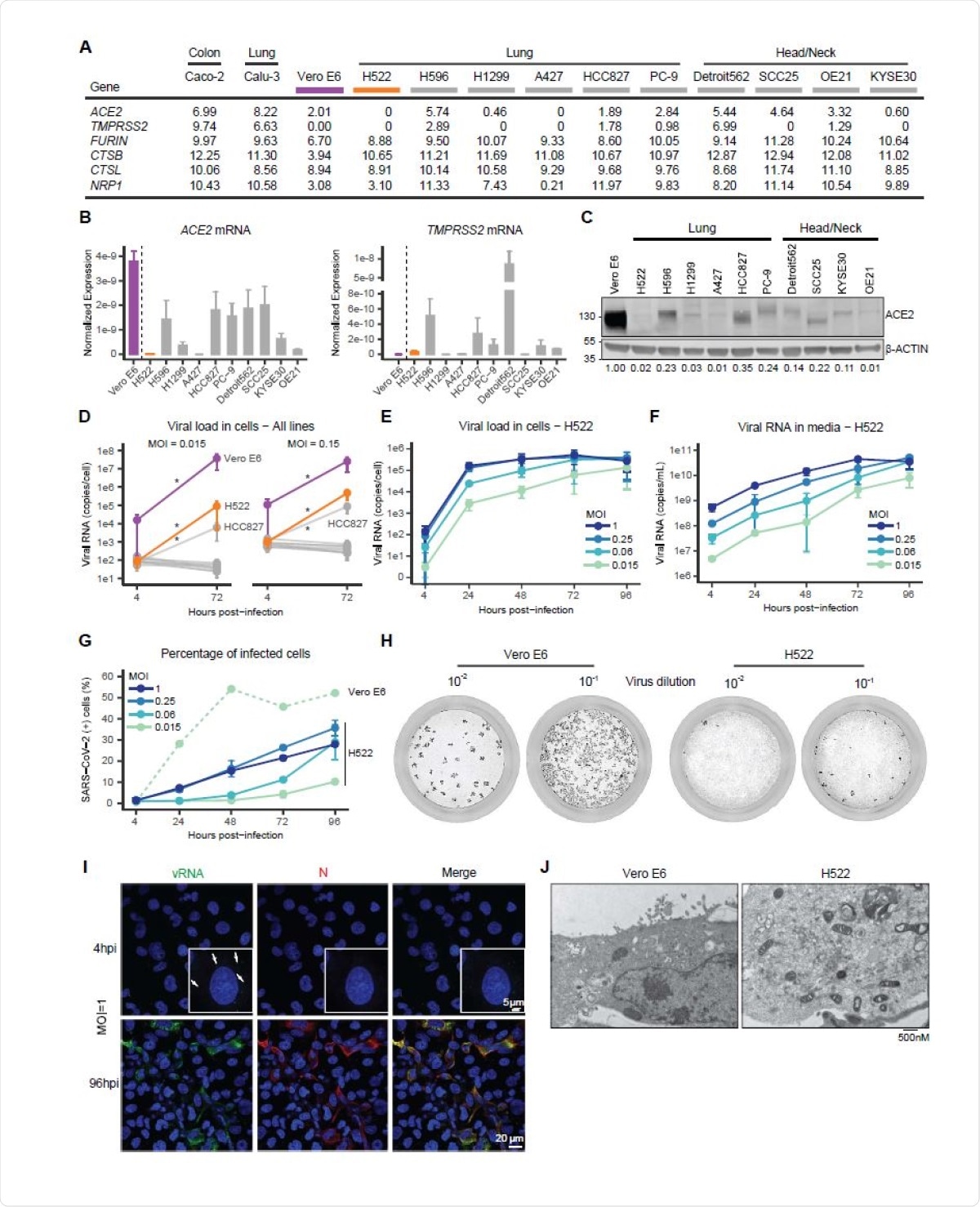
Expression of ACE2 across cell line models. A, Unsupervised hierarchical clustering of upper quartile-normalized RNA-seq reads. Normalized RNA-seq reads were aligned to the GRCh38 and Vervet-African green monkey genomes and quantified with Salmon (v1.3.0). As indicated, RNA-seq data were generated at UNC or Washington University or obtained from the Sequence Read Archive (SRA). B, Read counts for ACE2, GAPDH and ACTB across the indicated cell models.
With the help of CRISPR editing and neutralizing antibodies, the researchers confirmed the ACE2 independent nature of H522 infection, which is a significant finding that suggests an alternative receptor in a cell line of lung origin. Since ACE2 expression is relatively low in the respiratory system, it is possible that alternative receptors and/or attachment factors support viral entry. As recent findings by studies established that NRP1 and heparan sulfate are positive mediators of ACE2-dependent SARS-CoV-2 entry, it is not likely that they mediate ACE2-independent entry into H522 cells.
Taken together, H522 cells provide an alternative in vitro model to study SARS-CoV-2 infection and host innate immune responses.”
Another significant observation of this study is the inability of several lung and head/neck cancer cell lines to support replication of SARS-CoV-2 despite expressing ACE2 and TMPRSS2. The only exception was the HCC287 cells, which were less permissive to SARS-CoV-2 infection compared to the H522 and Vero E6 cells, though they express ACE2 and TMPRSS2. It is not clear yet if sialic acids act as entry receptors and/or attachment factors for SARS-CoV-2 in H522 cells.
The independence of virus replication from ACE2/TMPRSS2 in these cells indicates the utilization of an alternative receptor and entry pathway which may have functional relevance in understanding disease pathogenesis in vivo.”
Temporally resolved transcriptomic and proteomic profiling showed alterations in the cell cycle and the antiviral host cell response, including activation of type I interferon signaling mediated by MDA5. Chemical screens reveal important roles of clathrin-mediated endocytosis and endosomal cathepsins in the infection of H522 cells by SARS-CoV-2. These findings demonstrate the utilization of an alternative host cell receptor for SARS-CoV-2, and this may affect SARS-CoV-2 tropism and, thus human disease pathogenesis.
The inevitable emergence of novel coronaviruses utilizing variable entry pathways further underscores the importance of the H522 cell line model.”

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Maritza Puray-Chavez, Kyle M. LaPak, Travis P. Schrank, Jennifer L. Elliott, Dhaval P. Bhatt, Megan J. Agajanian, Ria Jasuja, Dana Q. Lawson, Keanu Davis, Paul W. Rothlauf, Heejoon Jo, Nakyung Lee, Kasyap Tenneti, Jenna E. Eschbach, Christian Shema Mugisha, Hung R. Vuong, Adam L. Bailey, D. Neil Hayes, Sean P.J. Whelan, Amjad Horani, Steven L. Brody, Dennis Goldfarb, M. Ben Major, Sebla B. Kutluay (2021) Systematic analysis of SARS-CoV-2 infection of an ACE2-negative human airway cell. bioRxiv preprint server. doi: https://doi.org/10.1101/2021.03.01.433431, https://www.biorxiv.org/content/10.1101/2021.03.01.433431v1
- Peer reviewed and published scientific report.
Puray-Chavez, Maritza, Kyle M. LaPak, Travis P. Schrank, Jennifer L. Elliott, Dhaval P. Bhatt, Megan J. Agajanian, Ria Jasuja, et al. 2021. “Systematic Analysis of SARS-CoV-2 Infection of an ACE2-Negative Human Airway Cell.” Cell Reports 36 (2): 109364. https://doi.org/10.1016/j.celrep.2021.109364. https://www.sciencedirect.com/science/article/pii/S2211124721007622.